Detailed information of TA system
Overview
TA module
| Type | I | Classification (family/domain) | hok-sok/- |
| Location | 2565..2828 | Replicon | chromosome |
| Accession | X03774 | ||
| Organism | Hafnia alvei | ||
| T1TAdb ID | TA07129 | ||
Toxin (Protein)
| Gene name | hokH | Uniprot ID | - |
| Locus tag | - | Protein ID | - |
| Coordinates | 2565..2723 (-) | Length | 53 a.a. |
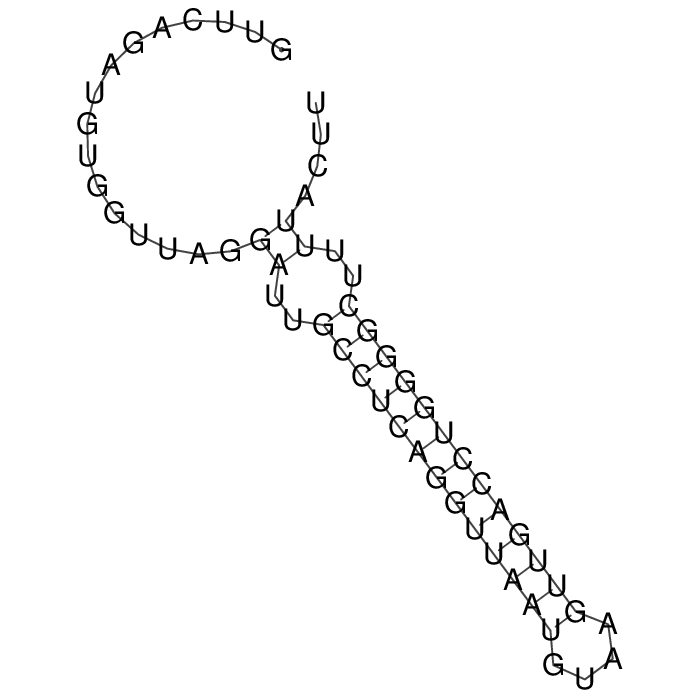
Antitoxin (RNA)
| Gene name | sokH | ||
| Locus tag | - | ||
| Coordinates | 2771..2828 (+) |
Genomic Context
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| Locus_0 | 185..2404 | + | 2220 | CAA27400.1 | - | - |
| - | 2565..2723 | - | 159 | - | - | Toxin |
| - | 2771..2828 | + | 58 | - | - | Antitoxin |
Associated MGEs
| MGE detail |
Similar MGEs |
Relative position |
MGE Type | Cargo ARG | Virulence gene | Coordinates | Length (bp) |
|---|
Relative position:
(1) inside: TA loci is completely located inside the MGE;
(2) overlap: TA loci is partially overlapped with the MGE;
(3) flank: The TA loci is located in the 5 kb flanking regions of MGE.
Sequences
Toxin
Download Length: 53 a.a. Molecular weight: 6065.37 Da Isoelectric Point: 9.2226
>T10210 - X03774:c2723-2565 [Hafnia alvei]
MSRKLLLYGLIVMCFTLLIFTWMVRGSLCELRIKQGKTEVAAFLNYEDKHAL
MSRKLLLYGLIVMCFTLLIFTWMVRGSLCELRIKQGKTEVAAFLNYEDKHAL
Download Length: 159 bp
>T10210 X03774:c2723-2565 [Hafnia alvei]
ATGTCGCGTAAGCTGTTGCTTTACGGATTAATAGTGATGTGTTTTACGCTATTGATCTTTACCTGGATGGTACGTGGTTC
GCTGTGTGAGCTACGAATTAAGCAGGGAAAAACAGAGGTTGCCGCGTTCTTAAACTACGAAGATAAGCACGCGTTGTAA
ATGTCGCGTAAGCTGTTGCTTTACGGATTAATAGTGATGTGTTTTACGCTATTGATCTTTACCTGGATGGTACGTGGTTC
GCTGTGTGAGCTACGAATTAAGCAGGGAAAAACAGAGGTTGCCGCGTTCTTAAACTACGAAGATAAGCACGCGTTGTAA
Antitoxin
Download Length: 58 bp
>AT10210 X03774:2771-2828 [Hafnia alvei]
GTTCAGATGTGGTTAGGATTGCCTCAGGTTAATGTAAGTTGACCTGGGGCTTTTACTT
GTTCAGATGTGGTTAGGATTGCCTCAGGTTAATGTAAGTTGACCTGGGGCTTTTACTT
Similar Proteins
Only experimentally validated proteins are listed.
| Protein | Organism | Identities (%) | Coverage (%) | Ha-value |
|---|---|---|---|---|
| T10208 | Escherichia coli C |
68.182 |
88 |
0.6 |
| T10124 | Erwinia amylovora ATCC 49946 |
62.069 |
90.625 |
0.563 |
| T6370 | Escherichia coli |
52 |
100 |
0.52 |
| T6363 | uncultured bacterium |
47.917 |
92.308 |
0.442 |
| T6324 | Escherichia coli |
47.917 |
92.308 |
0.442 |
| T10039 | Escherichia coli O157:H7 str. Sakai |
42.857 |
96.078 |
0.412 |
| T6369 | Escherichia coli K-12 |
43.478 |
93.878 |
0.408 |
| T10209 | Escherichia coli strain ECOR24 |
39.583 |
96 |
0.38 |
| T10211 | E.coli NADPH-sulfite reductase flavoprotein component (cysJ) NADPH-sulfite reductase hemoprotein component (cysI) and 3' |
40.909 |
89.796 |
0.367 |
Multiple sequence alignment
| Protein | Organism | Identities (%) | Coverage (%) | Ha-value |
|---|
References
(1) K Pedersen et al. (1999) Multiple hok genes on the chromosome of Escherichia coli. Molecular Microbiology 32(5):1090-102. [PubMed:10361310]