Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 116199 |
| Name | oriT_SRCM100995|unnamed4 |
| Organism | Lactiplantibacillus plantarum strain SRCM100995 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP028279 (35338..35358 [+], 21 nt) |
| oriT length | 21 nt |
| IRs (inverted repeats) | _ |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 21 nt
>oriT_SRCM100995|unnamed4
AGTGGGGTGTAAGTGCGCATT
AGTGGGGTGTAAGTGCGCATT
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file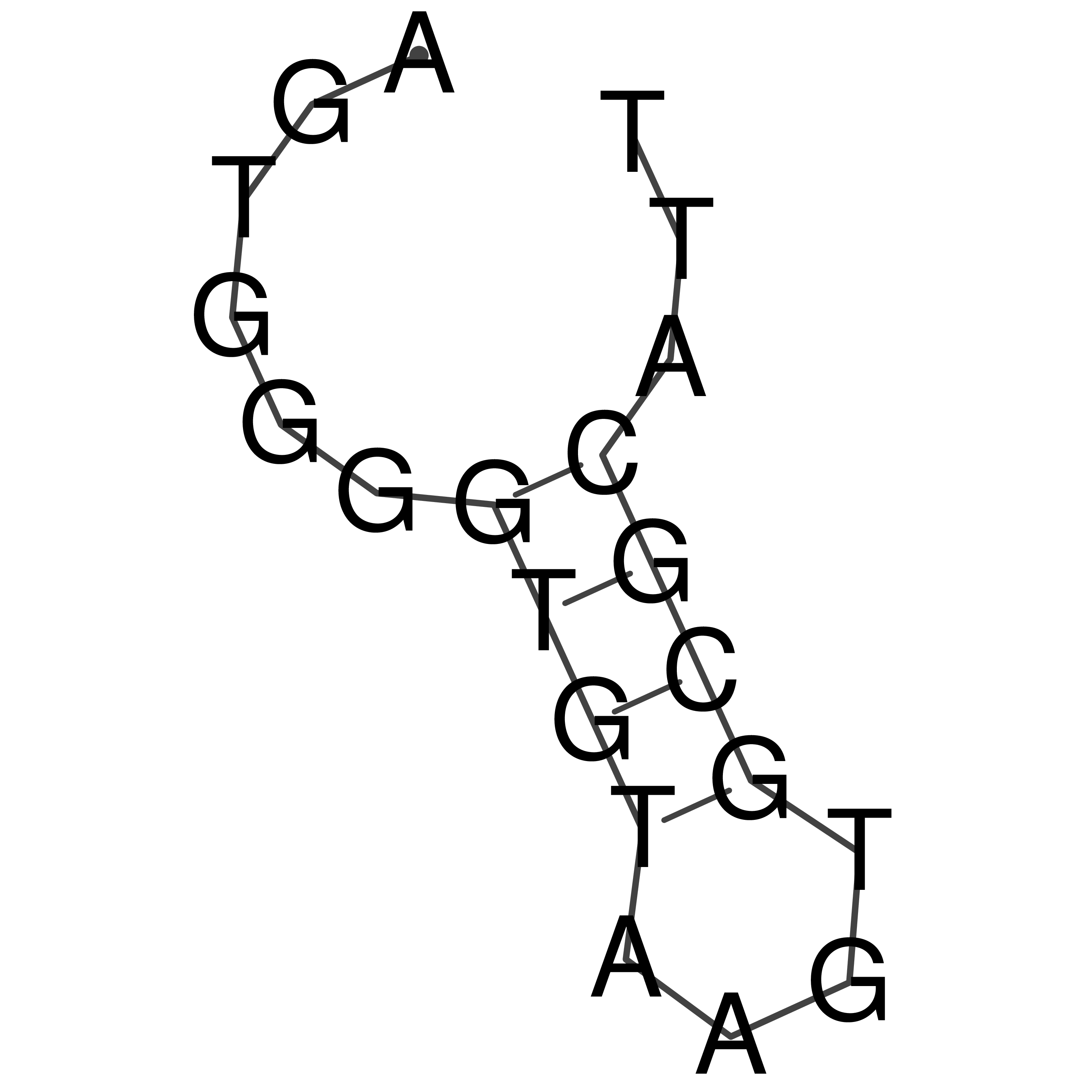
Relaxase
| ID | 10358 | GenBank | WP_160248129 |
| Name | mobQ_C7M34_RS16135_SRCM100995|unnamed4 |
UniProt ID | _ |
| Length | 687 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 687 a.a. Molecular weight: 80412.07 Da Isoelectric Point: 9.4536
>WP_160248129.1 MobQ family relaxase [Lactiplantibacillus plantarum]
MAIFHMSFSNISAGKGRSAIASAAYRSGEKLFDDKEGRHYFYARSIMPESFILTPKNSPEWASDREQLWN
EVEKKDRKSNSRYAKEFNVALPVELSESEQKELLTKYVQENFVDEGMVADVAIHRDHPDNPHAHVMLTNR
PFNSDGTWGIKSKKQYILDDNGNKTYTGTSKYPKSRKILMVDWDKKEKITEWRHNWAASVNQALEQKNIP
DRISEKSFVEQGIADTPMQHEGINSKRHERKAFNQQVKNYRKSKAGYKNMQEKVVNRGHLDSLSKHFSFN
EKKVVKELSHELKTYISLESLDDKRRLLFNWKNSTLIKHAVGEDVTKQLLTINQQESSLKKADELLNKVV
DRTTKKLYPELDFEQTTAAERRELIKETNSEQTIFKGSELNERLMNIRDDLLTRQLLTFTKRPYVGWKLL
MQQEKEVKIKLKYTLMIHDDSLESLEHVDQGLLEKYSPTEQQKITRAVKDLRTIMAVKQVIKTQYHEVLK
RAFPKGDLDELPMTKQEQAYTAVMYYDPVLKPCQAETIEQWQANPPQVFSPQEHQQGLAYLSGQLSLDQL
ENHHLQRVLKHDGTKQLFLGECKADPTIKNSQIEKIQKQLKGQQAKDDQYRKENIGHYQPLNYKPVSPDY
YLKTAFSDAIMTVLYARDEDYQRQKQERGLKETEWEMTKKQRQHQTRNRHEDGGMHL
MAIFHMSFSNISAGKGRSAIASAAYRSGEKLFDDKEGRHYFYARSIMPESFILTPKNSPEWASDREQLWN
EVEKKDRKSNSRYAKEFNVALPVELSESEQKELLTKYVQENFVDEGMVADVAIHRDHPDNPHAHVMLTNR
PFNSDGTWGIKSKKQYILDDNGNKTYTGTSKYPKSRKILMVDWDKKEKITEWRHNWAASVNQALEQKNIP
DRISEKSFVEQGIADTPMQHEGINSKRHERKAFNQQVKNYRKSKAGYKNMQEKVVNRGHLDSLSKHFSFN
EKKVVKELSHELKTYISLESLDDKRRLLFNWKNSTLIKHAVGEDVTKQLLTINQQESSLKKADELLNKVV
DRTTKKLYPELDFEQTTAAERRELIKETNSEQTIFKGSELNERLMNIRDDLLTRQLLTFTKRPYVGWKLL
MQQEKEVKIKLKYTLMIHDDSLESLEHVDQGLLEKYSPTEQQKITRAVKDLRTIMAVKQVIKTQYHEVLK
RAFPKGDLDELPMTKQEQAYTAVMYYDPVLKPCQAETIEQWQANPPQVFSPQEHQQGLAYLSGQLSLDQL
ENHHLQRVLKHDGTKQLFLGECKADPTIKNSQIEKIQKQLKGQQAKDDQYRKENIGHYQPLNYKPVSPDY
YLKTAFSDAIMTVLYARDEDYQRQKQERGLKETEWEMTKKQRQHQTRNRHEDGGMHL
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Host bacterium
| ID | 16632 | GenBank | NZ_CP028279 |
| Plasmid name | SRCM100995|unnamed4 | Incompatibility group | - |
| Plasmid size | 48008 bp | Coordinate of oriT [Strand] | 35338..35358 [+] |
| Host baterium | Lactiplantibacillus plantarum strain SRCM100995 |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |