Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 118944 |
| Name | oriT_DY2415|p1 |
| Organism | Raoultella sp. DY2415 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP111116 (54641..54690 [-], 50 nt) |
| oriT length | 50 nt |
| IRs (inverted repeats) | 7..13, 18..24 (GCAAAAT..ATTTTGC) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 50 nt
>oriT_DY2415|p1
AAATCTGCAAAATATTAATTTTGCGTGGGGTGTGGTCATTTTGTGGTGAG
AAATCTGCAAAATATTAATTTTGCGTGGGGTGTGGTCATTTTGTGGTGAG
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file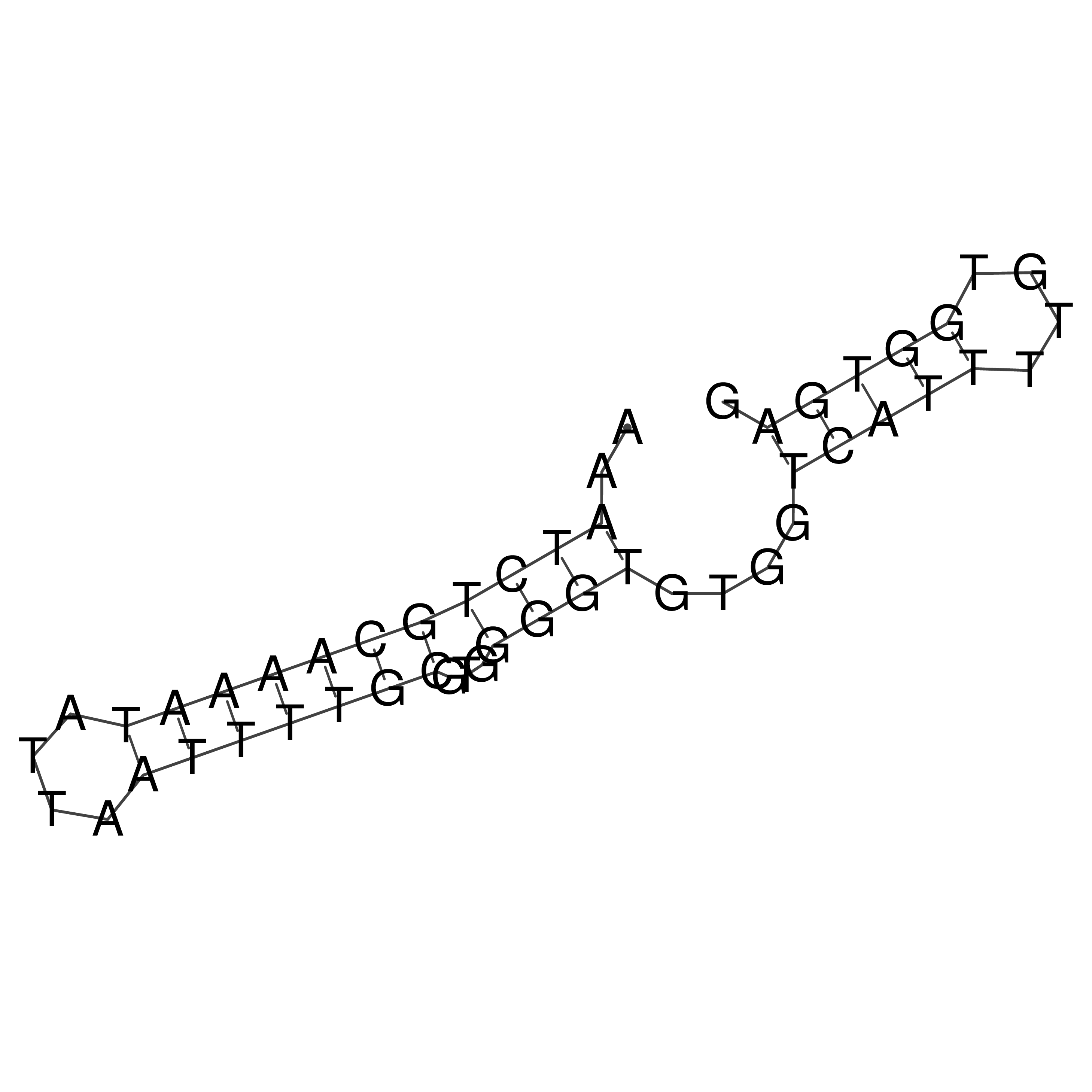
Relaxase
| ID | 12004 | GenBank | WP_110522042 |
| Name | traI_OQV00_RS26750_DY2415|p1 |
UniProt ID | _ |
| Length | 1750 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1750 a.a. Molecular weight: 189911.63 Da Isoelectric Point: 6.3072
>WP_110522042.1 MULTISPECIES: conjugative transfer relaxase/helicase TraI [Klebsiella/Raoultella group]
MMSIGSVKSAGSASNYYTDKDNYYVIGSMDERWQGKGAEALGLDGKVDKQVFTDLLKGKLPDGSDLTRMQ
DGVNKHRPGYDLTFSAPKSVSVMAMIGGDKRLIDAHNQAVTEAVKQVETLAATRVMTDGKSKTVLTGNLI
VAKFNHDTNRNQEPQLHTHAVVINATQNGGKWQSLGTDTVGKTGFIENVYANQIAFGKLYREVLKPLVEK
LGYETEVTGKHGMWEMKDVPVEPFSTRSQEVREAAGPDASLKSRDVATLDTRKSKETLDPAEKMVEWVNT
LKESGFDIHSYREDADKRAAEINRAPVSPAKTDGPDISDVVTKAIAGLSDRKVQFTYADMLARTVSQLEA
KPGVFELARTGIDAAIEREQLIPLDREKGLFTSNIHVLDELSVKALSQEVQRQNHVSVTPDASVVRHAPF
SDAVSVLAQDRPAMGIVYGQGGATGQRERVAELTLMAREQGRDVHILAADNRSRDFLAGDARLVGETVTG
KSALQDGTAFIPGGTLIVDQAEKLSLKETLSLLDGAMRHNVQVLLSDSGKRSGTGSAVTVLKESGVNTYR
WQGGNQTTADIISEPDKNVRYSRLAQEFAVSVREGQESVAQISGTREQNALNNVIRDTLKNEGVLGAKEI
TVTALTPVWLDSKSRGVRDYYREGMVMERWDPETRKHDRFLIDRVTASSNMLTLRDKEGERLDLKVSAVG
SQWTLFRADKLLVADGERLSVLGKIPETRLKGGESVTVLKAEAGQLTVQRPGQKTTQILAVGAGPFDGIK
VGHGWVESPGRSVSETATVFASLTQRELDNATLNQLAQSGSHIRLYSAQDAARTTEKLARHTAFSVVSEQ
LKARAGETELDAAIASQKARLHTPAEQAIHLSIPLLESDNLTFTRPQLLATAMESGGGKVPMADIDRSIQ
AQIKAGSLLNVPVAPGQGNDLLISRQTWDAEKSILTRVLEGKEAVAPLMDSVPGSLLSDLTSGQRASTRM
ILETTDRFTVVQGYAGVGKTTQFRAVTGAINLLPEETRPRVIGLGPTHRAVGEMQSAGVDAQTTASFLHD
TQLLQRNGQTPDFSNTLFLLDESSMVGLTDTAKALSLIAAGGGRAVLSGDTDQLQSIAPGQPFRLMQQRS
AADVAIMKEIVRQVPELRPAVYSLIDRDINTALATIEKVTPEQVPRKDGAWVPENSVVEFTPAQEEAIQK
SLNKGESVPAGQPDTLYGALVKDYTGRTPEAQSQTLIITHLNDDRRALNSLIHNARRENGETGREEVTLP
VLVTSNIRDGELRRLSTWTAHRGAVALVDNVYHRIGSVDKENQLITLTDSEGKERLISPREASAEGVTLY
