Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 116688 |
| Name | oriT_pUC109C |
| Organism | Lactococcus cremoris strain UC109 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP034576 (26738..26874 [+], 137 nt) |
| oriT length | 137 nt |
| IRs (inverted repeats) | 21..28, 42..49 (AAAAGGGG..CCCCTTTT) 25..30, 39..44 (GGGGAA..TTCCCC) |
| Location of nic site | 104..105 |
| Conserved sequence flanking the nic site |
GCTTGCAGTA |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 137 nt
>oriT_pUC109C
CGACACAATCCAAAGGGGATAAAAGGGGAAAGTGAAACTTCCCCCTTTTCAAGCCACATTGTAATACAAGAACGAAGTGCTTTGTATTACAATGTGAAAGCTTGCAGTATTTATGGGTTTATATTTTCTATTTTGTT
CGACACAATCCAAAGGGGATAAAAGGGGAAAGTGAAACTTCCCCCTTTTCAAGCCACATTGTAATACAAGAACGAAGTGCTTTGTATTACAATGTGAAAGCTTGCAGTATTTATGGGTTTATATTTTCTATTTTGTT
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file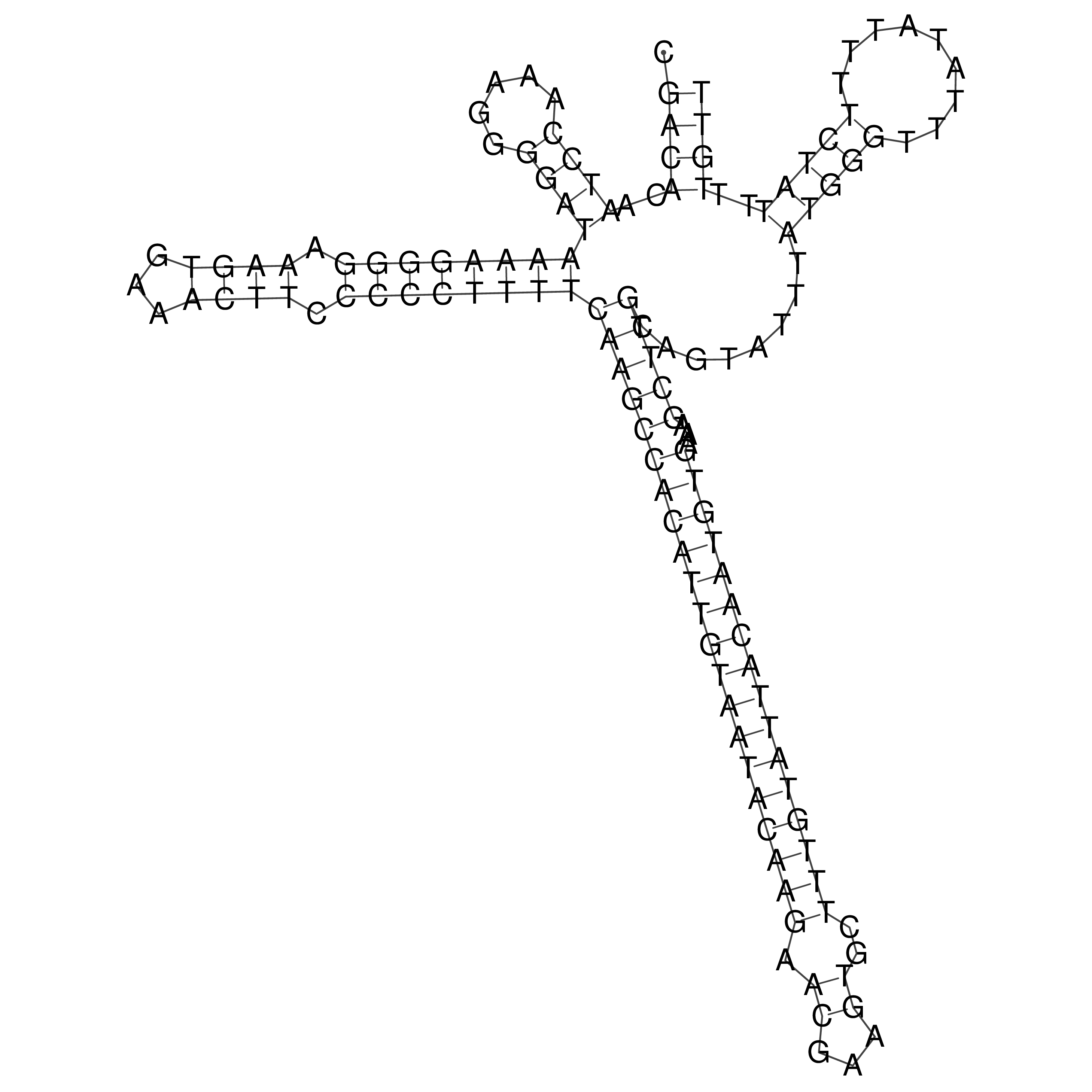
T4CP
| ID | 12437 | GenBank | WP_021214672 |
| Name | tcpA_LLUC109_RS12330_pUC109C |
UniProt ID | _ |
| Length | 358 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 358 a.a. Molecular weight: 40830.21 Da Isoelectric Point: 7.4643
>WP_021214672.1 MULTISPECIES: cell division protein FtsK [Lactococcus]
MTNSQLALYLLQSLNMALGSQIEGETSYTNSFDVKVQEDGFLFLPRMPSGYIIDNDLYFKIFLIANACLY
PRYTLLKQNSAYFIPINTDDIHTQRGLFFPWKVGISKRLVINDLDFFVATQHKPYIPIMENLKLNYDNVT
SIAIAGNSGSGKSYTLTYFLSMLKPLSDLIIIDPKFDTPSRWARENKIPVIHPERNRSKSDFVAEINESL
SQTLNIIYKRQEILYDNPRHQFSHLTIVIDEVLALSEGTNKNIKDSFFSLISQIALLGRATKVHLLLVSQ
RFDYQSIPVSVREQLNVLIQIGNINKKTVQFLFPDLDPEGIVIPLGKGTGLIQIIDNEHPYQVLPLLCPT
YYTKKGIL
MTNSQLALYLLQSLNMALGSQIEGETSYTNSFDVKVQEDGFLFLPRMPSGYIIDNDLYFKIFLIANACLY
PRYTLLKQNSAYFIPINTDDIHTQRGLFFPWKVGISKRLVINDLDFFVATQHKPYIPIMENLKLNYDNVT
SIAIAGNSGSGKSYTLTYFLSMLKPLSDLIIIDPKFDTPSRWARENKIPVIHPERNRSKSDFVAEINESL
SQTLNIIYKRQEILYDNPRHQFSHLTIVIDEVLALSEGTNKNIKDSFFSLISQIALLGRATKVHLLLVSQ
RFDYQSIPVSVREQLNVLIQIGNINKKTVQFLFPDLDPEGIVIPLGKGTGLIQIIDNEHPYQVLPLLCPT
YYTKKGIL
Protein domains
No domain identified.
Protein structure
No available structure.
Host bacterium
| ID | 17121 | GenBank | NZ_CP034576 |
| Plasmid name | pUC109C | Incompatibility group | - |
| Plasmid size | 27874 bp | Coordinate of oriT [Strand] | 26738..26874 [+] |
| Host baterium | Lactococcus cremoris strain UC109 |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |