Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 118094 |
| Name | oriT_PCH555|unnamed1 |
| Organism | Enterococcus faecalis strain PCH555 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP090559 (4138..4187 [-], 50 nt) |
| oriT length | 50 nt |
| IRs (inverted repeats) | _ |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 50 nt
>oriT_PCH555|unnamed1
AATATCGCAACATGCTAGCATGTTGCTCTGCTCGCAAAAAGAAAGCCTAC
AATATCGCAACATGCTAGCATGTTGCTCTGCTCGCAAAAAGAAAGCCTAC
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file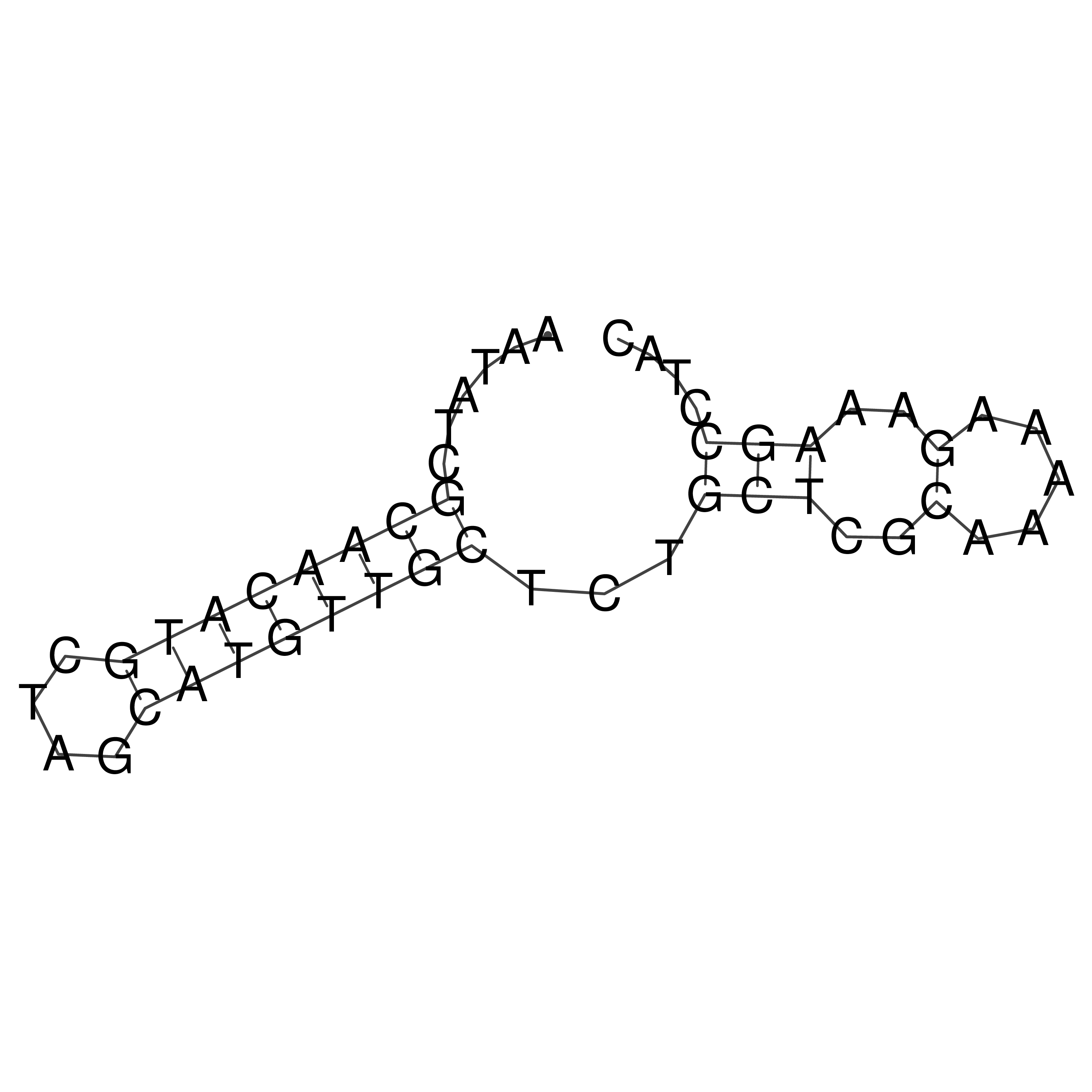
Relaxase
| ID | 11477 | GenBank | WP_010784125 |
| Name | Relaxase_LZP99_RS13460_PCH555|unnamed1 |
UniProt ID | _ |
| Length | 561 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 561 a.a. Molecular weight: 65427.18 Da Isoelectric Point: 6.1241
>WP_010784125.1 relaxase/mobilization nuclease domain-containing protein [Enterococcus faecalis]
MVYTKHFVIHTFDKLNNACSYIENAEKTEVTNDNPSEHLEHLFQYIVNDDKTYMKKLVSGHGIVDPTNPY
EEFKLTKLQAAIQRKIGYTFDPKSERLLPPTLTELEKGNAVLAHHLIQSFSPEDDLTPEKTHEIGYNTVM
ELTGGEYEFVIATHVDKEHLHNHIIFSSTNLKTGKAFRWQKGTKRVFEQISDKIAAKEGAKIIEKSPKNS
HKKYTMWETESLYKEKIKSRLDFLLEQSSSINDFLEKAEALNLSVDFSKKWTTYRLLDEPQIKNTRSRSL
SKNDPTRYNYEKIVERLKENKSVLTLKEVVEKYVEKGEKSKNEFDYQLTIDEWQISHKTTRGYYVNVDFG
FGERGKVFIGAYKVEPLENGQYNIYVKRKDFFYLMNEKNADRNRYMTGETLIKQLRLYNGQTPLKKEPVM
RTIDELVNAINFLAANEIEDTRQLELLEEKLEAAFLEAEETLETLDEKMLELHQLSNLLLENELNGELEL
IQKKLKTLLPDATLAEFSYEDVRGEIEAIKTSQSLLENKLERTRNEINQLHEIQAVQKKETANQNNVKPK
L
MVYTKHFVIHTFDKLNNACSYIENAEKTEVTNDNPSEHLEHLFQYIVNDDKTYMKKLVSGHGIVDPTNPY
EEFKLTKLQAAIQRKIGYTFDPKSERLLPPTLTELEKGNAVLAHHLIQSFSPEDDLTPEKTHEIGYNTVM
ELTGGEYEFVIATHVDKEHLHNHIIFSSTNLKTGKAFRWQKGTKRVFEQISDKIAAKEGAKIIEKSPKNS
HKKYTMWETESLYKEKIKSRLDFLLEQSSSINDFLEKAEALNLSVDFSKKWTTYRLLDEPQIKNTRSRSL
SKNDPTRYNYEKIVERLKENKSVLTLKEVVEKYVEKGEKSKNEFDYQLTIDEWQISHKTTRGYYVNVDFG
FGERGKVFIGAYKVEPLENGQYNIYVKRKDFFYLMNEKNADRNRYMTGETLIKQLRLYNGQTPLKKEPVM
RTIDELVNAINFLAANEIEDTRQLELLEEKLEAAFLEAEETLETLDEKMLELHQLSNLLLENELNGELEL
IQKKLKTLLPDATLAEFSYEDVRGEIEAIKTSQSLLENKLERTRNEINQLHEIQAVQKKETANQNNVKPK
L
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Auxiliary protein
| ID | 6595 | GenBank | WP_010784124 |
| Name | WP_010784124_PCH555|unnamed1 |
UniProt ID | _ |
| Length | 118 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
Auxiliary protein sequence
Download Length: 118 a.a. Molecular weight: 13987.11 Da Isoelectric Point: 10.1993
>WP_010784124.1 MULTISPECIES: plasmid mobilization relaxosome protein MobC [Enterococcus]
MENKQKLKRPIQRIVRLSEEENNLIKRKIEKSFFPNFQNFALHLLIQGEIRHVDYSELNRLTTEIHKIGI
NINQMARLANQFHEISSEDIKDLTDKVHSLNALVQSELNKLIKRKEQS
MENKQKLKRPIQRIVRLSEEENNLIKRKIEKSFFPNFQNFALHLLIQGEIRHVDYSELNRLTTEIHKIGI
NINQMARLANQFHEISSEDIKDLTDKVHSLNALVQSELNKLIKRKEQS
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4CP
| ID | 13413 | GenBank | WP_048948780 |
| Name | t4cp2_LZP99_RS13480_PCH555|unnamed1 |
UniProt ID | _ |
| Length | 609 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 609 a.a. Molecular weight: 70085.17 Da Isoelectric Point: 6.4057
>WP_048948780.1 MULTISPECIES: VirD4-like conjugal transfer protein, CD1115 family [Enterococcus]
MQTYKKELKPYLYLGIFLFVIGFGIGNFFWLLPGTDYFSKTNYLFTKDILTYIEQTFYRFFITPSLLGFS
VGILGFLLGLLMYVRDNDRGIYRHGEEYGSARFATPAEMKKYEDPIPENNIIVSKHVKISLFNKRLPIKL
QKNKNIAILGDSGAAKTLAFIKTNLMQRHASFITTDPDGGILPEIGLLLKKGKYKIKVLDLNTLSNSDTF
NVFEYMHSELDVDRVLEAVTEATKEVEKTTGDDFWIKAEGLLVRSLIAYLWFDGRRNDYTPHLGMIADML
RHLKRKDKNVPSPVEEWFEELNVAIPDNYAYRQWTLFNNLYESETRASVLGIAAARYSVFDHEEVVNMIR
TDTMDIDSWNEVKTAVFIAIPETNTSFNFIAALMFATVGEVLRNKSDQVRLGKRALEEGKELLHVRFLID
EFANIGRIPHFEKMLTTFRKREMSFTIVLQSLNQLQTLYRNGWQNILNGCACLLYLGGDEEETTKYLSAR
AGKQTLSIRKHSMNKGNQGGGSENRDKYGRDLIDRSEVALIGGDECLVFISKEHVFKDKKFYAFQHDRAD
ELATGPTDDKWYTWRRYMTDEEAILDKISATDIIDHGEITSELKEETHV
MQTYKKELKPYLYLGIFLFVIGFGIGNFFWLLPGTDYFSKTNYLFTKDILTYIEQTFYRFFITPSLLGFS
VGILGFLLGLLMYVRDNDRGIYRHGEEYGSARFATPAEMKKYEDPIPENNIIVSKHVKISLFNKRLPIKL
QKNKNIAILGDSGAAKTLAFIKTNLMQRHASFITTDPDGGILPEIGLLLKKGKYKIKVLDLNTLSNSDTF
NVFEYMHSELDVDRVLEAVTEATKEVEKTTGDDFWIKAEGLLVRSLIAYLWFDGRRNDYTPHLGMIADML
RHLKRKDKNVPSPVEEWFEELNVAIPDNYAYRQWTLFNNLYESETRASVLGIAAARYSVFDHEEVVNMIR
TDTMDIDSWNEVKTAVFIAIPETNTSFNFIAALMFATVGEVLRNKSDQVRLGKRALEEGKELLHVRFLID
EFANIGRIPHFEKMLTTFRKREMSFTIVLQSLNQLQTLYRNGWQNILNGCACLLYLGGDEEETTKYLSAR
AGKQTLSIRKHSMNKGNQGGGSENRDKYGRDLIDRSEVALIGGDECLVFISKEHVFKDKKFYAFQHDRAD
ELATGPTDDKWYTWRRYMTDEEAILDKISATDIIDHGEITSELKEETHV
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Host bacterium
| ID | 18527 | GenBank | NZ_CP090559 |
| Plasmid name | PCH555|unnamed1 | Incompatibility group | - |
| Plasmid size | 77536 bp | Coordinate of oriT [Strand] | 4138..4187 [-] |
| Host baterium | Enterococcus faecalis strain PCH555 |
Cargo genes
| Drug resistance gene | ClpL |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | AcrIIA21, AcrIIA9 |