REEAITVSQGDKMRFSKSDTERGYVANSVWEVKSVSGDSVTLSDGKTTRTMNPKADEAQRHIDLAYAITA
HGAQGASEPYAIALEGVAGGRERMASFESAYVGLSRMKQHVQVYTDSREGWIKAINHSPEKATAHDILAP
RNDRAVKTAELLFGRAKPLDETAAGRAALQQSGLARGSSPGKFISPGKKYPKPHVALPAFDKNGKDAGIW
LSPLTDKDGCLEAIGGEGRVMGNEAARFVGLQNSRNGESLLAGNMSEGVRMARDNPDSGVVVRLAGDDRP
WNPGAITGGRVWADPVPVFPQTGADVPLPPEVLAQRAAEEVQRQQMEKQAEQTAREVAGDGRKNGEPDDR
IKEVIGDVIRGLERDKPGGGKTVLPDDPQTHRQEMAVQQVVSESLQRERLQQMERDMVRDLSREKTLGGD
MMSIGSVKSAGSASNYYTDKDNYYVIGSMDERWQGKGAEALGLDGKVDKQVFTDLLKGKLPDGSDLTRMQ
DGVNKHRPGYDLTFSAPKSVSVMAMIGGDKRLIDAHNQAVTEAVKQVETLAATRVMTDGKSKTVLTGNLI
VAKFNHDTNRNQEPQLHTHAVVINATQNGGKWQSLGTDTVGKTGFIENVYANQIAFGKLYREVLKPLVEK
LGYETEVTGKHGMWEMKDVPVEPFSTRSQEVREAAGPDASLKSRDVATLDTRKSKETLDPAEKMVEWVNT
LKESGFDIHSYREDADKRAAEINRAPVSPAKTDGPDISDVVTKAIAGLSDRKVQFTYADMLARTVSQLEA
KPGVFELARTGIDAAIEREQLIPLDREKGLFTSNIHVLDELSVKALSQEVQRQNHVSVTPDASVVRHAPF
SDAVSVLAQDRPAMGIVYGQGGATGQRERVAELTLMAREQGRDVHILAADNRSRDFLAGDARLVGETVTG
KSALQDGTAFIPGGTLIVDQAEKLSLKETLSLLDGAMRHNVQVLLSDSGKRSGTGSAVTVLKESGVNTYR
WQGGNQTTADIISEPDKNVRYSRLAQEFAVSVREGQESVAQISGTREQNALNNVIRDTLKNEGVLGAKEI
TVTALTPVWLDSKSRGVRDYYREGMVMERWDPETRKHDRFLIDRVTASSNMLTLRDKEGERLDLKVSAVG
SQWTLFRADKLLVADGERLSVLGKIPETRLKGGESVTVLKAEAGQLTVQRPGQKTTQILAVGAGPFDGIK
VGHGWVESPGRSVSETATVFASLTQRELDNATLNQLAQSGSHIRLYSAQDAARTTEKLARHTAFSVVSEQ
LKARAGETELDAAIASQKARLHTPAEQAIHLSIPLLESDNLTFTRPQLLATAMESGGGKVPMADIDRSIQ
AQIKAGSLLNVPVAPGQGNDLLISRQTWDAEKSILTRVLEGKEAVAPLMDSVPGSLLSDLTSGQRASTRM
ILETTDRFTVVQGYAGVGKTTQFRAVTGAINLLPEETRPRVIGLGPTHRAVGEMQSAGVDAQTTASFLHD
TQLLQRNGQTPDFSNTLFLLDESSMVGLTDTAKALSLIAAGGGRAVLSGDTDQLQSIAPGQPFRLMQQRS
AADVAIMKEIVRQVPELRPAVYSLIDRDINTALATIEKVTPEQVPRKDGAWVPENSVVEFTPAQEEAIQK
SLNKGESVPAGQPDTLYGALVKDYTGRTPEAQSQTLIITHLNDDRRALNSLIHNARRENGETGREEVTLP
VLVTSNIRDGELRRLSTWTAHRGAVALVDNVYHRIGSVDKENQLITLTDSEGKERLISPREASAEGVTLY
REEAITVSQGDKMRFSKSDTERGYVANSVWEVKSVSGDSVTLSDGKTTRTMNPKADEAQRHIDLAYAITA
HGAQGASEPYAIALEGVAGGRERMASFESAYVGLSRMKQHVQVYTDSREGWIKAINHSPEKATAHDILAP
RNDRAVKTAELLFGRAKPLDETAAGRAALQQSGLARGSSPGKFISPGKKYPKPHVALPAFDKNGKDAGIW
LSPLTDKDGCLEAIGGEGRVMGNEAARFVGLQNSRNGESLLAGNMSEGVRMARDNPDSGVVVRLAGDDRP
WNPGAITGGRVWADPVPVFPQTGADVPLPPEVLAQRAAEEVQRQQMEKQAEQTAREVAGDGRKNGEPDDR
IKEVIGDVIRGLERDKPGGGKTVLPDDPQTHRQEMAVQQVVSESLQRERLQQMERDMVRDLSREKTLGGD
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4CP
| ID | 14057 | GenBank | WP_266280848 |
| Name | traD_OQV00_RS26745_DY2415|p1 |
UniProt ID | _ |
| Length | 764 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 764 a.a. Molecular weight: 85399.59 Da Isoelectric Point: 4.9674
>WP_266280848.1 type IV conjugative transfer system coupling protein TraD [Raoultella sp. DY2415]
MSFNAKDMTQGGQIANMRFRMFGQIANIIFYVLFILFWVLCGLMLMYRLSWQTFVNGCVYWWCTTLAPML
DIIRSQPVYTISYYGKSLEYNAEQILADKYTIWCGEQLWTAFVFAAIVSLTICVVTFFVASWVLGRQGKL
QSEDENTGGRQLSDKPKDVARQMKRDGAASDIKIGDLPILKNSEIQNFCLHGTVGSGKSEVIRRLLNYVR
ARGDMAIIYDRSCEFVKSYYDPSLDKILNPLDSRCAAWDLWKECLTLPDFDNISNTLIPMGTKEDPFWQG
SGRTIFAEGAYLMRGDKDRSYEKLVDTMLSIKIDKLRAYLQNTPAANLVEEKIEKTAISIRAVLTNYVKA
IRYLQGIEKNGEPFTIRDWMRGVREDQPNGWLFISSNADTHASLKPVISMWLSIAIRGLLAMGENRNRRV
WIFADELPTLHKLPDLVEILPEARKFGGCYVFGIQSYAQLEDIYGVKPAATLFDVMNTRAFFRSPSKEIA
EFAAGEIGEKEILKASEQYSYGADPVRDGVSTGKEKERETLVSYSDIQTLPDLSCYVTLPGPYPAVKLAL
KYNPRPKIAEGFIQRRLDTRIDERLSAQLEAREAEGSLVRALFTPDEPVSEPAEEKAGVQPEQTKPAKDS
VLPVPVKTSSMAAEGAGKTPDVPLSGAPVTRVKTVPLIRPKPAASVTGTAAAASTAAVSATAGGTEQELA
QQPVEADQDMLPAGMNEDGVIEDMQAYDAWLEGEQIQRDMQRREEVNINHSRSEQDDIEIGGNF
MSFNAKDMTQGGQIANMRFRMFGQIANIIFYVLFILFWVLCGLMLMYRLSWQTFVNGCVYWWCTTLAPML
DIIRSQPVYTISYYGKSLEYNAEQILADKYTIWCGEQLWTAFVFAAIVSLTICVVTFFVASWVLGRQGKL
QSEDENTGGRQLSDKPKDVARQMKRDGAASDIKIGDLPILKNSEIQNFCLHGTVGSGKSEVIRRLLNYVR
ARGDMAIIYDRSCEFVKSYYDPSLDKILNPLDSRCAAWDLWKECLTLPDFDNISNTLIPMGTKEDPFWQG
SGRTIFAEGAYLMRGDKDRSYEKLVDTMLSIKIDKLRAYLQNTPAANLVEEKIEKTAISIRAVLTNYVKA
IRYLQGIEKNGEPFTIRDWMRGVREDQPNGWLFISSNADTHASLKPVISMWLSIAIRGLLAMGENRNRRV
WIFADELPTLHKLPDLVEILPEARKFGGCYVFGIQSYAQLEDIYGVKPAATLFDVMNTRAFFRSPSKEIA
EFAAGEIGEKEILKASEQYSYGADPVRDGVSTGKEKERETLVSYSDIQTLPDLSCYVTLPGPYPAVKLAL
KYNPRPKIAEGFIQRRLDTRIDERLSAQLEAREAEGSLVRALFTPDEPVSEPAEEKAGVQPEQTKPAKDS
VLPVPVKTSSMAAEGAGKTPDVPLSGAPVTRVKTVPLIRPKPAASVTGTAAAASTAAVSATAGGTEQELA
QQPVEADQDMLPAGMNEDGVIEDMQAYDAWLEGEQIQRDMQRREEVNINHSRSEQDDIEIGGNF
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 54082..82011
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| OQV00_RS26545 | 50084..50515 | + | 432 | WP_154949013 | conjugation system SOS inhibitor PsiB | - |
| OQV00_RS26550 | 50512..51243 | + | 732 | WP_266280977 | plasmid SOS inhibition protein A | - |
| OQV00_RS26555 | 51240..51467 | + | 228 | WP_154949017 | hypothetical protein | - |
| OQV00_RS26560 | 51467..51706 | + | 240 | WP_154949019 | hypothetical protein | - |
| OQV00_RS26565 | 51882..52034 | + | 153 | WP_224222653 | DUF5431 family protein | - |
| OQV00_RS26570 | 51979..52098 | + | 120 | WP_223156217 | type I toxin-antitoxin system Hok family toxin | - |
| OQV00_RS26575 | 52866..53687 | + | 822 | WP_266280980 | DUF932 domain-containing protein | - |
| OQV00_RS26580 | 53720..54049 | + | 330 | WP_266280983 | DUF5983 family protein | - |
| OQV00_RS26585 | 54082..54567 | - | 486 | WP_266281003 | transglycosylase SLT domain-containing protein | virB1 |
| OQV00_RS26590 | 54998..55390 | + | 393 | WP_032736874 | conjugal transfer relaxosome DNA-binding protein TraM | - |
| OQV00_RS26595 | 55628..56323 | + | 696 | WP_154925432 | sigma factor-like helix-turn-helix DNA-binding protein | - |
| OQV00_RS26600 | 56414..56611 | + | 198 | WP_154925430 | TraY domain-containing protein | - |
| OQV00_RS26605 | 56676..57041 | + | 366 | WP_154925428 | type IV conjugative transfer system pilin TraA | - |
| OQV00_RS26610 | 57055..57360 | + | 306 | WP_154925426 | type IV conjugative transfer system protein TraL | traL |
| OQV00_RS26615 | 57380..57946 | + | 567 | WP_154925424 | type IV conjugative transfer system protein TraE | traE |
| OQV00_RS26620 | 57933..58667 | + | 735 | WP_154925422 | type-F conjugative transfer system secretin TraK | traK |
| OQV00_RS26625 | 58667..60088 | + | 1422 | WP_266280988 | F-type conjugal transfer pilus assembly protein TraB | traB |
| OQV00_RS26630 | 60081..60464 | + | 384 | WP_154925419 | conjugal transfer protein TraP | - |
| OQV00_RS26635 | 60445..60669 | + | 225 | WP_154925417 | hypothetical protein | - |
| OQV00_RS26640 | 60692..61261 | + | 570 | WP_154925415 | type IV conjugative transfer system lipoprotein TraV | traV |
| OQV00_RS26645 | 61375..61707 | + | 333 | WP_154925413 | hypothetical protein | - |
| OQV00_RS27105 | 61707..62112 | + | 406 | Protein_66 | hypothetical protein | - |
| OQV00_RS26650 | 62109..62459 | + | 351 | WP_154925411 | hypothetical protein | - |
| OQV00_RS26655 | 62607..65246 | + | 2640 | WP_154925409 | type IV secretion system protein TraC | virb4 |
| OQV00_RS26660 | 65246..65638 | + | 393 | WP_154925407 | type-F conjugative transfer system protein TrbI | - |
| OQV00_RS26665 | 65635..66285 | + | 651 | WP_154925405 | type-F conjugative transfer system protein TraW | traW |
| OQV00_RS26670 | 66307..67299 | + | 993 | WP_154925403 | conjugal transfer pilus assembly protein TraU | traU |
| OQV00_RS26675 | 67312..67938 | + | 627 | WP_154925401 | type-F conjugative transfer system pilin assembly protein TrbC | trbC |
| OQV00_RS26680 | 67935..69917 | + | 1983 | WP_154949027 | type-F conjugative transfer system mating-pair stabilization protein TraN | traN |
| OQV00_RS26685 | 69910..70521 | + | 612 | WP_154925397 | hypothetical protein | - |
| OQV00_RS26690 | 70511..70747 | + | 237 | WP_154925395 | conjugal transfer protein TrbE | - |
| OQV00_RS26695 | 70966..71292 | + | 327 | WP_154925393 | hypothetical protein | - |
| OQV00_RS26700 | 71315..72067 | + | 753 | WP_154925391 | type-F conjugative transfer system pilin assembly protein TraF | traF |
| OQV00_RS26705 | 72078..72518 | + | 441 | WP_154925389 | hypothetical protein | - |
| OQV00_RS26710 | 72481..72840 | - | 360 | WP_177951992 | hypothetical protein | - |
| OQV00_RS26715 | 72929..73165 | + | 237 | WP_154925387 | type-F conjugative transfer system pilin chaperone TraQ | - |
| OQV00_RS26720 | 73140..73715 | + | 576 | WP_250210061 | type-F conjugative transfer system pilin assembly thiol-disulfide isomerase TrbB | traF |
| OQV00_RS26725 | 73702..75072 | + | 1371 | WP_154925382 | conjugal transfer pilus assembly protein TraH | traH |
| OQV00_RS26730 | 75072..77966 | + | 2895 | WP_154925380 | conjugal transfer mating-pair stabilization protein TraG | traG |
| OQV00_RS26735 | 77963..78478 | + | 516 | WP_154925378 | conjugal transfer protein TraS | - |
| OQV00_RS26740 | 78790..79521 | + | 732 | WP_154925376 | conjugal transfer complement resistance protein TraT | - |
| OQV00_RS26745 | 79717..82011 | + | 2295 | WP_266280848 | type IV conjugative transfer system coupling protein TraD | virb4 |
Host bacterium
| ID | 19376 | GenBank | NZ_CP111116 |
| Plasmid name | DY2415|p1 | Incompatibility group | IncFIB |
| Plasmid size | 152330 bp | Coordinate of oriT [Strand] | 54641..54690 [-] |
| Host baterium | Raoultella sp. DY2415 |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |