
| 7 | |
| Agrobacterium tumefaciens plasmid Ti (NC_002377) | |
| plasmid Ti [Browse all T4SS(s) in this replicon] | |
| NC_002377 | |
| 151232..172619 | |
| VirB | |
| effector translocation and conjugation | |
| Type IVA; Type P | |
T4SS components |
| Component | VirB1 | VirB2 | VirB3 | VirB4 | VirB5 | VirB6 | VirB7 | VirB8 | VirB9 |
| Number | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| Component | VirB10 | VirB11 | VirD4 | ||||||
| Number | 1 | 1 | 1 |
The information of T4SS components from NC_002377 | ||||||
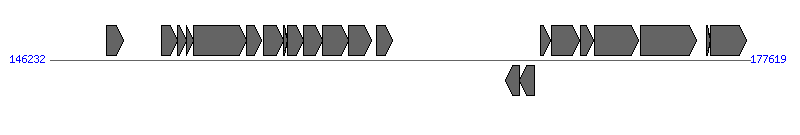 | ||||||
| # | Locus tag (Gene) | Coordinates [+/-], size (bp) | Protein GI | Product | Component | |
| 1 | pTi_126 (virJ) | 148762..149529 [+], 768 | 10955142 | hypothetical protein | ||
| 2 | pTi_127 (virB1) | 151232..151951 [+], 720 | 10955143 | type IV secretion system lytic transglycosylase VirB1 | VirB1 | |
| 3 | pTi_128 (virB2) | 151966..152331 [+], 366 | 10955144 | type IV secretion system pilin subunit VirB2 | VirB2 | |
| 4 | pTi_129 (virB3) | 152331..152657 [+], 327 | 10955145 | type IV secretion system protein VirB3 | VirB3 | |
| 5 | pTi_130 (virB4) | 152657..155026 [+], 2370 | 10955146 | type IV secretion system protein VirB4 | VirB4 | |
| 6 | pTi_131 (virB5) | 155041..155703 [+], 663 | 10955147 | type IV secretion system protein VirB5 | VirB5 | |
| 7 | pTi_132 (virB6) | 155800..156687 [+], 888 | 10955148 | type IV secretion system protein VirB6 | VirB6 | |
| 8 | pTi_133 (virB7) | 156721..156888 [+], 168 | 10955149 | type IV secretion system lipoprotein VirB7 | VirB7 | |
| 9 | pTi_134 (virB8) | 156896..157588 [+], 693 | 10955150 | type IV secretion system protein VirB8 | VirB8 | |
| 10 | pTi_135 (virB9) | 157585..158466 [+], 882 | 10955151 | type IV secretion system protein VirB9 | VirB9 | |
| 11 | pTi_136 (virB10) | 158463..159596 [+], 1134 | 10955152 | type IV secretion system protein VirB10 | VirB10 | |
| 12 | pTi_137 (virB11) | 159636..160667 [+], 1032 | 10955153 | type IV secretion system protein VirB11 | VirB11 | |
| 13 | pTi_138 (virG) | 160877..161602 [+], 726 | 10955154 | two-component response regulator VirG | ||
| 14 | pTi_139 (virC2) | 166669..167277 [-], 609 | 10955155 | putative crown gall tumor protein VirC2 | ||
| 15 | pTi_140 (virC1) | 167280..167975 [-], 696 | 10955156 | putative crown gall tumor protein VirC1 | ||
| 16 | pTi_141 (virD1) | 168244..168687 [+], 444 | 10955157 | type IV secretion system T-DNA border endonuclease VirD1 | ||
| 17 | pTi_142 (virD2) | 168721..169995 [+], 1275 | 10955158 | type IV secretion system T-DNA border endonuclease VirD2 | ||
| 18 | pTi_143 (virD3) | 170015..170620 [+], 606 | 10955159 | hypothetical protein | ||
| 19 | pTi_144 (virD4) | 170649..172619 [+], 1971 | 10955160 | type IV secretion system protein VirD4 | VirD4 | |
| 20 | pTi_145 (virD5) | 172711..175212 [+], 2502 | 10955161 | hypothetical protein | ||
| 21 | pTi_146 (virE1) | 175665..175862 [+], 198 | 10955162 | type IV secretion system chaperone VirE1 | ||
| 22 | pTi_147 (virE2) | 175867..177468 [+], 1602 | 10955163 | type IV secretion system single-stranded DNA binding protein VirE2 | ||
Download FASTA format files |
| Proteins Genes |
| This system delivers both DNA and proteins into plant cell. The transferred DNA (T-DNA) region: NC_002377: 1..22878) |
Effectors |
| VirD2, VirD5, VirE2, VirE3, VirF |
The information of protein effectors | ||||||
| # | Locus tag (Gene) | Coordinates [+/-], size (bp) | Protein GI | Product | * | |
| 1 | pTi_142 (virD2) | 168721..169995 [+], 1275 | 10955158 | type IV secretion system T-DNA border endonuclease VirD2 | ||
| 2 | pTi_145 (virD5) | 172711..175212 [+], 2502 | 10955161 | hypothetical protein | ||
| 3 | pTi_147 (virE2) | 175867..177468 [+], 1602 | 10955163 | type IV secretion system single-stranded DNA binding protein VirE2 | ||
| 4 | pTi_148 (virE3) | 177534..179552 [+], 2019 | 10955164 | hypothetical protein | ||
| 5 | pTi_152 (virF) | 184764..185372 [+], 609 | 10955168 | hypothetical protein | ||
Download FASTA format files |
| Proteins Genes |
| # | Accessory protein(GI) | motif(s) | Substrate(s) | Function | Reference |
| 1 | VirC1 (10955156) | ND | VirD2 | VirC1 pairwise interacts with the processing factors VirD2 relaxase, VirC2, and VirD1 to stimulate generation of multiple copies per cell of the T-DNA substrate. VirC1 promotes conjugative T-DNA transfer. | (1) PubMed: 17505518 |
| 2 | VirE1 chaperone (10955162) | CT50aa sufficient (interacts with the VirD4 T4CP. CT100aa(VirE2) sufficient for the T4CP interation) | VirE2 | Translocation of the VirE2 effector through the A. tumefaciens VirB/VirD4 T4SS requires VirE1. | (2) PubMed: 19946141 |
|
(1) Atmakuri K; Cascales E; Burton OT; Banta LM; Christie PJ (2007). Agrobacterium ParA/MinD-like VirC1 spatially coordinates early conjugative DNA transfer reactions. EMBO J. 26(10):2540-51. [PudMed:17505518] |
| (2) Alvarez-Martinez CE; Christie PJ (2009). Biological diversity of prokaryotic type IV secretion systems. Microbiol Mol Biol Rev. 73(4):775-808. [PudMed:19946141] |
| (1) Ream W (2009). Agrobacterium tumefaciens and A. rhizogenes use different proteins to transport bacterial DNA into the plant cell nucleus. Microb Biotechnol. 2(4):416-27. [PudMed:21255274] |