Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 121129 |
| Name | oriT_pR1 |
| Organism | Xanthomonas arboricola pv. pruni strain R1 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP091080 (11014..11150 [-], 137 nt) |
| oriT length | 137 nt |
| IRs (inverted repeats) | 28..33, 37..42 (ACATGC..GCATGT) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 137 nt
>oriT_pR1
CGAATGGGCAGGCGGGGATTGGCAGGAACATGCGTAGCATGTGTATGACAATCCGCATCCTGTATTGCTCAACCACTCTACTATCATATCAGCAGTTTAGGTGCACGGGACGCACTTTGCAACAAGCATAAAAGGCG
CGAATGGGCAGGCGGGGATTGGCAGGAACATGCGTAGCATGTGTATGACAATCCGCATCCTGTATTGCTCAACCACTCTACTATCATATCAGCAGTTTAGGTGCACGGGACGCACTTTGCAACAAGCATAAAAGGCG
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file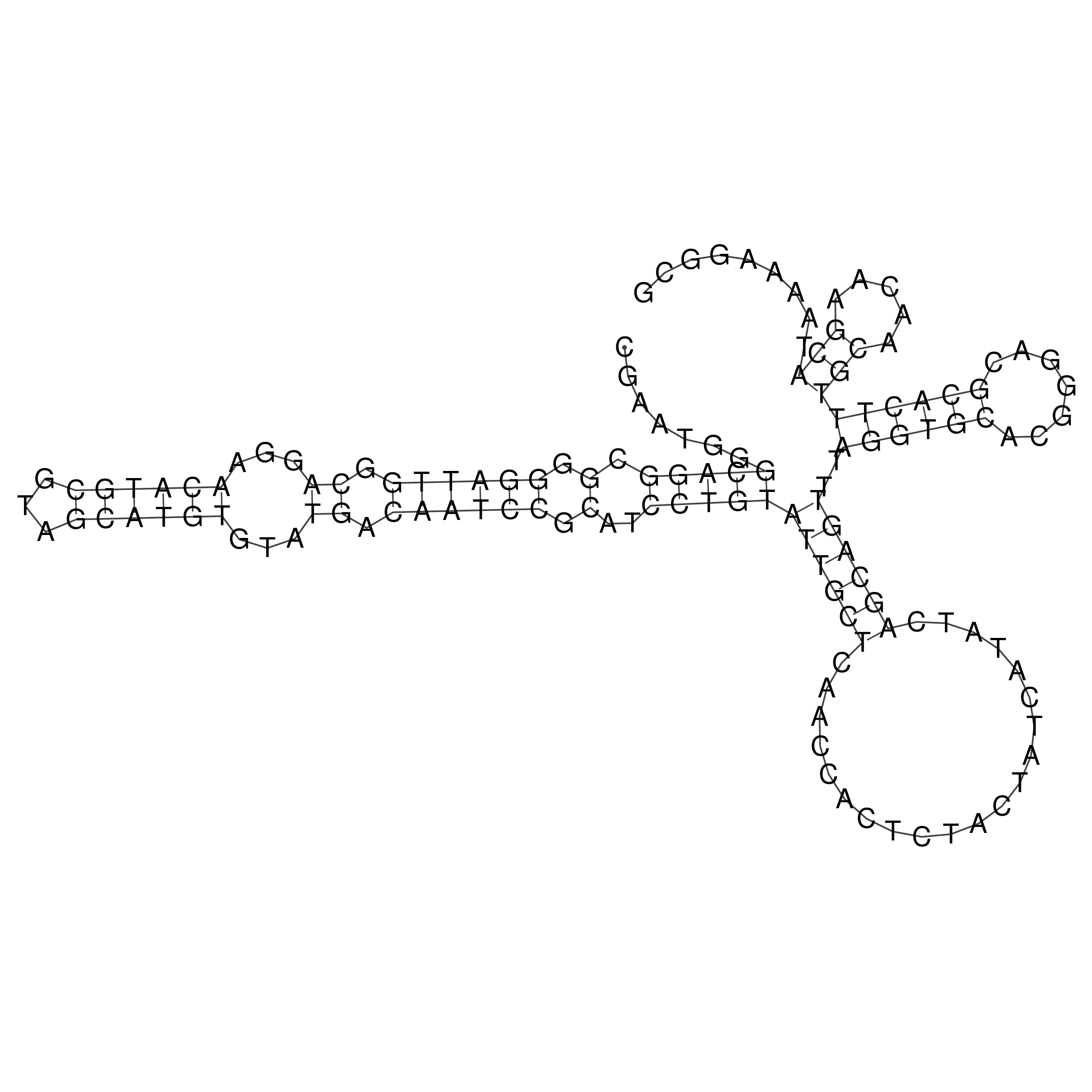
Relaxase
| ID | 13339 | GenBank | WP_047353284 |
| Name | traI_K9U01_RS22240_pR1 |
UniProt ID | A0A2H5CQB3 |
| Length | 890 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 890 a.a. Molecular weight: 101914.42 Da Isoelectric Point: 9.7193
>WP_047353284.1 MULTISPECIES: TraI/MobA(P) family conjugative relaxase [Pseudomonadota]
MIVKKVPNPKKSASKAQRIGQLTSYVRSPESESPQEKCLYAGARGFMMDDPKSQTAEMIALSQEAVRSKD
TINHYVLSWREGEQPSPEQVEEAVSIFMDELGWKDHQAIYGLHSDTDNIHLHIVINRVHPETLKIVEKNR
GFDIELAHKAIARIEHAQGWQREQNGRYQVLENGELGRAPYDPEKPRQPDQKKRDMENRTGEKSAHRIAI
EDGAAIIKQAQTWEQLHRELAAKGMRYEKTGSGATVFVGDVGVKASDVDRNASLAKMQKRLGEYQPAPQR
QQVAPREPEPIKPDVPGWKDYITGRKAHYAEKNADKLAQDKRQEQERKQLAEQQKARRDELMRGNWKGKG
EVLNAMRSVIAAEQAAEKAALKEKHQKEREQHRQRFRPYPDLEQWQRMQKSPELAEQWRHRASEPQRIEG
DRSEPPTPRDIRAYQPEIVGQQVHYSRKEEAGRGGGVSFVDKGKSIDIHDWRNRDSTLAALQLSAQKWGS
FTVMGNDEYKAMCGKLAAEHGFKITNPELQESIQQERQRIQQERVQAMKSEQLKQFERYAEAVGAERYRV
TSIKMREDGSKQTFILDKKDGITRGFTPQEIEQRTPEMQRLQRRGENLYYTPLSDKKHHILIDDMNREKL
ERLIRDGYQPAAVLESSPGNYQAIITVPKLGTAHDKDVGNRLSDALNREYGDPKLSGAIHPHRAPGYENR
KPKHQREDGSYPEVRLLKAERRECIKALALSSQIDAEYQRQAALKAQQPERSKAKPALELAAASGSAIDA
YQRHYRDVLKRQRGGEVDLSRLDSMIAVRMRVTGHDQAAIEGAIRQCAPATRQKDEGRDWNDYAQRTARY
AYSAAGDRQAAELGKYRQQWEKLEGREPVRQQEQAKAQKIERDNSPGMSL
MIVKKVPNPKKSASKAQRIGQLTSYVRSPESESPQEKCLYAGARGFMMDDPKSQTAEMIALSQEAVRSKD
TINHYVLSWREGEQPSPEQVEEAVSIFMDELGWKDHQAIYGLHSDTDNIHLHIVINRVHPETLKIVEKNR
GFDIELAHKAIARIEHAQGWQREQNGRYQVLENGELGRAPYDPEKPRQPDQKKRDMENRTGEKSAHRIAI
EDGAAIIKQAQTWEQLHRELAAKGMRYEKTGSGATVFVGDVGVKASDVDRNASLAKMQKRLGEYQPAPQR
QQVAPREPEPIKPDVPGWKDYITGRKAHYAEKNADKLAQDKRQEQERKQLAEQQKARRDELMRGNWKGKG
EVLNAMRSVIAAEQAAEKAALKEKHQKEREQHRQRFRPYPDLEQWQRMQKSPELAEQWRHRASEPQRIEG
DRSEPPTPRDIRAYQPEIVGQQVHYSRKEEAGRGGGVSFVDKGKSIDIHDWRNRDSTLAALQLSAQKWGS
FTVMGNDEYKAMCGKLAAEHGFKITNPELQESIQQERQRIQQERVQAMKSEQLKQFERYAEAVGAERYRV
TSIKMREDGSKQTFILDKKDGITRGFTPQEIEQRTPEMQRLQRRGENLYYTPLSDKKHHILIDDMNREKL
ERLIRDGYQPAAVLESSPGNYQAIITVPKLGTAHDKDVGNRLSDALNREYGDPKLSGAIHPHRAPGYENR
KPKHQREDGSYPEVRLLKAERRECIKALALSSQIDAEYQRQAALKAQQPERSKAKPALELAAASGSAIDA
YQRHYRDVLKRQRGGEVDLSRLDSMIAVRMRVTGHDQAAIEGAIRQCAPATRQKDEGRDWNDYAQRTARY
AYSAAGDRQAAELGKYRQQWEKLEGREPVRQQEQAKAQKIERDNSPGMSL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A2H5CQB3 |
Host bacterium
| ID | 21557 | GenBank | NZ_CP091080 |
| Plasmid name | pR1 | Incompatibility group | IncQ2 |
| Plasmid size | 20174 bp | Coordinate of oriT [Strand] | 11014..11150 [-] |
| Host baterium | Xanthomonas arboricola pv. pruni strain R1 |
Cargo genes
| Drug resistance gene | tet(C), aph(6)-Id, aph(3'')-Ib |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |