Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 200122 |
| Name | oriT_ICEValE0601 |
| Organism | Vibrio alginolyticus strain E0601 sequence |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | KT072768 (5548..5846 [+], 299 nt) |
| oriT length | 299 nt |
| IRs (inverted repeats) | 234..242, 254..262 (AAAGCCAAA..TTTGGCTTT) 45..50, 52..57 (CAAACG..CGTTTG) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 299 nt
>oriT_ICEValE0601
GCTCTGTTTGGCGGCGAATGACTGAGTCAAAAAAATCGAGACGCCAAACGTCGTTTGCATTCTGGCCTGAACTCGCCAAACGGTTTGTATCTTCATGGCGATACGTCTTTTTAGGCGCTTTTAAGTGAAATCGGCCTGTATCCCTTGTCATGTATGGGATTGCGCGAGTTGATTTATATCGAGACGCCAAACAGTGATTGTTACGGCAGTTTTACGTTTGGCGTTTCGATCCAAAAGCCAAACGGATAGTGGTTTTGGCTTTCGGGGTTAATTGGATGGGGAAATTGGTTTGGTAGAAA
GCTCTGTTTGGCGGCGAATGACTGAGTCAAAAAAATCGAGACGCCAAACGTCGTTTGCATTCTGGCCTGAACTCGCCAAACGGTTTGTATCTTCATGGCGATACGTCTTTTTAGGCGCTTTTAAGTGAAATCGGCCTGTATCCCTTGTCATGTATGGGATTGCGCGAGTTGATTTATATCGAGACGCCAAACAGTGATTGTTACGGCAGTTTTACGTTTGGCGTTTCGATCCAAAAGCCAAACGGATAGTGGTTTTGGCTTTCGGGGTTAATTGGATGGGGAAATTGGTTTGGTAGAAA
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file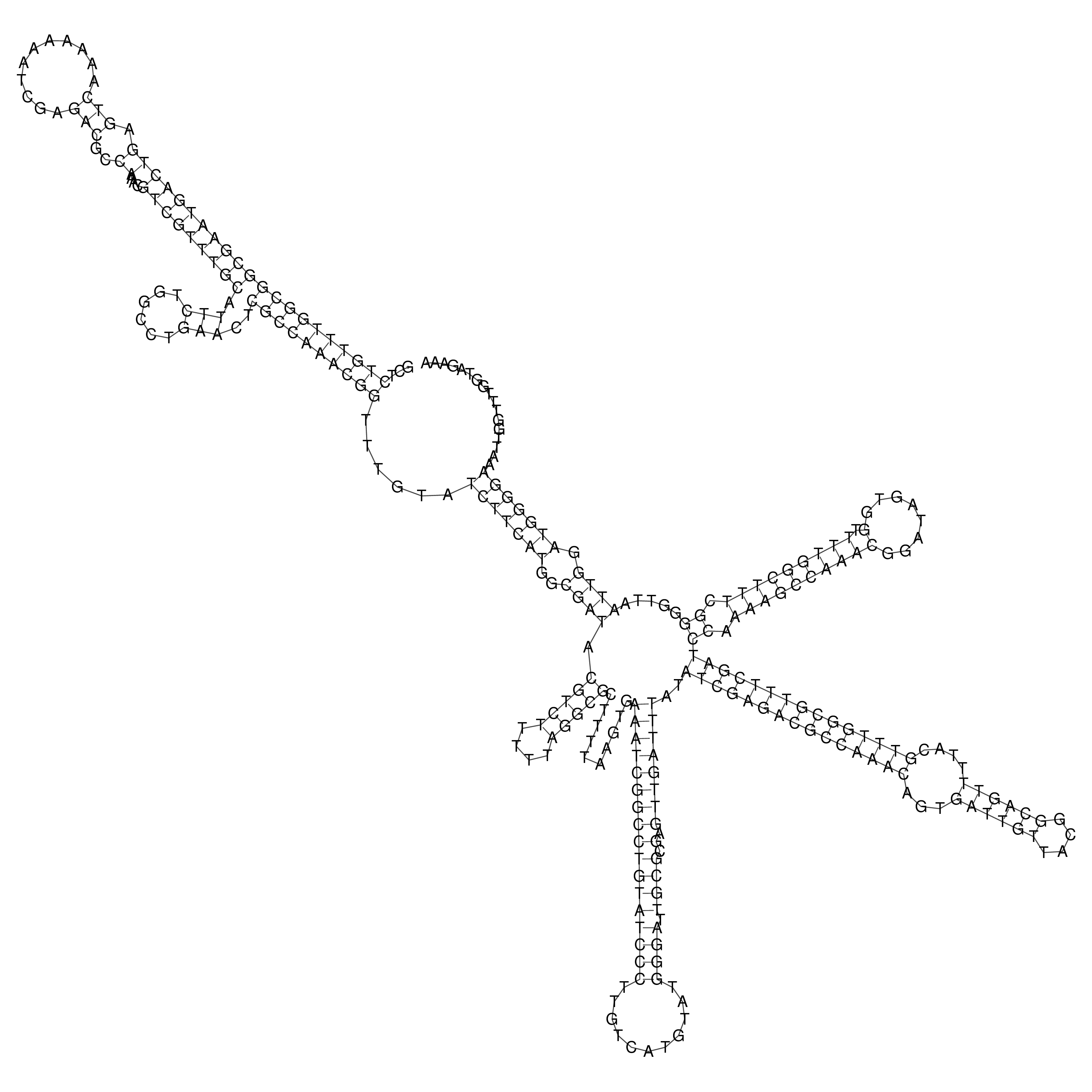
Relaxase
| ID | 14275 | GenBank | ALF34804 |
| Name | traI_ICEValE0601_046_ICEValE0601 |
UniProt ID | _ |
| Length | 726 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 726 a.a. Molecular weight: 80776.67 Da Isoelectric Point: 5.0517
>ALF34804.1 Conjugative transfer protein, TraI [Vibrio alginolyticus]
MPFTKDDTTFAGVFMLSRFLKSQHQPESTESEIPRYPPFLKGLPAASPEDLQSTQDELISKLRQVLGFNQ
RDFQRLIQPCIDHLAAYVHLLPASEHHHHSGAGGLLRHSLEVAFWAAQAAEGIIFVASGTPVEKKEQEPR
WRVAAALGGLFHDIGKPVSDLSITDEDGRYLWNPFLETLSQWTTNNSIERYFIRWRDGRCKRHEQFSILV
LNRVMTPELLAWLTQPGPEILQAMLEAIGNTDPEHVLSKLVIEADQTSVQRDLKAQRISVDDNALGVPVE
RYLLDAMRRLLASSQWLVNQRDARVWIRKSNQSTNLYLVWKSAAKEIIELLAKDKIPGIPRDPDTLADIL
IERGLATKSASNERYESFAPEVLIKDGKPIWLPMLHISEADLLFSSNVPSGVTLFNKSEWEATQQTQAEQ
QSRSSEHSDLPEASSSIEHGNSSESPSTNSSDQDDELRLASDVNNPQANENAPGDGCEKPNNSYDGAISN
NVNQHDAEALNLPESLAWLPEASSALVVVGEQILFRYPDAVRPWCAPRKLLAELSRLDWLELDPANPTRK
ARTITTKDGVQEQGLLLKVSISKGLTSLIDLPKQDAESAAAIQNEEALLRPSRTKTTNAQAKEPATRAER
KQKPIVANANSSTDPKHAQRQQMVSFVKDLPILLTDGDYPDVDHSADGIRVTIQTLRQVANEHGIPAGQL
LRGISASDECQFDEGETVLFTAHAER
MPFTKDDTTFAGVFMLSRFLKSQHQPESTESEIPRYPPFLKGLPAASPEDLQSTQDELISKLRQVLGFNQ
RDFQRLIQPCIDHLAAYVHLLPASEHHHHSGAGGLLRHSLEVAFWAAQAAEGIIFVASGTPVEKKEQEPR
WRVAAALGGLFHDIGKPVSDLSITDEDGRYLWNPFLETLSQWTTNNSIERYFIRWRDGRCKRHEQFSILV
LNRVMTPELLAWLTQPGPEILQAMLEAIGNTDPEHVLSKLVIEADQTSVQRDLKAQRISVDDNALGVPVE
RYLLDAMRRLLASSQWLVNQRDARVWIRKSNQSTNLYLVWKSAAKEIIELLAKDKIPGIPRDPDTLADIL
IERGLATKSASNERYESFAPEVLIKDGKPIWLPMLHISEADLLFSSNVPSGVTLFNKSEWEATQQTQAEQ
QSRSSEHSDLPEASSSIEHGNSSESPSTNSSDQDDELRLASDVNNPQANENAPGDGCEKPNNSYDGAISN
NVNQHDAEALNLPESLAWLPEASSALVVVGEQILFRYPDAVRPWCAPRKLLAELSRLDWLELDPANPTRK
ARTITTKDGVQEQGLLLKVSISKGLTSLIDLPKQDAESAAAIQNEEALLRPSRTKTTNAQAKEPATRAER
KQKPIVANANSSTDPKHAQRQQMVSFVKDLPILLTDGDYPDVDHSADGIRVTIQTLRQVANEHGIPAGQL
LRGISASDECQFDEGETVLFTAHAER
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4CP
| ID | 16536 | GenBank | ALF34805 |
| Name | traD_ICEValE0601_047_ICEValE0601 |
UniProt ID | _ |
| Length | 606 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 606 a.a. Molecular weight: 67084.85 Da Isoelectric Point: 6.5130
>ALF34805.1 Conjugative transfer protein, TraD [Vibrio alginolyticus]
MKENAYEMPWRTNYEAMAAAGWLVGATGAIAAELLTELPPEPFWWMTGISSGMALYRLPEAYRLYKLQKG
LKGKPLAFMELSHLQKVMAKHPDELWLGYGFEWDQRHAQRAYEILKRDKQTLLNQGHGKQMGSTWIHGVE
PKEEDVYQPVGHTEGHTLIVGTTGAGKTRCFDAMITQAILRNEAVIIIDPKGDKELKDNAQRACIAAGSP
ERFVYFHPGFPEHSVRLNPLRNFNRGTEIASRIAALIPSETGADPFKAFGQMALNNIVQGLLLTSQRPDL
KTLRRFLEGGPEGLVVKAVTAWGEQVYPNFSVEIKRFTEKANTLAKQAMAMLLFYYERIQPVAANTDLEG
LLSMFEHDRTHYSKMVASLMPVLNMLTSSELGPLLSPIANDVDDSRLITDSGRIINNAQVAYIGLDSLTD
AMVGSAIGSLLLSDLTAVAGDRYNYGVENRPVNIFIDEAAEVVNDPFIQLLNKGRGAKMRCVIATQTFAD
FAARTGSEAKARQVLGNINNLIALRVMDAETQQYITDNLPKTRLQYIMQTQGMSSNSDSPALFTGNHGER
LMEEEGDMFPPQLLGQLSNLEYIAKLSGGRVIKGRIPILTSSTQAA
MKENAYEMPWRTNYEAMAAAGWLVGATGAIAAELLTELPPEPFWWMTGISSGMALYRLPEAYRLYKLQKG
LKGKPLAFMELSHLQKVMAKHPDELWLGYGFEWDQRHAQRAYEILKRDKQTLLNQGHGKQMGSTWIHGVE
PKEEDVYQPVGHTEGHTLIVGTTGAGKTRCFDAMITQAILRNEAVIIIDPKGDKELKDNAQRACIAAGSP
ERFVYFHPGFPEHSVRLNPLRNFNRGTEIASRIAALIPSETGADPFKAFGQMALNNIVQGLLLTSQRPDL
KTLRRFLEGGPEGLVVKAVTAWGEQVYPNFSVEIKRFTEKANTLAKQAMAMLLFYYERIQPVAANTDLEG
LLSMFEHDRTHYSKMVASLMPVLNMLTSSELGPLLSPIANDVDDSRLITDSGRIINNAQVAYIGLDSLTD
AMVGSAIGSLLLSDLTAVAGDRYNYGVENRPVNIFIDEAAEVVNDPFIQLLNKGRGAKMRCVIATQTFAD
FAARTGSEAKARQVLGNINNLIALRVMDAETQQYITDNLPKTRLQYIMQTQGMSSNSDSPALFTGNHGER
LMEEEGDMFPPQLLGQLSNLEYIAKLSGGRVIKGRIPILTSSTQAA
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
| ID | 16537 | GenBank | ALF34817 |
| Name | traC_ICEValE0601_061_ICEValE0601 |
UniProt ID | _ |
| Length | 799 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 799 a.a. Molecular weight: 90131.69 Da Isoelectric Point: 5.9714
>ALF34817.1 Conjugative transfer pilus assembly protein, TraC [Vibrio alginolyticus]
MKASLYSGQRASELLPVLAYSDDEQLFFMEDQSVGFGFLCDPLPGGDESVADRVNVLLNNDWPTDTLLQF
GLYSSPDIQTDLQRMMGLRHRQSDPLLRASIRKRADFLDGGTVQPIEESTQTQVRNFQLIVTCKLPLENP
IPTERELSRASALRASFSQALATVGFRVTEMTDRNWLAALSAQLNWGKDASWRNPSLIRSEADKPLREQV
LDYDRAIKVDSQGLMLGDYRVKTLSFKRLPERIWFGHAASFAGDMMTGSRGLRGSFLLNVTIHFPSAEAM
RSRLETKRQWAVNQAYGPMLKFVPVLAAKKKGFDVLFEALQEGDRPIRANMTLTLFSPTEEASISSVSNA
RTYFKELGFELMEDKYFCLPIFLNALPFGADRQAMNDLFRFKTMATRHVIPLLPLFADWKGTGTPVINFV
SRNGQIMSVSLYDSGSNYNCCIAAQSGSGKSFLVNEIISSYLSEGGQCWVIDVGRSYEKLCEVYDGEFLQ
FGRDSGICLNPFEIVEDYDEEADVLVGLLAAMAAPTQSLSDFQMANLKRQTRELWEKKGRAMLVDDVAAA
LKNHEDRRVQDVGEQLYPFTTQGEYGRFFNGHNNIRFKNRFTVLELEELKGRKHLQQVVLLQLIYQIQQE
MYLGERDRRKIVFIDEAWDLLTQGDVGKFIETGYRRFRKYGGSAVTVTQSVNDLYDSPTGKAIAENSANM
YLLGQKAETINALKKEGRLPLGEGGYEYLKTVHTVTGVYSEIFFITEMGTGIGRLIVDPFHKLLYSSRAE
DVNAIKQLTRKGLSVADAISQLLKERGYE
MKASLYSGQRASELLPVLAYSDDEQLFFMEDQSVGFGFLCDPLPGGDESVADRVNVLLNNDWPTDTLLQF
GLYSSPDIQTDLQRMMGLRHRQSDPLLRASIRKRADFLDGGTVQPIEESTQTQVRNFQLIVTCKLPLENP
IPTERELSRASALRASFSQALATVGFRVTEMTDRNWLAALSAQLNWGKDASWRNPSLIRSEADKPLREQV
LDYDRAIKVDSQGLMLGDYRVKTLSFKRLPERIWFGHAASFAGDMMTGSRGLRGSFLLNVTIHFPSAEAM
RSRLETKRQWAVNQAYGPMLKFVPVLAAKKKGFDVLFEALQEGDRPIRANMTLTLFSPTEEASISSVSNA
RTYFKELGFELMEDKYFCLPIFLNALPFGADRQAMNDLFRFKTMATRHVIPLLPLFADWKGTGTPVINFV
SRNGQIMSVSLYDSGSNYNCCIAAQSGSGKSFLVNEIISSYLSEGGQCWVIDVGRSYEKLCEVYDGEFLQ
FGRDSGICLNPFEIVEDYDEEADVLVGLLAAMAAPTQSLSDFQMANLKRQTRELWEKKGRAMLVDDVAAA
LKNHEDRRVQDVGEQLYPFTTQGEYGRFFNGHNNIRFKNRFTVLELEELKGRKHLQQVVLLQLIYQIQQE
MYLGERDRRKIVFIDEAWDLLTQGDVGKFIETGYRRFRKYGGSAVTVTQSVNDLYDSPTGKAIAENSANM
YLLGQKAETINALKKEGRLPLGEGGYEYLKTVHTVTGVYSEIFFITEMGTGIGRLIVDPFHKLLYSSRAE
DVNAIKQLTRKGLSVADAISQLLKERGYE
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 40445..60725
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| ICEValE0601_044 | 36115..37824 | + | 1710 | ALF34802 | Type III restriction endonuclease | - |
| ICEValE0601_045 | 37860..38162 | - | 303 | ALF34803 | Hypothetical protein | - |
| ICEValE0601_046 | 38216..40396 | + | 2181 | ALF34804 | Conjugative transfer protein, TraI | - |
| ICEValE0601_047 | 40445..42265 | + | 1821 | ALF34805 | Conjugative transfer protein, TraD | virb4 |
| ICEValE0601_048 | 42275..42835 | + | 561 | ALF34806 | Conjugative transfer protein, S091 | - |
| ICEValE0601_049 | 42822..43457 | + | 636 | ALF34807 | Conjugative transfer protein, TraJ | tfc7 |
| ICEValE0601_050 | 43484..44071 | - | 588 | ALF34808 | Hypothetical protein | - |
| ICEValE0601_051 | 44360..44641 | + | 282 | ALF34809 | Conjugative transfer pilus assembly protein, TraL | traL |
| ICEValE0601_052 | 44650..45264 | + | 615 | ALF34810 | Conjugative transfer pilus assembly protein, TraE | traE |
| ICEValE0601_053 | 45248..46144 | + | 897 | ALF34811 | Conjugative transfer pilus assembly protein, TraK | traK |
| ICEValE0601_054 | 46147..47436 | + | 1290 | ALF34812 | Conjugative transfer pilus assembly protein, TraB | traB |
| ICEValE0601_055 | 47511..48083 | + | 573 | ALF34813 | Conjugative transfer protein, TraV | traV |
| ICEValE0601_056 | 48080..48466 | + | 387 | ALF34814 | Conjugative transfer protein, TraA | - |
| ICEValE0601_057 | 48507..49013 | - | 507 | ALF34867 | Acetyltransferase | - |
| ICEValE0601_058 | 49004..49270 | - | 267 | ALF34879 | Hypothetical protein | - |
| ICEValE0601_059 | 49581..50684 | - | 1104 | ALF34815 | Transposase | - |
| ICEValE0601_060 | 50955..51647 | + | 693 | ALF34816 | Thiol:disulfide involved in conjugative transfer,DsbC | trbB |
| ICEValE0601_061 | 51648..54047 | + | 2400 | ALF34817 | Conjugative transfer pilus assembly protein, TraC | virb4 |
| ICEValE0601_062 | 54040..54387 | + | 348 | ALF34818 | Conjugative transfer protein 345 | - |
| ICEValE0601_063 | 54440..54883 | + | 444 | ALF34819 | Conjugative signal peptidase, TrhF | - |
| ICEValE0601_064 | 54894..56018 | + | 1125 | ALF34820 | Conjugative transfer pilus assembly protein, TraW | traW |
| ICEValE0601_065 | 56050..57030 | + | 981 | ALF34821 | Conjugative transfer pilus assembly protein, TraU | traU |
| ICEValE0601_066 | 57033..60725 | + | 3693 | ALF34822 | Conjugative transfer protein, TraN | traN |
| ICEValE0601_067 | 60956..61288 | + | 333 | ALF34823 | Hypothetical protein | - |
| ICEValE0601_068 | 61319..62008 | + | 690 | ALF34824 | Hypothetical protein | - |
| ICEValE0601_069 | 62324..62752 | + | 429 | ALF34825 | Transposase | - |
| ICEValE0601_070 | 62798..63697 | - | 900 | ALF34868 | Transposase | - |
| ICEValE0601_071 | 63929..64231 | - | 303 | ALF34877 | Hypothetical protein | - |
Host bacterium
| ID | 124 | Element type | ICE (Integrative and conjugative element) |
| Element name | ICEValE0601 | GenBank | KT072768 |
| Element size | 106165 bp | Coordinate of oriT [Strand] | 5548..5846 [+] |
| Host bacterium | Vibrio alginolyticus strain E0601 sequence | Coordinate of element | 1..106165 |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |