Detailed information of T6SS
Note: only experimentally validated proteins are displayed in the graph.
Summary
| T6SS ID | T6SS00004 |
| T6SS type | Type i1 |
| Strain | Pseudomonas aeruginosa PAO1 |
| Replicon (RefSeq) | chromosome (H2-T6SS) (NC_002516) |
| Location | 1797958..1827821 |
| Function | anti-eukaryotic; anti-bacterial; effector translocation; virulence |
| Comment | H2-T6SS modulates internalization in epithelial cells; virulence in the worm model; virulence (slow killing) in Caenorhabditis elegans model; PldA achieves its antibacterial activity by degrading phosphatidylethanolamine; PldA contributes to the internalization of P. aeruginosa into Human Epithelial Cells and activates the PI3K/Akt Pathway during P. aeruginosa Infection. |
| References | [1] Russell AB et al (2013) Diverse type VI secretion phospholipases are functionally plastic antibacterial effectors. Nature. 496(7446):508-12. [PudMed:23552891] [2] Osipiuk J et al (2011) Crystal structure of secretory protein Hcp3 from Pseudomonas aeruginosa. J Struct Funct Genomics. 12(1):21-6. [PudMed:21476004] [3] Sana TG et al (2013) Divergent Control of Two Type VI Secretion Systems by RpoN in Pseudomonas aeruginosa. PLoS One. 8(10):e76030. [PudMed:24204589] [4] Sana TG et al (2015) Internalization of Pseudomonas aeruginosa Strain PAO1 into Epithelial Cells Is Promoted by Interaction of a T6SS Effector with the Microtubule Network. MBio. 6(3):e00712. [PudMed:26037124] [5] Sana TG et al (2012) The Second Type VI Secretion System of Pseudomonas aeruginosa Strain PAO1 Is Regulated by Quorum Sensing and Fur and Modulates Internalization in Epithelial Cells. J Biol Chem. 287(32):27095-105. [PudMed:22665491] [6] Jiang F et al (2014) A Pseudomonas aeruginosa Type VI Secretion Phospholipase D Effector Targets Both Prokaryotic and Eukaryotic Cells. Cell Host Microbe. 15(5):600-10. [PudMed:24832454] [7] Barret M et al (2011) Genomic analysis of the type VI secretion systems in Pseudomonas spp.: novel clusters and putative effectors uncovered. Microbiology. 157(Pt 6):1726-39. [PudMed:21474537] [8] Bleves S et al (2010) Protein secretion systems in Pseudomonas aeruginosa: A wealth of pathogenic weapons. Int J Med Microbiol. 300(8):534-43. [PudMed:20947426] [9] Hachani A et al (2011) Type VI secretion system in Pseudomonas aeruginosa: secretion and multimerization of VgrG proteins. J Biol Chem. 286(14):12317-27. [PudMed:21325275] |
T6SS components and genome coordinates
| Component | FHA | Sfa2 | Stk1 | Stp1 | TssA | TssB | TssC | TssE | TssF | TssG | TssH | TssJ | TssK | TssL | TssM |
| Number | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
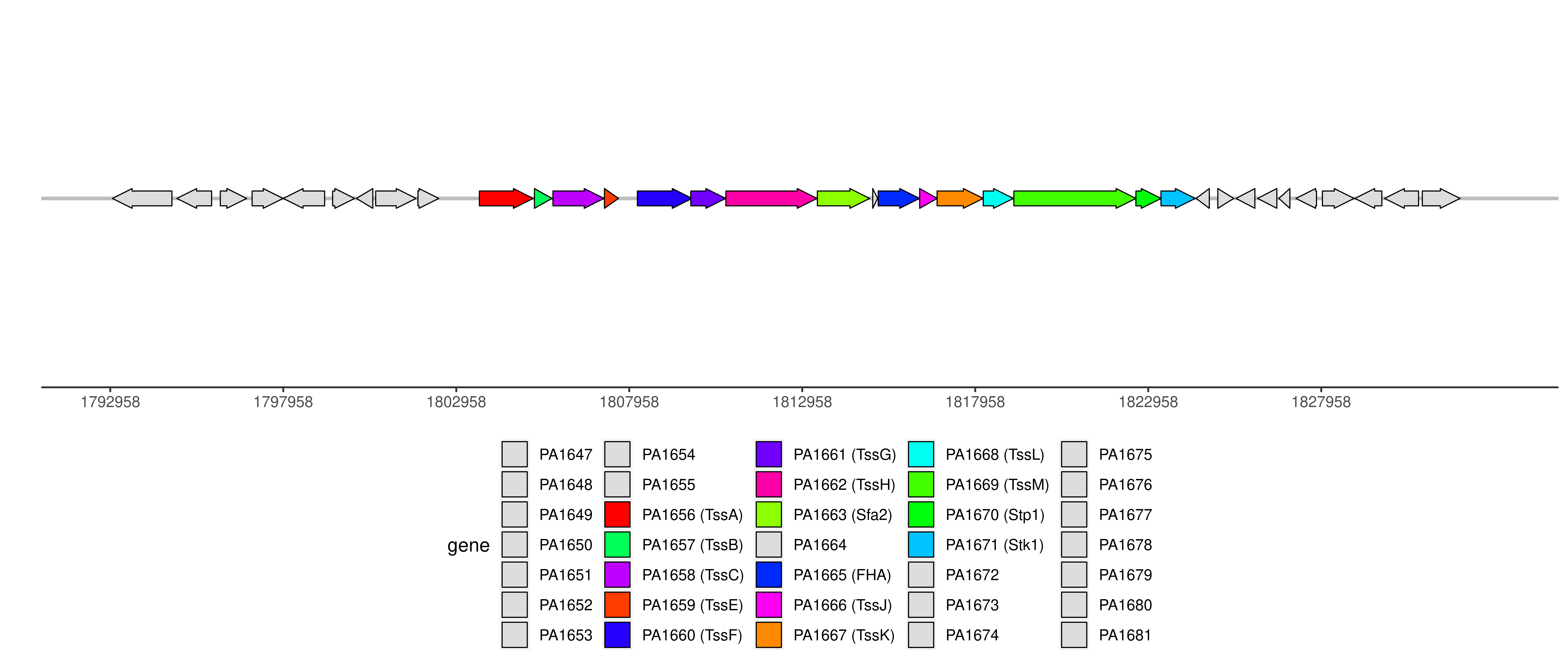
| Component ID | Component | Locus tag (Gene) | Coordinate | Strand | Size (bp) | NCBI ID | Product |
|---|---|---|---|---|---|---|---|
| PA1647 | 1793019..1794740 | - | 1722 | NP_250338.1 | sulfate transporter | ||
| PA1648 | 1794883..1795887 | - | 1005 | NP_250339.1 | oxidoreductase | ||
| PA1649 | 1796138..1796899 | + | 762 | NP_250340.1 | short-chain dehydrogenase | ||
| PA1650 | 1797058..1797951 | + | 894 | NP_250341.1 | transporter | ||
| PA1651 | 1797958..1799154 | - | 1197 | NP_250342.1 | transporter | ||
| PA1652 | 1799385..1800020 | + | 636 | NP_250343.1 | hypothetical protein | ||
| PA1653 | 1800071..1800547 | - | 477 | NP_250344.1 | transcriptional regulator | ||
| PA1654 | 1800629..1801795 | + | 1167 | NP_250345.1 | aminotransferase | ||
| PA1655 | 1801851..1802453 | + | 603 | NP_250346.1 | glutathione S-transferase | ||
| T6CP008377 | TssA | PA1656 | 1803626..1805182 | + | 1557 | NP_250347.1 | hypothetical protein |
| T6CP000011 | TssB | PA1657 | 1805218..1805724 | + | 507 | NP_250348.1 | hypothetical protein |
| T6CP000012 | TssC | PA1658 | 1805753..1807228 | + | 1476 | NP_250349.1 | hypothetical protein |
| T6CP008378 | TssE | PA1659 | 1807241..1807648 | + | 408 | NP_250350.1 | hypothetical protein |
| T6CP000013 | TssF | PA1660 | 1808193..1809773 | + | 1581 | NP_250351.1 | hypothetical protein |
| T6CP000014 | TssG | PA1661 | 1809737..1810744 | + | 1008 | NP_250352.1 | hypothetical protein |
| T6CP000015 | TssH | PA1662 | 1810751..1813384 | + | 2634 | NP_250353.1 | ClpA/B-type protease |
| T6CP012421 | Sfa2 | PA1663 | 1813395..1814906 | + | 1512 | NP_250354.1 | transcriptional regulator |
| PA1664 | 1814995..1815135 | + | 141 | NP_250355.1 | hypothetical protein | ||
| T6CP011362 | FHA | PA1665 | 1815153..1816346 | + | 1194 | NP_250356.1 | hypothetical protein |
| T6CP000016 | TssJ | PA1666 | 1816352..1816858 | + | 507 | NP_250357.1 | hypothetical protein |
| T6CP000017 | TssK | PA1667 | 1816855..1818186 | + | 1332 | NP_250358.1 | hypothetical protein |
| T6CP000018 | TssL | PA1668 | 1818189..1819058 | + | 870 | NP_250359.1 | hypothetical protein |
| T6CP000019 | TssM | PA1669 | 1819074..1822601 | + | 3528 | NP_250360.1 | hypothetical protein |
| T6CP012422 | Stp1 | PA1670 (stp1) | 1822601..1823329 | + | 729 | NP_250361.1 | serine/threonine phosphoprotein phosphatase Stp1 |
| T6CP012423 | Stk1 | PA1671 (stk1) | 1823326..1824315 | + | 990 | NP_250362.1 | serine-threonine kinase Stk1 |
| PA1672 | 1824343..1824723 | - | 381 | NP_250363.1 | hypothetical protein | ||
| PA1673 | 1824969..1825430 | + | 462 | NP_250364.1 | bacteriohemerythrin | ||
| PA1674 (folE2) | 1825495..1826040 | - | 546 | NP_250365.1 | GTP cyclohydrolase I | ||
| PA1675 | 1826118..1826675 | - | 558 | NP_250366.1 | hypothetical protein | ||
| PA1676 | 1826732..1827052 | - | 321 | NP_250367.1 | hypothetical protein | ||
| PA1677 | 1827225..1827821 | - | 597 | NP_250368.1 | hypothetical protein | ||
| PA1678 | 1827992..1828906 | + | 915 | NP_250369.1 | 50S ribosomal protein L3 glutamine methyltransferase | ||
| PA1679 | 1828931..1829710 | - | 780 | NP_250370.1 | hypothetical protein | ||
| PA1680 | 1829788..1830771 | - | 984 | NP_250371.1 | hypothetical protein | ||
| PA1681 (aroC) | 1830879..1831970 | + | 1092 | NP_250372.1 | chorismate synthase |
Download FASTA format files
Note: in the 'Target' column, ↓ means the target is positively regulated by the regulator, while ↑ indicates negatively regulation.
| Immunity protein ID | Locus tag (Gene) | Cognate effector | Organism | Replicon (Accession) | Coordinates (Strand) | NCBI ID |
|---|---|---|---|---|---|---|
| IMU00080 |
PA3488 (Tli5) | EFF00019 |
Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 3906391..3907089 (+) | 15598684 |
| IMU00314 |
PA1509 (TplEi) | EFF01463 |
Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 1638652..1639794 (-) | 15596706 |
| IMU00315 |
PA0261 (Tli3) | EFF01450; EFF01496 |
Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 293301..293798 (-) | 15595458 |
| Accessory protein ID | Locus tag (Gene) | Type | Organism | Replicon (Accession) | Coordinates (Strand) | NCBI ID | Related structure protein | Related effector |
|---|---|---|---|---|---|---|---|---|
| ACP00005 |
PA0259 (tla3) | adaptor | Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 289562..291004 (-) | 15595456 | PA0262 (VgrG2b) | PA0260 (Tle3) |
| ACP00043 |
PA4370 (IcmP) | chaperone | Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 4898193..4899533 (+) | 15599566 | - | PA1863 (ModA) |
| ACP00044 |
PA1862 (ModB) | chaperone | Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 2021712..2022398 (-) | 15597059 | - | PA1863 (ModA) |
| ACP00045 |
PA1861 (ModC) | chaperone | Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 2020625..2021710 (-) | 15597058 | - | PA1863 (ModA) |