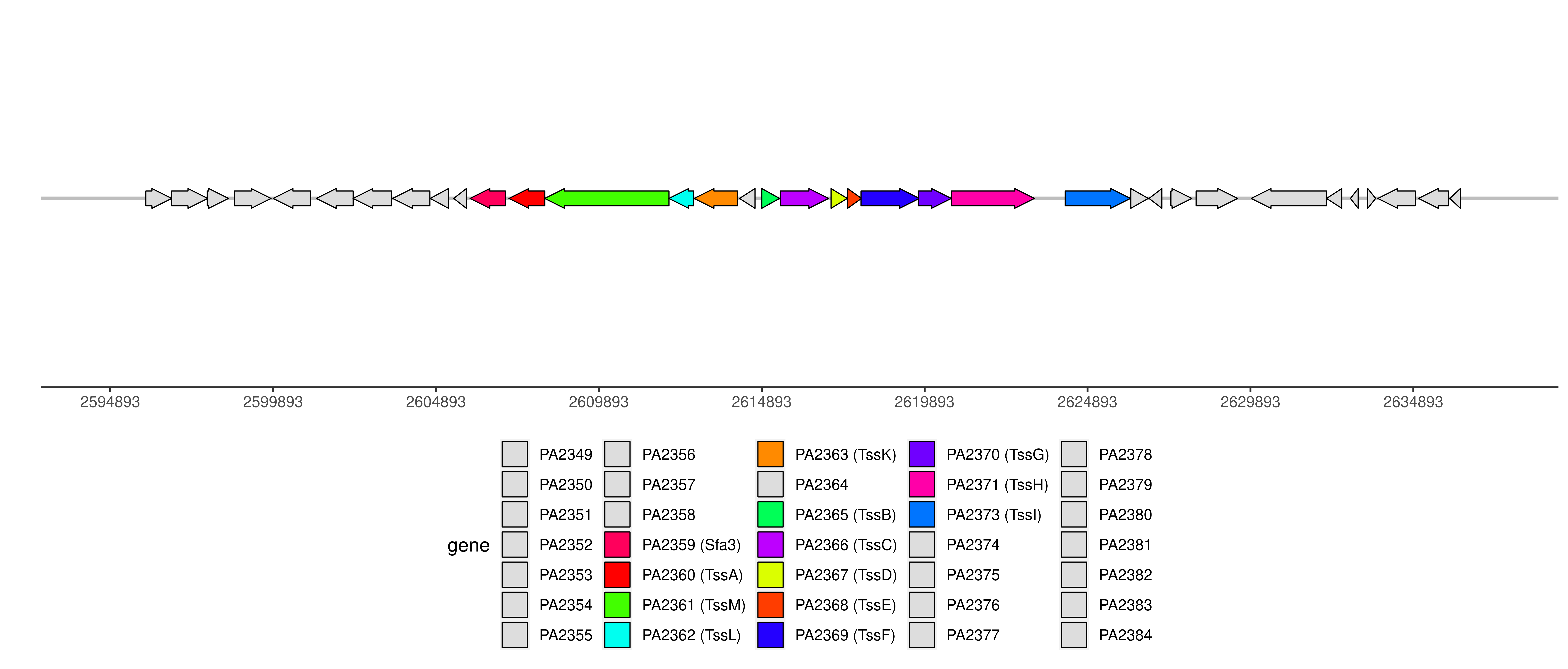
Detailed information of T6SS
Note: only experimentally validated proteins are displayed in the graph.
Summary
| T6SS ID | T6SS00005 |
| T6SS type | Type i4a |
| Strain | Pseudomonas aeruginosa PAO1 |
| Replicon (RefSeq) | chromosome (H3-T6SS) (NC_002516) |
| Location | 2599893..2632231 |
| Function | anti-eukaryotic; anti-bacterial; effector translocation; virulence |
| Comment | H3-T6SS is required for virulence in the worm model; PldB exerts an antibacterial activity and facilitates eukaryotic cells by activation of the PI3K/Akt pathway; PldB contributes to the internalization of P. aeruginosa into Human Epithelial cells. |
| References | [1] Hachani A et al (2011) Type VI secretion system in Pseudomonas aeruginosa: secretion and multimerization of VgrG proteins. J Biol Chem. 286(14):12317-27. [PudMed:21325275] [2] Kefala K et al (2012) Purification, crystallization and preliminary X-ray diffraction analysis of the C-terminal fragment of the MvfR protein from Pseudomonas aeruginosa. Acta Crystallogr Sect F Struct Biol Cryst Commun. 68(Pt 6):695-7. [PudMed:22684073] [3] Osipiuk J et al (2011) Crystal structure of secretory protein Hcp3 from Pseudomonas aeruginosa. J Struct Funct Genomics. 12(1):21-6. [PudMed:21476004] [4] Sana TG et al (2013) Divergent Control of Two Type VI Secretion Systems by RpoN in Pseudomonas aeruginosa. PLoS One. 8(10):e76030. [PudMed:24204589] [5] Jiang F et al (2014) A Pseudomonas aeruginosa Type VI Secretion Phospholipase D Effector Targets Both Prokaryotic and Eukaryotic Cells. Cell Host Microbe. 15(5):600-10. [PudMed:24832454] [6] Barret M et al (2011) Genomic analysis of the type VI secretion systems in Pseudomonas spp.: novel clusters and putative effectors uncovered. Microbiology. 157(Pt 6):1726-39. [PudMed:21474537] [7] Bleves S et al (2010) Protein secretion systems in Pseudomonas aeruginosa: A wealth of pathogenic weapons. Int J Med Microbiol. 300(8):534-43. [PudMed:20947426] [8] Chen Z et al (2014) Cloning, purification, crystallization and preliminary X-ray studies of the putative type VI secretion immunity protein Tli5 (PA5088) from Pseudomonas aeruginosa. Acta Crystallogr F Struct Biol Commun. 70(Pt 7):903-5. [PudMed:25005085] |
T6SS components and genome coordinates
| Component | PAAR | Sfa3 | TecT | TssA | TssB | TssC | TssD | TssE | TssF | TssG | TssH | TssI | TssK | TssL | TssM |
| Number | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| Component ID | Component | Locus tag (Gene) | Coordinate | Strand | Size (bp) | NCBI ID | Product |
|---|---|---|---|---|---|---|---|
| PA2349 | 2595985..2596779 | + | 795 | NP_251039.1 | hypothetical protein | ||
| PA2350 | 2596776..2597885 | + | 1110 | NP_251040.1 | methionine ABC transporter ATP-binding protein | ||
| PA2351 | 2597869..2598522 | + | 654 | NP_251041.1 | ABC transporter permease | ||
| PA2352 | 2598700..2599827 | + | 1128 | NP_251042.1 | glycerophosphoryl diester phosphodiesterase | ||
| PA2353 | 2599893..2601047 | - | 1155 | NP_251043.1 | hypothetical protein | ||
| PA2354 | 2601219..2602349 | - | 1131 | NP_251044.1 | transcriptional regulator | ||
| PA2355 | 2602346..2603530 | - | 1185 | NP_251045.1 | FMNH2-dependent monooxygenase | ||
| PA2356 (msuD) | 2603560..2604705 | - | 1146 | NP_251046.1 | methanesulfonate monooxygenase | ||
| PA2357 (msuE) | 2604715..2605275 | - | 561 | NP_251047.1 | FMN reductase MsuE | ||
| PA2358 | 2605435..2605824 | - | 390 | NP_251048.1 | hypothetical protein | ||
| T6CP012424 | Sfa3 | PA2359 | 2605937..2607022 | - | 1086 | NP_251049.1 | transcriptional regulator |
| T6CP000020 | TssA | PA2360 | 2607132..2608232 | - | 1101 | NP_251050.1 | hypothetical protein |
| T6CP000021 | TssM | PA2361 | 2608229..2612044 | - | 3816 | NP_251051.1 | hypothetical protein |
| T6CP008379 | TssL | PA2362 | 2612041..2612799 | - | 759 | NP_251052.1 | hypothetical protein |
| T6CP000022 | TssK | PA2363 | 2612817..2614148 | - | 1332 | NP_251053.1 | hypothetical protein |
| PA2364 | 2614208..2614684 | - | 477 | NP_251054.1 | hypothetical protein | ||
| T6CP000023 | TssB | PA2365 | 2614893..2615438 | + | 546 | NP_251055.1 | hypothetical protein |
| T6CP000024 | TssC | PA2366 | 2615461..2616945 | + | 1485 | NP_251056.1 | uricase |
| T6CP000025 | TssD | PA2367 | 2617019..2617516 | + | 498 | NP_251057.1 | hypothetical protein |
| T6CP008380 | TssE | PA2368 | 2617529..2617954 | + | 426 | NP_251058.1 | hypothetical protein |
| T6CP000026 | TssF | PA2369 | 2617938..2619731 | + | 1794 | NP_251059.1 | hypothetical protein |
| T6CP000027 | TssG | PA2370 | 2619695..2620711 | + | 1017 | NP_251060.1 | hypothetical protein |
| T6CP000028 | TssH | PA2371 | 2620713..2623262 | + | 2550 | NP_251061.1 | ClpA/B-type protease |
| T6CP000029 | TssI | PA2373 | 2624204..2626210 | + | 2007 | NP_251063.1 | hypothetical protein |
| PA2374 | 2626221..2626757 | + | 537 | NP_251064.1 | hypothetical protein | ||
| PA2375 | 2626780..2627175 | - | 396 | NP_251065.1 | hypothetical protein | ||
| PA2376 | 2627452..2628093 | + | 642 | NP_251066.1 | transcriptional regulator | ||
| PA2377 | 2628225..2629499 | + | 1275 | NP_251067.1 | hypothetical protein | ||
| PA2378 | 2629916..2632231 | - | 2316 | NP_251068.1 | aldehyde dehydrogenase | ||
| PA2379 | 2632228..2632698 | - | 471 | NP_251069.1 | oxidoreductase | ||
| PA2380 | 2632963..2633199 | - | 237 | NP_251070.1 | hypothetical protein | ||
| PA2381 | 2633494..2633736 | + | 243 | NP_251071.1 | hypothetical protein | ||
| PA2382 (lldA) | 2633803..2634954 | - | 1152 | NP_251072.1 | L-lactate dehydrogenase | ||
| PA2383 | 2635051..2635971 | - | 921 | NP_251073.1 | transcriptional regulator | ||
| PA2384 | 2636013..2636336 | - | 324 | NP_251074.1 | hypothetical protein |
Download FASTA format files
Note: in the 'Target' column, ↓ means the target is positively regulated by the regulator, while ↑ indicates negatively regulation.
| Regulator ID | Locus tag (Gene) | Regulator type | Regulatory pathway | Organism | Replicon (Accession) | Coordinates (Strand) | NCBI ID | Target (Regulation) |
|---|---|---|---|---|---|---|---|---|
| REG0018 |
PA4462 (RpoN) | protein | Global regulator | Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 4992870..4994363 (+) | 15599658 | PA1512 (hcpA) promotor (↓); PA5267 (hcpB) promotor (↓) |
| REG0070 |
pa1663 (sfa2) | protein | Global regulator; QS regulatory pathway | Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 1813395..1814906 (+) | 15596860 | PA4462 (rpoN) (↑); T6SS transcript (↑); T6SS transcript (↓) |
| Effector ID | Locus tag (Gene) | Classification | Cognate immunity protein | Organism | Replicon (Accession) | Coordinates (Strand) | NCBI ID |
|---|---|---|---|---|---|---|---|
| EFF01466 |
PA2374 | Tie | - | Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 2626221..2626757 (+) | 15597570 |
| EFF01482 |
PA3907 (TseT) | Tde | IMU00309 |
Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 4375230..4376015 (+) | 15599102 |
| EFF01279 |
PA5089 (PldB) | Tle | IMU00044 |
Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 5727239..5729476 (-) | 15600282 |
| EFF01893 |
PA0256 | Tse | - | Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 287188..288120 (-) | 15595453 |
| Immunity protein ID | Locus tag (Gene) | Cognate effector | Organism | Replicon (Accession) | Coordinates (Strand) | NCBI ID |
|---|---|---|---|---|---|---|
| IMU00044 |
PA5086 | EFF01279 |
Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 5724601..5725242 (-) | 15600279 |
| IMU00045 |
PA5087 | EFF01279 |
Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 5725479..5726348 (-) | 15600280 |
| IMU00046 |
PA5088 | EFF01279 |
Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 5726357..5727238 (-) | 15600281 |
| IMU00309 |
PA3908 | EFF01482 |
Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 4376012..4376731 (+) | 15599103 |
| Accessory protein ID | Locus tag (Gene) | Type | Organism | Replicon (Accession) | Coordinates (Strand) | NCBI ID | Related structure protein | Related effector |
|---|---|---|---|---|---|---|---|---|
| ACP00006 |
PA3906 (co-TecT) | chaperone | Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 4374850..4375233 (+) | 15599101 | PA3905 (TecT) | - |
| ACP00007 |
PA3905 (tecT) | chaperone | Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 4374330..4374857 (+) | 15599100 | PA3904 (PAAR4) | PA3907 (TseT) |
| ACP00008 |
PA3790 (oprC) | chaperone | Pseudomonas aeruginosa PAO1 | chromosome (NC_002516) | 4247703..4249874 (-) | 15598985 | - | PA4922 (Azu) |