Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 200376 |
| Name | oriT_ICEPmiChnBCP11 |
| Organism | Proteus mirabilis integrating |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | MG773277 (3772..4070 [+], 299 nt) |
| oriT length | 299 nt |
| IRs (inverted repeats) | 234..242, 254..262 (AAAGCCAAA..TTTGGCTTT) 45..50, 52..57 (CAAACG..CGTTTG) 7..12, 26..31 (TTTGGC..GCCAAA) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 299 nt
>oriT_ICEPmiChnBCP11
GCTCTGTTTGGCGGCTGGTGACCTAGCCAAAAAAATTGAGACGCCAAACGTCGTTTGCATTCTGGCCTGAACTTGCCAAACGGTTTGTATCTTCATGGCGATACGTCTTTTTAGGTGTTTTTAAGTGAAATCAGCCTGTATCCCTTGTCGGGTATGGGATTGAGCGAGTCGATTTATATCGAGACGCCAAACAGTGATTGTGACGGCAGTTTTACGTTTGGCGTTTCGATCCAAAAGCCAAACGGATAGTGGTTTTGGCTTTAGGGGTTAATTGGATGGGGAAATTGGTTTGGTAGAAA
GCTCTGTTTGGCGGCTGGTGACCTAGCCAAAAAAATTGAGACGCCAAACGTCGTTTGCATTCTGGCCTGAACTTGCCAAACGGTTTGTATCTTCATGGCGATACGTCTTTTTAGGTGTTTTTAAGTGAAATCAGCCTGTATCCCTTGTCGGGTATGGGATTGAGCGAGTCGATTTATATCGAGACGCCAAACAGTGATTGTGACGGCAGTTTTACGTTTGGCGTTTCGATCCAAAAGCCAAACGGATAGTGGTTTTGGCTTTAGGGGTTAATTGGATGGGGAAATTGGTTTGGTAGAAA
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file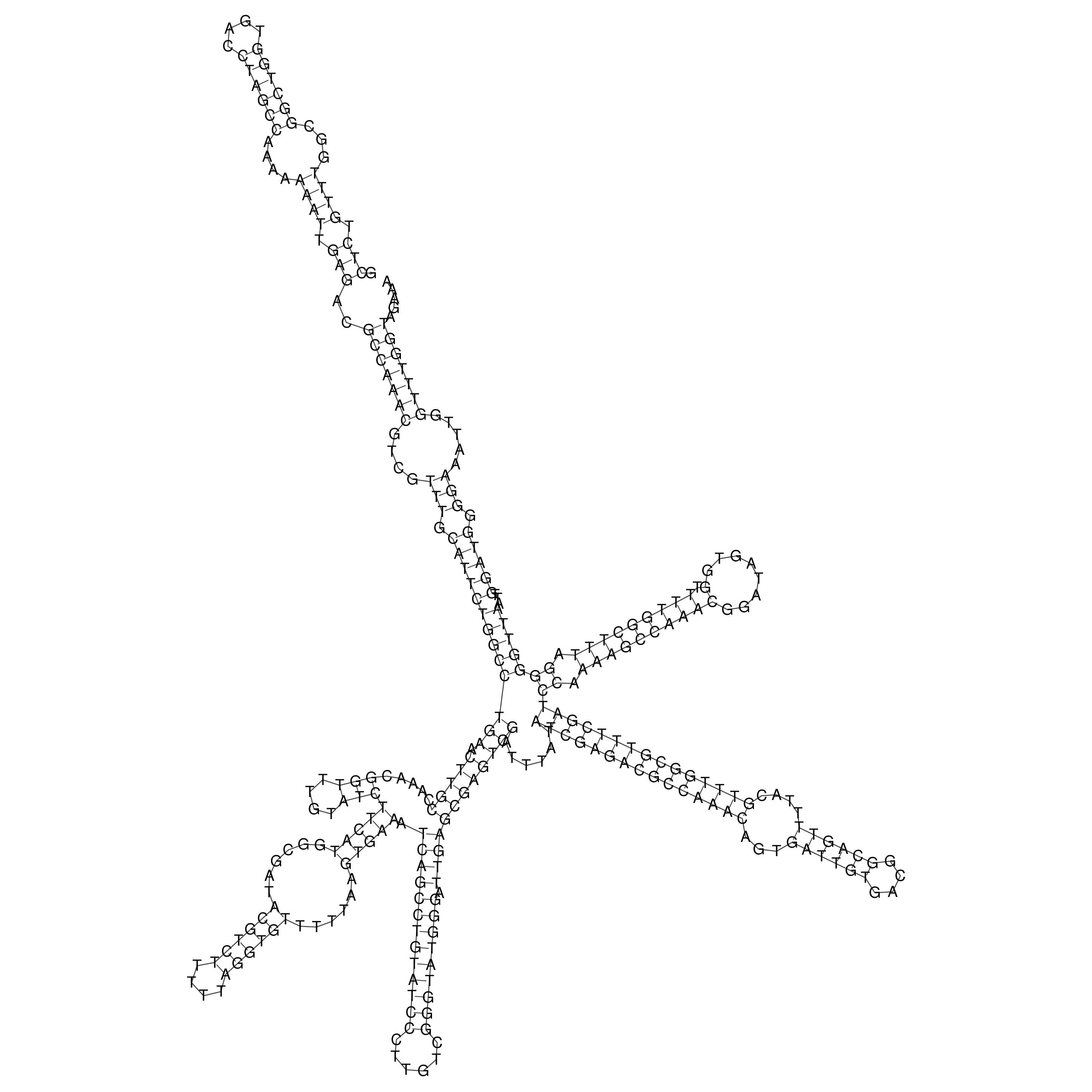
Relaxase
| ID | 14402 | GenBank | AWW18560 |
| Name | traI_-_ICEPmiChnBCP11 |
UniProt ID | _ |
| Length | 716 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 716 a.a. Molecular weight: 79760.84 Da Isoelectric Point: 5.2184
>AWW18560.1 TraI [Proteus mirabilis]
MFKNLFFQAKALPELSSQLDAEIPRYPPFLKGLPAASPEDLQSTQDELIAKLRQVLGFNQRDFQRLIQPC
IDHLAAYVHLLPASEHHHHSGAGGLLRHSLEVAFWAAQAAEGIIFVASGTPVEKKELEPRWRVAAALGGL
FHDIGKPVSDLSITDEDGRYQWNPFLETLSQWTSNNSIERYFIRWRDGRCKRHEQFSILVLNRVMTPELL
AWLTQPGPEILQAMLEAIGNTDPEHVLSKLVIEADQTSVQRDLKAQRISVDDNALGVPVERYLLDAMRRL
LASSQWLVNQRDARVWIRKSNQSTHLYLVWKSAAKDIIELLAKDKIPGIPRDPDTLADILIERGLATKSA
SNERYESLAPEVLIKDDKPIWLAMLHIVEADLLFSSNVPSSVRLFSKSEWEATPQTQAEPQSRSSEQPGL
PDASSSIEHGNSSESPSTKSSDQDDELRLASDVNNPQANENAPGDGCENPNNSYDGAISNNVNQHDAEAF
NLPESLAWLPEASSALVMVDEQIMIRYPDAVRHWCAPRKLLAELSRLDWLELDPANPTRKARTVTTKDGV
QEQGLLLKVSISKGLTALIDLPKQDAEPAAAIQNEEALQRPSRTKTTNAQAKEPATRAERKQKPIGPNAN
SSTDPKHAQRQQMVNFVKDLPILLTDGDYPYVDHSADGIRVTIQTLRQVAKEHGIPAGQLLRGISASDEC
QFDEGETVLFTARAER
MFKNLFFQAKALPELSSQLDAEIPRYPPFLKGLPAASPEDLQSTQDELIAKLRQVLGFNQRDFQRLIQPC
IDHLAAYVHLLPASEHHHHSGAGGLLRHSLEVAFWAAQAAEGIIFVASGTPVEKKELEPRWRVAAALGGL
FHDIGKPVSDLSITDEDGRYQWNPFLETLSQWTSNNSIERYFIRWRDGRCKRHEQFSILVLNRVMTPELL
AWLTQPGPEILQAMLEAIGNTDPEHVLSKLVIEADQTSVQRDLKAQRISVDDNALGVPVERYLLDAMRRL
LASSQWLVNQRDARVWIRKSNQSTHLYLVWKSAAKDIIELLAKDKIPGIPRDPDTLADILIERGLATKSA
SNERYESLAPEVLIKDDKPIWLAMLHIVEADLLFSSNVPSSVRLFSKSEWEATPQTQAEPQSRSSEQPGL
PDASSSIEHGNSSESPSTKSSDQDDELRLASDVNNPQANENAPGDGCENPNNSYDGAISNNVNQHDAEAF
NLPESLAWLPEASSALVMVDEQIMIRYPDAVRHWCAPRKLLAELSRLDWLELDPANPTRKARTVTTKDGV
QEQGLLLKVSISKGLTALIDLPKQDAEPAAAIQNEEALQRPSRTKTTNAQAKEPATRAERKQKPIGPNAN
SSTDPKHAQRQQMVNFVKDLPILLTDGDYPYVDHSADGIRVTIQTLRQVAKEHGIPAGQLLRGISASDEC
QFDEGETVLFTARAER
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4CP
| ID | 16914 | GenBank | AWW18561 |
| Name | traD_-_ICEPmiChnBCP11 |
UniProt ID | _ |
| Length | 606 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 606 a.a. Molecular weight: 67102.94 Da Isoelectric Point: 6.2511
>AWW18561.1 TraD [Proteus mirabilis]
MKENAYEMPWRTNYEAMAAAGWLVGATGAIAAEMLTELPPEPFWWMTGISSGMALYRLPEAYLLYKLQMG
LKGKPLAFMELSHLQKVMAKHPDELWLGYGFEWDQRHAQRAYEILKRDKQTLLNQGHGKQMGSTWIHGVE
PKEEDVYQPVGHTEGHTLIVGTTGAGKTRCFDAMITQAILRNEAVIIIDPKGDKELKDNAQRACIAAGSP
ERFVYFHPGFPEHSVRLNPLRNFNRGTEIASRIAALIPSETGADPFKAFGQMALNNIVQGLLLTSQRPDL
KTLRRFLEGGPEGLVVKAVTAWGEQVYPNFSVEIKRFTEKTNTLAKQAMAMLLFYYERIQPIAANTDLEG
LLSMFEHDRTHYSKMVASLMPVLNMLTSSELGPLLSPIANDVDDSRLITDSGRIINNAQVAYIGLDSLTD
AMVGSAIGSLLLSDLTAVAGDRYNYGVDNRPVNIFIDEAAEVVNDPFIQLLNKGRGAKMRCVIATQTFAD
FAARTGSEAKARQVLGNINNLIALRVMDAETQQYITDNLPKTRLQYIMQTQGMSSNSDSPALFTGNHGER
LMEEEGDMFPPQLLGQLPNLEYIAKLSGGRVIKGRIPILTSSTQAA
MKENAYEMPWRTNYEAMAAAGWLVGATGAIAAEMLTELPPEPFWWMTGISSGMALYRLPEAYLLYKLQMG
LKGKPLAFMELSHLQKVMAKHPDELWLGYGFEWDQRHAQRAYEILKRDKQTLLNQGHGKQMGSTWIHGVE
PKEEDVYQPVGHTEGHTLIVGTTGAGKTRCFDAMITQAILRNEAVIIIDPKGDKELKDNAQRACIAAGSP
ERFVYFHPGFPEHSVRLNPLRNFNRGTEIASRIAALIPSETGADPFKAFGQMALNNIVQGLLLTSQRPDL
KTLRRFLEGGPEGLVVKAVTAWGEQVYPNFSVEIKRFTEKTNTLAKQAMAMLLFYYERIQPIAANTDLEG
LLSMFEHDRTHYSKMVASLMPVLNMLTSSELGPLLSPIANDVDDSRLITDSGRIINNAQVAYIGLDSLTD
AMVGSAIGSLLLSDLTAVAGDRYNYGVDNRPVNIFIDEAAEVVNDPFIQLLNKGRGAKMRCVIATQTFAD
FAARTGSEAKARQVLGNINNLIALRVMDAETQQYITDNLPKTRLQYIMQTQGMSSNSDSPALFTGNHGER
LMEEEGDMFPPQLLGQLPNLEYIAKLSGGRVIKGRIPILTSSTQAA
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
| ID | 16915 | GenBank | AWW18575 |
| Name | traC_-_ICEPmiChnBCP11 |
UniProt ID | _ |
| Length | 799 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 799 a.a. Molecular weight: 90171.75 Da Isoelectric Point: 5.9727
>AWW18575.1 TraC [Proteus mirabilis]
MKASLYSGQRASELLPVLAYSDDEQLFFMEDQSVGFGFLCDPLPGGDESVADRVNVLLNNDWPKDTLLQF
GLYASPDIQTDLQRMMGLRHRQSDPLLRASIRKRADFLDGGTVQPIEESTQTQVRNFQLIVTCKLPLESP
IPTDRELSRASALRASFSQALATVGFRVTEMTDRNWLAALSAQLNWGKDASWRNPSPIRSEADKPLREQV
LDYDRAIKVDSQGLMLGDYRVKTLSFKRLPERIWFGHAASFAGDMMTGSRGLRGSFLLNVTIHFPSAEAM
RSRLETKRQWAVNQAYGPMLKFVPVLAAKKKGFDVLFEALQEGDRPIRANMTLTLFSPTEEASISSVSNA
RTYFKELGFELMEDKYFCLPIFLNALPFGADRQAMNDLFRFKTMATRHIIPLLPLFADWKGTGTPVINFV
SRNGQIMSVSLYDSGSNYNCCIAAQSGSGKSFLVNEIISSYLSEGGQCWVIDVGRSYEKLCEVYDGEFLQ
FGRDSGICLNPFEIVEDYDEEADVLVGLLAAMAAPTQSLTDFQMANLKRQTRELWEKKGRAMLVDDVAEA
LKNHEDRRVQDVGEQLYPFTTQGEYGRFFNGHNNIRFKNRFTVLELEELKGRKHLQQVVLLQLIYQIQQE
MYLGERDRRKIVFIDEAWDLLTQGDVGKFIETGYRRFRKYGGSAVTVTQSVNDLYDSPTGKAIAENSANM
YLLGQKAETINALKKEGRLPLGEGGYEYLKTVHTVTGVYSEIFFITEMGTGIGRLIVDPFHKLLYSSRAE
DVNAIKQLTRKGLSVADAISQLLKERGYE
MKASLYSGQRASELLPVLAYSDDEQLFFMEDQSVGFGFLCDPLPGGDESVADRVNVLLNNDWPKDTLLQF
GLYASPDIQTDLQRMMGLRHRQSDPLLRASIRKRADFLDGGTVQPIEESTQTQVRNFQLIVTCKLPLESP
IPTDRELSRASALRASFSQALATVGFRVTEMTDRNWLAALSAQLNWGKDASWRNPSPIRSEADKPLREQV
LDYDRAIKVDSQGLMLGDYRVKTLSFKRLPERIWFGHAASFAGDMMTGSRGLRGSFLLNVTIHFPSAEAM
RSRLETKRQWAVNQAYGPMLKFVPVLAAKKKGFDVLFEALQEGDRPIRANMTLTLFSPTEEASISSVSNA
RTYFKELGFELMEDKYFCLPIFLNALPFGADRQAMNDLFRFKTMATRHIIPLLPLFADWKGTGTPVINFV
SRNGQIMSVSLYDSGSNYNCCIAAQSGSGKSFLVNEIISSYLSEGGQCWVIDVGRSYEKLCEVYDGEFLQ
FGRDSGICLNPFEIVEDYDEEADVLVGLLAAMAAPTQSLTDFQMANLKRQTRELWEKKGRAMLVDDVAEA
LKNHEDRRVQDVGEQLYPFTTQGEYGRFFNGHNNIRFKNRFTVLELEELKGRKHLQQVVLLQLIYQIQQE
MYLGERDRRKIVFIDEAWDLLTQGDVGKFIETGYRRFRKYGGSAVTVTQSVNDLYDSPTGKAIAENSANM
YLLGQKAETINALKKEGRLPLGEGGYEYLKTVHTVTGVYSEIFFITEMGTGIGRLIVDPFHKLLYSSRAE
DVNAIKQLTRKGLSVADAISQLLKERGYE
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 38606..63427
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| Locus_26 | 35069..36250 | + | 1182 | AWW18559 | hypothetical protein | - |
| Locus_27 | 36407..38557 | + | 2151 | AWW18560 | TraI | - |
| Locus_28 | 38606..40426 | + | 1821 | AWW18561 | TraD | virb4 |
| Locus_29 | 40436..40996 | + | 561 | AWW18562 | hypothetical protein | - |
| s043 | 40983..41618 | + | 636 | AWW18563 | Conjugative transfer protein S043 | tfc7 |
| Locus_31 | 41690..42316 | + | 627 | AWW18564 | hypothetical protein | - |
| Locus_32 | 42528..43427 | + | 900 | AWW18565 | putative Zn peptidase | - |
| Locus_33 | 43564..43845 | + | 282 | AWW18566 | TraL | traL |
| Locus_34 | 44076..44468 | + | 393 | AWW18567 | TraE | traE |
| Locus_35 | 44449..45348 | + | 900 | AWW18568 | TraK | traK |
| Locus_36 | 45351..46640 | + | 1290 | AWW18569 | TraB | traB |
| Locus_37 | 46637..47287 | + | 651 | AWW18570 | HtdD | traV |
| Locus_38 | 47284..47670 | + | 387 | AWW18571 | TraA | - |
| Locus_39 | 47849..48682 | + | 834 | AWW18572 | Ynd | - |
| Locus_40 | 48675..49613 | + | 939 | AWW18573 | Ync | - |
| Locus_41 | 49745..50437 | + | 693 | AWW18574 | thiol:disulfide interchange protein DsbC | trbB |
| Locus_42 | 50437..52836 | + | 2400 | AWW18575 | TraC | virb4 |
| Locus_43 | 53160..53672 | + | 513 | AWW18576 | TrbI | - |
| Locus_44 | 53683..54807 | + | 1125 | AWW18577 | TraW | traW |
| Locus_45 | 54791..55819 | + | 1029 | AWW18578 | TraU | traU |
| Locus_46 | 55822..59514 | + | 3693 | AWW18579 | TraN | traN |
| Locus_47 | 59973..61391 | + | 1419 | AWW18580 | hypothetical protein | - |
| Locus_48 | 61388..63427 | + | 2040 | AWW18581 | hypothetical protein | virb4 |
| Locus_49 | 64552..65766 | - | 1215 | AWW18582 | putative transposase TnpA | - |
| Locus_50 | 65863..66375 | - | 513 | AWW18583 | putative lipoprotein signal peptidase LspA | - |
| Locus_51 | 66379..67275 | - | 897 | AWW18584 | putative cation-efflux family protein | - |
| Locus_52 | 67371..67778 | + | 408 | AWW18585 | putative MerR family regulator | - |
Host bacterium
| ID | 333 | Element type | ICE (Integrative and conjugative element) |
| Element name | ICEPmiChnBCP11 | GenBank | MG773277 |
| Element size | 141263 bp | Coordinate of oriT [Strand] | 3772..4070 [+] |
| Host bacterium | Proteus mirabilis integrating | Coordinate of element | 1..141263 |
Cargo genes
| Drug resistance gene | floR, aph(6)-Id, aph(3'')-Ib, sul2, tet(C), cfr, blaCTX-M-65, aph(3')-Ia, aac(6')-Ib-cr, blaOXA-1, catB3, ARR-3, qacE, sul1, fosA3, aph(4)-Ia, aac(3)-IVa, dfrA32, ere(A), aadA2 |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |