Detailed information of TA system
Overview
TA module
| Type | I | Classification (family/domain) | ldrD-rdlD/- |
| Location | 1271280..1271500 | Replicon | chromosome |
| Accession | NC_007779 | ||
| Organism | Escherichia coli str. K-12 substr. W3110 | ||
| T1TAdb ID | TA06175 | ||
Toxin (Protein)
| Gene name | ldrD | Uniprot ID | J7R083 |
| Locus tag | Y75_p1189 | Protein ID | WP_000170963.1 |
| Coordinates | 1271280..1271387 (-) | Length | 36 a.a. |
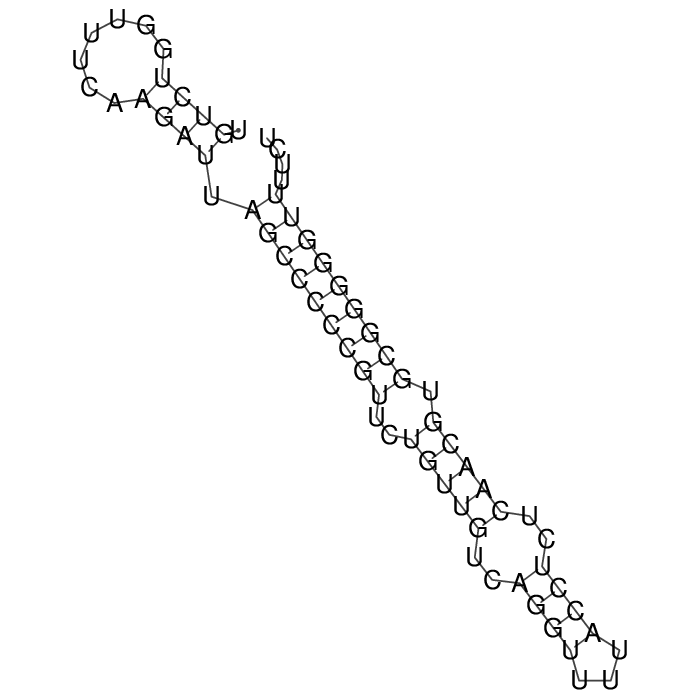
Antitoxin (RNA)
| Gene name | rdlD | ||
| Locus tag | Y75_s0009 | ||
| Coordinates | 1271434..1271500 (+) |
Genomic Context
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| Y75_RS06335 (Y75_p1183) | 1266589..1267671 | + | 1083 | WP_000804726.1 | peptide chain release factor 1 | - |
| Y75_RS06340 (Y75_p1184) | 1267671..1268504 | + | 834 | WP_000456467.1 | peptide chain release factor N(5)-glutamine methyltransferase | - |
| Y75_RS06345 (Y75_p1185) | 1268501..1268893 | + | 393 | WP_000200374.1 | invasion regulator SirB2 | - |
| Y75_RS06350 (Y75_p1186) | 1268897..1269706 | + | 810 | WP_001257044.1 | invasion regulator SirB1 | - |
| Y75_RS06355 (Y75_p1187) | 1269742..1270596 | + | 855 | WP_000811065.1 | 3-deoxy-8-phosphooctulonate synthase | - |
| Y75_RS06360 (Y75_p1188) | 1270745..1270852 | - | 108 | WP_000170955.1 | small toxic polypeptide LdrA/LdrC | - |
| Y75_RS06365 (Y75_p1189) | 1271280..1271387 | - | 108 | WP_000170963.1 | small toxic polypeptide LdrB | Toxin |
| - | 1271434..1271500 | + | 67 | - | - | Antitoxin |
| Y75_RS06370 (Y75_p1190) | 1271815..1271922 | - | 108 | WP_000170955.1 | small toxic polypeptide LdrA/LdrC | - |
| Y75_RS06375 (Y75_p1191) | 1272326..1273426 | - | 1101 | WP_000063607.1 | sodium-potassium/proton antiporter ChaA | - |
| Y75_RS06380 (Y75_p1192) | 1273696..1273926 | + | 231 | WP_001146444.1 | putative cation transport regulator ChaB | - |
| Y75_RS06385 (Y75_p1193) | 1274084..1274779 | + | 696 | WP_001336325.1 | glutathione-specific gamma-glutamylcyclotransferase | - |
| Y75_RS06390 (Y75_p1194) | 1274823..1275176 | - | 354 | WP_001169669.1 | DsrE/F sulfur relay family protein YchN | - |
Associated MGEs
| MGE detail |
Similar MGEs |
Relative position |
MGE Type | Cargo ARG | Virulence gene | Coordinates | Length (bp) |
|---|
Relative position:
(1) inside: TA loci is completely located inside the MGE;
(2) overlap: TA loci is partially overlapped with the MGE;
(3) flank: The TA loci is located in the 5 kb flanking regions of MGE.
Sequences
Toxin
Download Length: 36 a.a. Molecular weight: 3971.74 Da Isoelectric Point: 11.4779
>T6342 WP_000170963.1 NC_007779:c1271387-1271280 [Escherichia coli str. K-12 substr. W3110]
MTLAQFAMTFWHDLAAPILAGIITAAIVGWWRNRK
MTLAQFAMTFWHDLAAPILAGIITAAIVGWWRNRK
Download Length: 108 bp
>T6342 NC_007779:c1271387-1271280 [Escherichia coli str. K-12 substr. W3110]
ATGACGCTCGCGCAGTTTGCCATGACTTTCTGGCACGACCTGGCGGCACCGATCCTGGCGGGAATTATTACCGCAGCGAT
TGTCGGCTGGTGGCGTAACCGGAAGTAA
ATGACGCTCGCGCAGTTTGCCATGACTTTCTGGCACGACCTGGCGGCACCGATCCTGGCGGGAATTATTACCGCAGCGAT
TGTCGGCTGGTGGCGTAACCGGAAGTAA
Antitoxin
Download Length: 67 bp
>AT6342 NC_007779:1271434-1271500 [Escherichia coli str. K-12 substr. W3110]
TGTCTGGTTTCAAGATTAGCCCCCGTTCTGTTGTCAGGTTTTACCTCTCAACGTGCGGGGGTTTTCT
TGTCTGGTTTCAAGATTAGCCCCCGTTCTGTTGTCAGGTTTTACCTCTCAACGTGCGGGGGTTTTCT
Similar Proteins
Only experimentally validated proteins are listed.
Multiple sequence alignment
| Protein | Organism | Identities (%) | Coverage (%) | Ha-value |
|---|
References
(1) Mitsuoki Kawano et al. (2002) Molecular characterization of long direct repeat (LDR) sequences expressing a stable mRNA encoding for a 35-amino-acid cell-killing peptide and a cis-encoded small antisense RNA in Escherichia coli. Molecular Microbiology 45(2):333-49. [PubMed:12123448]
(2) Elizabeth M Fozo et al. (2010) Abundance of type I toxin-antitoxin systems in bacteria: searches for new candidates and discovery of novel families. Nucleic Acids Research 38(11):3743-59. [PubMed:20156992]