Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 118047 |
| Name | oriT_pC212158-KPC_75k |
| Organism | Klebsiella aerogenes strain C212158 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP139398 (52177..52282 [+], 106 nt) |
| oriT length | 106 nt |
| IRs (inverted repeats) | 30..35, 41..46 (CCGCCG..CGGCGG) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 106 nt
>oriT_pC212158-KPC_75k
AGTACGGGACAAGATGTGTTTTTGGAGTACCGCCGACACGCGGCGGCCGTCCAAAAACGTCTTGGTCAGGGCAAGCCCCGACACCCCCTAACGAGGTTAGCTATCT
AGTACGGGACAAGATGTGTTTTTGGAGTACCGCCGACACGCGGCGGCCGTCCAAAAACGTCTTGGTCAGGGCAAGCCCCGACACCCCCTAACGAGGTTAGCTATCT
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file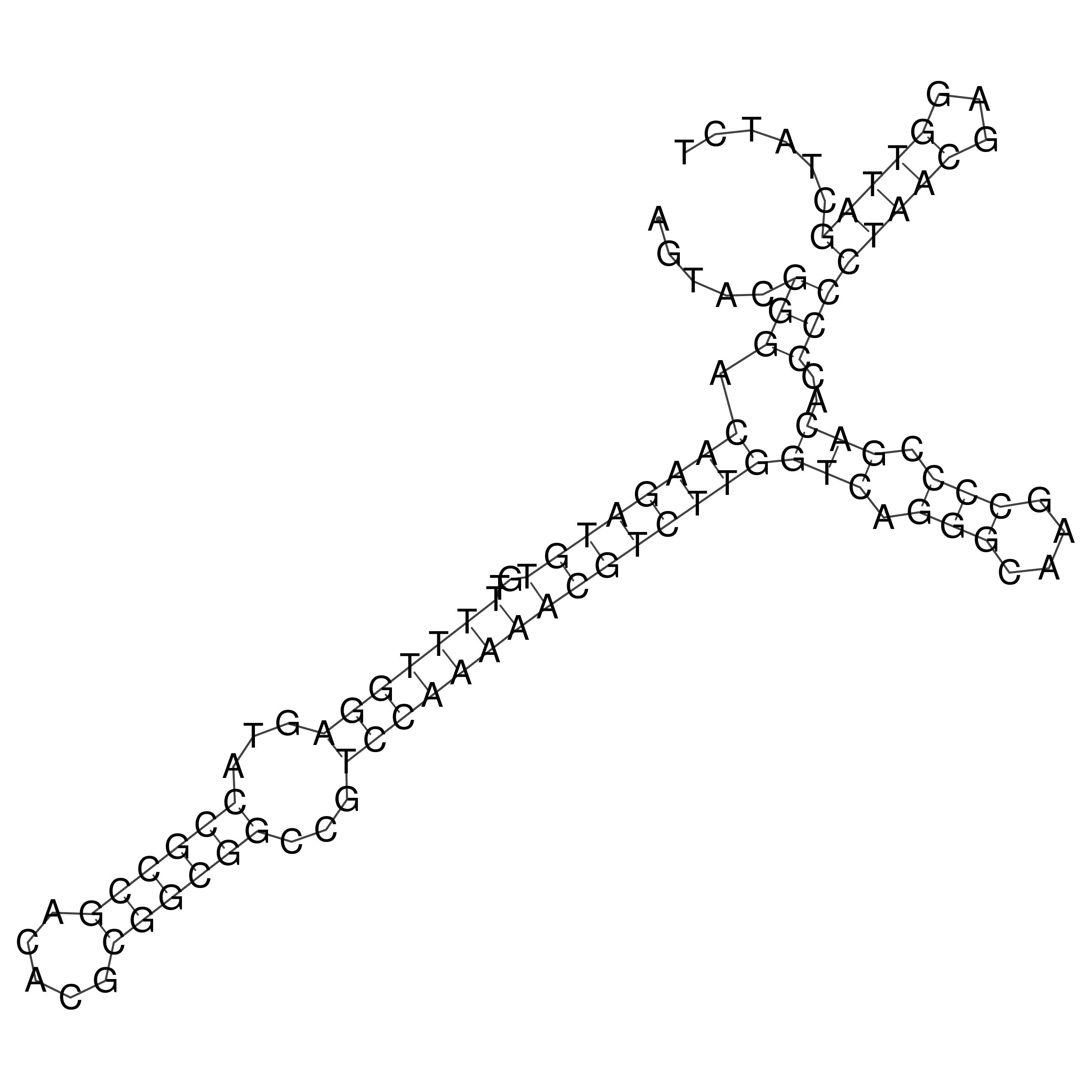
Relaxase
| ID | 11449 | GenBank | WP_004187323 |
| Name | traI_SM904_RS26170_pC212158-KPC_75k |
UniProt ID | A0A3U8KUL3 |
| Length | 659 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 659 a.a. Molecular weight: 75179.89 Da Isoelectric Point: 9.9948
>WP_004187323.1 MULTISPECIES: TraI/MobA(P) family conjugative relaxase [Enterobacterales]
MVIPKIIEGRRDKKSSFGQLIKYMADKPSQELTDTVQPTPETALAVKSDMFEGLNNYLTRKQKVISTPVD
VEPGVQRVVVGDVTCQYNTFSLDGAAQEMNSVSQQSTRCKDPVMHYVLSWPDYEKPNDDQVFDSVKFTLA
SMGMSDHQYVAAIHRDTDNLHVHVAVNRINPQTYKAASSSFTKDTLHQACRLLELKNGWSHSNGAYVVND
RQQIVRNPHSKKERGNWRSLDRINKMENKEGVETLYRYIVGDEQVGGSRQNLIHVSAGLREAKSWDDVHK
TFADIGLRVEKAQGKKGYVITHEHQNQKTAVKASLVFNKAQYTLKSMEERFGEYQPSHIEPAKVSVFKTA
YTPGAYRRDANKRLQRKIERAEERMLLKGRYRAYRNNLPIYSPDKDRIADEYRKIAQHTRLVKNNVRHSV
SDPHTRKLMYNLAEFKRLQAVANLRLSLREERNGFRAANPRLSYREWVEQEALKGDKAALSQMRGFAYSS
RKKEKYKQQLVEQIGFNRTFNAITSHDRDDVAVMASARHGVKPRLLKDGTVIFERDGKPVAADRGHIVLT
ESNGIDKEKTADLAIALTIAGKAKSVRVDGDGEFKELCCNRIVDAAVNHNHPVAQGITFTDAAQQAYAQN
EKHRLIREQNNSKNEMQFRSESDDKFNPK
MVIPKIIEGRRDKKSSFGQLIKYMADKPSQELTDTVQPTPETALAVKSDMFEGLNNYLTRKQKVISTPVD
VEPGVQRVVVGDVTCQYNTFSLDGAAQEMNSVSQQSTRCKDPVMHYVLSWPDYEKPNDDQVFDSVKFTLA
SMGMSDHQYVAAIHRDTDNLHVHVAVNRINPQTYKAASSSFTKDTLHQACRLLELKNGWSHSNGAYVVND
RQQIVRNPHSKKERGNWRSLDRINKMENKEGVETLYRYIVGDEQVGGSRQNLIHVSAGLREAKSWDDVHK
TFADIGLRVEKAQGKKGYVITHEHQNQKTAVKASLVFNKAQYTLKSMEERFGEYQPSHIEPAKVSVFKTA
YTPGAYRRDANKRLQRKIERAEERMLLKGRYRAYRNNLPIYSPDKDRIADEYRKIAQHTRLVKNNVRHSV
SDPHTRKLMYNLAEFKRLQAVANLRLSLREERNGFRAANPRLSYREWVEQEALKGDKAALSQMRGFAYSS
RKKEKYKQQLVEQIGFNRTFNAITSHDRDDVAVMASARHGVKPRLLKDGTVIFERDGKPVAADRGHIVLT
ESNGIDKEKTADLAIALTIAGKAKSVRVDGDGEFKELCCNRIVDAAVNHNHPVAQGITFTDAAQQAYAQN
EKHRLIREQNNSKNEMQFRSESDDKFNPK
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A3U8KUL3 |
T4CP
| ID | 13378 | GenBank | WP_011091092 |
| Name | t4cp2_SM904_RS25885_pC212158-KPC_75k |
UniProt ID | _ |
| Length | 695 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 695 a.a. Molecular weight: 79491.81 Da Isoelectric Point: 4.8270
>WP_011091092.1 MULTISPECIES: hypothetical protein [Gammaproteobacteria]
MQQESVDSKKVIRPDGSFMEWFLTPNVQFGLLAILTVLGLFLPFTMLLSILILPPLMVAFTDRKFRPPLR
MPKDCEMFDDTLTTETQAEYRLGPIRIPHKVRERKKAKGILYVGYERGRLFGRELWLNMTDLLRHMVFFG
TTGSGKTETFYGFIVNFLLWCRGYCLSDGKADNKLAFATWSLARRFGREDDYYVLNLLTGSIDRFVNLVK
QESIPAQSNSVNLFSVAPPTFIIQLMESMLPQVGGDSAQWQDTAKAMMSALINALCYKRARGELLLSQRT
IQKNMSLPAMAALYVEAKKNGWHSEGYAALESYLENTPGFLLANAEYPETWEARAFEQHNYMSRQFLKTL
SLFNETYGHVFPEDSGDIRMDDIFHNDRILIVMIPSLELSRGEAATLGRLYVTLQRMTISKDLGYQLEGK
KEEVLLTHALNNQAPYGLIYDELGQYFTSGMDTLSAQMRSLEKMGVFSSQDHPSLARGANGEVDSLIANT
RVKYFESIEDRKTFEILRETVGQDYYSELSGQEAEHGTFSKTVREGDLYQIREKDRVNIRELRKQTEGQG
VISFQDALVRSAAFYIPDNEKFSSELPMRINRFIEVLPPSPATLYSIYPERKDDELAETNPDAMHFPPIK
LFGYDKAANSLLEKIEVFYELQCGAISPEEMSVCLFELFADWLDGTDTDDVLYHQEDDLFPMIEM
MQQESVDSKKVIRPDGSFMEWFLTPNVQFGLLAILTVLGLFLPFTMLLSILILPPLMVAFTDRKFRPPLR
MPKDCEMFDDTLTTETQAEYRLGPIRIPHKVRERKKAKGILYVGYERGRLFGRELWLNMTDLLRHMVFFG
TTGSGKTETFYGFIVNFLLWCRGYCLSDGKADNKLAFATWSLARRFGREDDYYVLNLLTGSIDRFVNLVK
QESIPAQSNSVNLFSVAPPTFIIQLMESMLPQVGGDSAQWQDTAKAMMSALINALCYKRARGELLLSQRT
IQKNMSLPAMAALYVEAKKNGWHSEGYAALESYLENTPGFLLANAEYPETWEARAFEQHNYMSRQFLKTL
SLFNETYGHVFPEDSGDIRMDDIFHNDRILIVMIPSLELSRGEAATLGRLYVTLQRMTISKDLGYQLEGK
KEEVLLTHALNNQAPYGLIYDELGQYFTSGMDTLSAQMRSLEKMGVFSSQDHPSLARGANGEVDSLIANT
RVKYFESIEDRKTFEILRETVGQDYYSELSGQEAEHGTFSKTVREGDLYQIREKDRVNIRELRKQTEGQG
VISFQDALVRSAAFYIPDNEKFSSELPMRINRFIEVLPPSPATLYSIYPERKDDELAETNPDAMHFPPIK
LFGYDKAANSLLEKIEVFYELQCGAISPEEMSVCLFELFADWLDGTDTDDVLYHQEDDLFPMIEM
Protein domains
No domain identified.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 55078..74367
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| SM904_RS26150 (SM904_26150) | 50162..50473 | + | 312 | WP_011091065 | hypothetical protein | - |
| SM904_RS26155 (SM904_26155) | 50606..51142 | + | 537 | WP_011091066 | hypothetical protein | - |
| SM904_RS26160 (SM904_26160) | 51643..52008 | - | 366 | WP_126336485 | MobC | - |
| SM904_RS26165 (SM904_26165) | 52023..52601 | + | 579 | WP_223811519 | plasmid mobilization protein MobA | - |
| SM904_RS26170 (SM904_26170) | 52588..54567 | + | 1980 | WP_004187323 | TraI/MobA(P) family conjugative relaxase | - |
| SM904_RS26175 (SM904_26175) | 54581..55081 | + | 501 | WP_004187320 | DotD/TraH family lipoprotein | - |
| SM904_RS26180 (SM904_26180) | 55078..55857 | + | 780 | WP_004187315 | type IV secretory system conjugative DNA transfer family protein | traI |
| SM904_RS26185 (SM904_26185) | 55868..57031 | + | 1164 | WP_004187313 | plasmid transfer ATPase TraJ | virB11 |
| SM904_RS26190 (SM904_26190) | 57021..57281 | + | 261 | WP_004187310 | IcmT/TraK family protein | traK |
| SM904_RS26195 (SM904_26195) | 57306..60743 | + | 3438 | WP_225373171 | LPD7 domain-containing protein | - |
| SM904_RS26200 (SM904_26200) | 60709..61221 | + | 513 | WP_011091071 | hypothetical protein | traL |
| SM904_RS26205 (SM904_26205) | 61222..61434 | - | 213 | WP_305953721 | Hha/YmoA family nucleoid-associated regulatory protein | - |
| SM904_RS26210 (SM904_26210) | 61406..61816 | - | 411 | WP_004187465 | H-NS family nucleoid-associated regulatory protein | - |
| SM904_RS26215 (SM904_26215) | 61884..62597 | + | 714 | WP_004187467 | DotI/IcmL family type IV secretion protein | traM |
| SM904_RS26220 (SM904_26220) | 62606..63757 | + | 1152 | WP_004187471 | DotH/IcmK family type IV secretion protein | traN |
| SM904_RS26225 (SM904_26225) | 63769..65118 | + | 1350 | WP_004187474 | conjugal transfer protein TraO | traO |
| SM904_RS26230 (SM904_26230) | 65130..65834 | + | 705 | WP_015060002 | conjugal transfer protein TraP | traP |
| SM904_RS26235 (SM904_26235) | 65858..66388 | + | 531 | WP_004187478 | conjugal transfer protein TraQ | traQ |
| SM904_RS26240 (SM904_26240) | 66405..66794 | + | 390 | WP_004187479 | DUF6750 family protein | traR |
| SM904_RS26245 (SM904_26245) | 66840..67334 | + | 495 | WP_004187480 | hypothetical protein | - |
| SM904_RS26250 (SM904_26250) | 67331..70381 | + | 3051 | WP_004187482 | hypothetical protein | traU |
| SM904_RS26255 (SM904_26255) | 70378..71586 | + | 1209 | WP_011091082 | conjugal transfer protein TraW | traW |
| SM904_RS26260 (SM904_26260) | 71583..72179 | + | 597 | WP_138978268 | TraX-like protein | - |
| SM904_RS26265 (SM904_26265) | 72172..74367 | + | 2196 | WP_015062834 | DotA/TraY family protein | traY |
| SM904_RS26270 (SM904_26270) | 74369..75022 | + | 654 | WP_044706695 | hypothetical protein | - |
| SM904_RS26275 (SM904_26275) | 75103..75333 | + | 231 | WP_011091085 | IncL/M type plasmid replication protein RepC | - |
Host bacterium
| ID | 18480 | GenBank | NZ_CP139398 |
| Plasmid name | pC212158-KPC_75k | Incompatibility group | IncL/M |
| Plasmid size | 75628 bp | Coordinate of oriT [Strand] | 52177..52282 [+] |
| Host baterium | Klebsiella aerogenes strain C212158 |
Cargo genes
| Drug resistance gene | blaTEM-1B, blaKPC-2, blaCTX-M-3 |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |