Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 115422 |
| Name | oriT_p707804-NDM |
| Organism | Leclercia adecarboxylata |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_MH909331 (157424..157678 [+], 255 nt) |
| oriT length | 255 nt |
| IRs (inverted repeats) | 153..158, 160..165 (AAAAGT..ACTTTT) 122..130, 135..143 (TTAAGGCTT..AAGCCTTAA) 49..55, 60..66 (GAATTTT..AAAATTC) |
| Location of nic site | 78..79 |
| Conserved sequence flanking the nic site |
TTTGGTTAAA |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 255 nt
>oriT_p707804-NDM
GGATTTAGGTTTTTTTTTAATCGCTTCACATTTCGTTAGCATGCGCAGGAATTTTTGATAAAATTCTGGTTAGTTTGGTTAAAAAGTGTTACAAGTAAGCGGTGTGGTTGAAGGGATAGATTTAAGGCTTATTCAAGCCTTAAGAAAATACTAAAAGTTACTTTTCACCCTACCGAACACCTAACAAAAAATCCATGTTGAAGATTTGAACAATTGTAATGGCGCAAGGACAATCAGCACATGTCAGAATCTGAT
GGATTTAGGTTTTTTTTTAATCGCTTCACATTTCGTTAGCATGCGCAGGAATTTTTGATAAAATTCTGGTTAGTTTGGTTAAAAAGTGTTACAAGTAAGCGGTGTGGTTGAAGGGATAGATTTAAGGCTTATTCAAGCCTTAAGAAAATACTAAAAGTTACTTTTCACCCTACCGAACACCTAACAAAAAATCCATGTTGAAGATTTGAACAATTGTAATGGCGCAAGGACAATCAGCACATGTCAGAATCTGAT
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file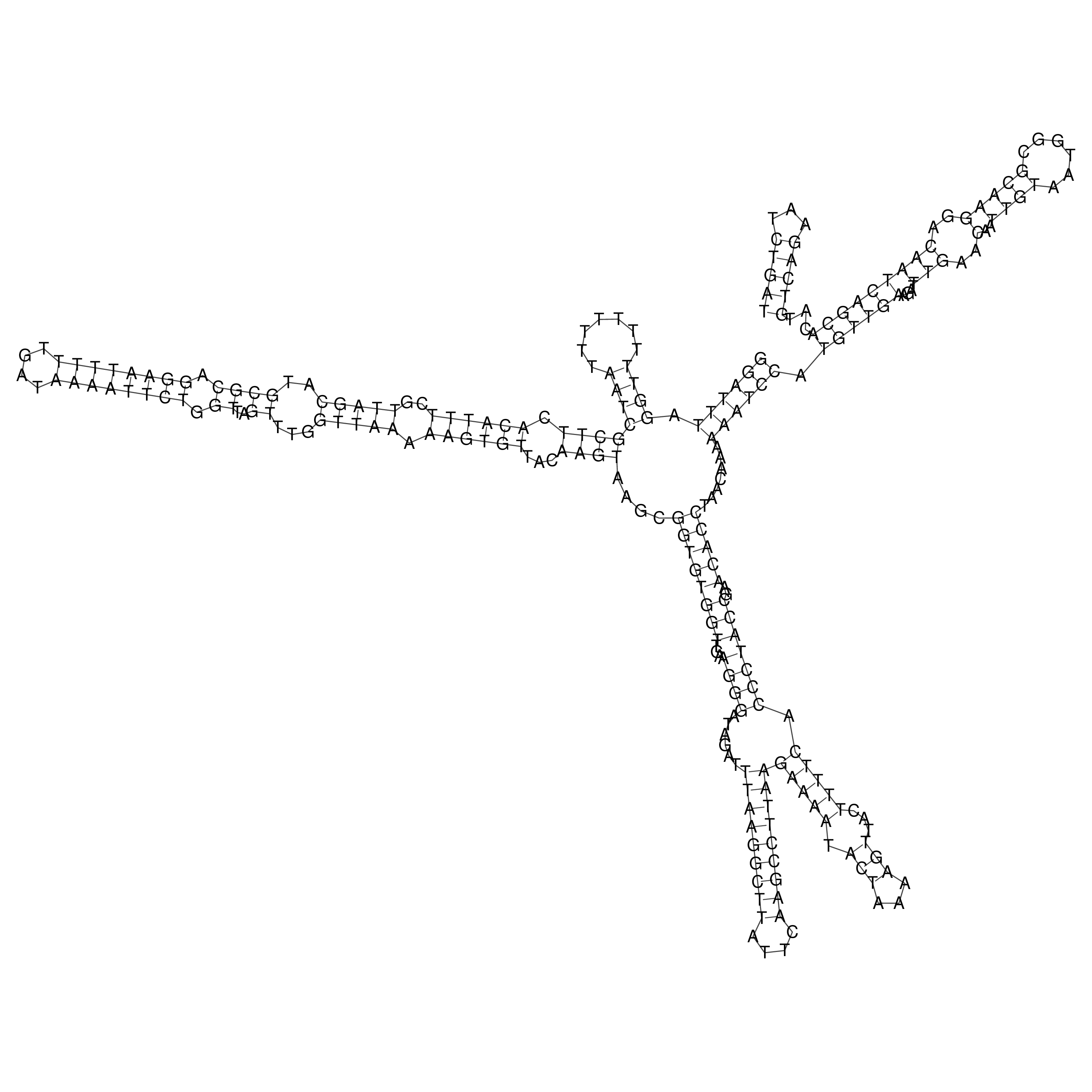
Relaxase
| ID | 9870 | GenBank | WP_012477394 |
| Name | mobH_HTU92_RS00805_p707804-NDM |
UniProt ID | A0A2Z2E7B6 |
| Length | 1050 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1050 a.a. Molecular weight: 118067.47 Da Isoelectric Point: 4.6347
>WP_012477394.1 MULTISPECIES: MobH family relaxase [Enterobacterales]
MMMNFRALYLCIKRILGIFSSQENDATSVMIEDISSLSPFAQILGDQKYTVPDHPNPEVLKFIEYPTRPT
GIQTFNEQSILSLYREKLHSISMMLAISDSDIRDDAYTFTNLVLKPLVEYVRWIHLLPASENHHHNGIGG
LLSHSLEVAILSLKNAHHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLASPIIWT
PSSQSLLDWARENDVVEYEIHWRKRIHNQHNIWSSVFLERILNPVCLAFLDRVNKERVYSKMITALNVYT
DGNDFLSKCVRTADFYSTGTDLNVLRDPIMGLRSNDAAARAISTIKHNFTSININNYNAKPMHIIIVNGE
VYLNENAFLDFVLNDFELHKYNFPQGEAGKTVLVESLVQRGYVEPYDDERVVHYFIPGIYSENEISNIFR
NGIGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRDVITKITDT
VETAVLKVNDLGRSSASIDVDIHSKKNEGSSDDFEKKAESDNEIDNDTQIVKSEGEEAADPVIPDIEESE
DESAKDTESHVLVNQLHELLLSAPLSNDYIVCVDAVPYLNIDTTMALLPGLDEKAFSEEPYFQLTFREGS
LDGMWIVRDIDDLRLVQLGDNCAGFQLTYHEPRRPTTLKSLFNTSMYQALVINDESSVENSAPRPKQTLE
LPPPRVNAVEEHSGDVEYHGTDSASATGPLKTEAVEYEHYQHLFEKEDEEHEIIDYTDFSQLSVSRPEVG
SCATSSSVHNEKLLSEPSELPELNREQNADPQGTNERSMDVSVGQENSEPDTEGNCPPPAEVVYSQTEAA
ATSVMASEEPALPPVLEESNGEHAPTDAKGHHLSPALARLFAPTAPVEKQNPKRNRNKSSDKAEVQKPAS
PVSGHNLNSKVFASTESDQNGEFSLISEGDVTELEFVEIALVLHQILSKMEVAFKRKRKNRFMVSTPNTL
YLTQSCVEKFGSQLEAQDLFNKLPQYLVNSGAVINTKCHAFNMPTLLAASDRAKVDIERIINNLKEAGNL
MMMNFRALYLCIKRILGIFSSQENDATSVMIEDISSLSPFAQILGDQKYTVPDHPNPEVLKFIEYPTRPT
GIQTFNEQSILSLYREKLHSISMMLAISDSDIRDDAYTFTNLVLKPLVEYVRWIHLLPASENHHHNGIGG
LLSHSLEVAILSLKNAHHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLASPIIWT
PSSQSLLDWARENDVVEYEIHWRKRIHNQHNIWSSVFLERILNPVCLAFLDRVNKERVYSKMITALNVYT
DGNDFLSKCVRTADFYSTGTDLNVLRDPIMGLRSNDAAARAISTIKHNFTSININNYNAKPMHIIIVNGE
VYLNENAFLDFVLNDFELHKYNFPQGEAGKTVLVESLVQRGYVEPYDDERVVHYFIPGIYSENEISNIFR
NGIGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRDVITKITDT
VETAVLKVNDLGRSSASIDVDIHSKKNEGSSDDFEKKAESDNEIDNDTQIVKSEGEEAADPVIPDIEESE
DESAKDTESHVLVNQLHELLLSAPLSNDYIVCVDAVPYLNIDTTMALLPGLDEKAFSEEPYFQLTFREGS
LDGMWIVRDIDDLRLVQLGDNCAGFQLTYHEPRRPTTLKSLFNTSMYQALVINDESSVENSAPRPKQTLE
LPPPRVNAVEEHSGDVEYHGTDSASATGPLKTEAVEYEHYQHLFEKEDEEHEIIDYTDFSQLSVSRPEVG
SCATSSSVHNEKLLSEPSELPELNREQNADPQGTNERSMDVSVGQENSEPDTEGNCPPPAEVVYSQTEAA
ATSVMASEEPALPPVLEESNGEHAPTDAKGHHLSPALARLFAPTAPVEKQNPKRNRNKSSDKAEVQKPAS
PVSGHNLNSKVFASTESDQNGEFSLISEGDVTELEFVEIALVLHQILSKMEVAFKRKRKNRFMVSTPNTL
YLTQSCVEKFGSQLEAQDLFNKLPQYLVNSGAVINTKCHAFNMPTLLAASDRAKVDIERIINNLKEAGNL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A2Z2E7B6 |
T4CP
| ID | 11507 | GenBank | WP_001284073 |
| Name | traD_HTU92_RS00800_p707804-NDM |
UniProt ID | _ |
| Length | 694 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 694 a.a. Molecular weight: 78123.91 Da Isoelectric Point: 8.0177
>WP_001284073.1 MULTISPECIES: conjugative transfer system coupling protein TraD [Enterobacterales]
MSDSKRTNLHAQENFYRPILEYRSASILLICSVSMLYMGLSSDGLDIAPIVLFTSILLFLLCLYRCKTAA
PFLMAHWRVFKRHFMFVSLDSLRVINKSNFFSNERKYRQLVQDYQNKNKDIPERKSYFCDGFEWGPEHAD
RAYQIANLSSDKREIELPFVFNPIKRHFDAMARKMGGSNAIFAVERREPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVVVVIDPKNDAEWRESLMEEAKTLGLPFYKFHPGQPASSVCIDVCNTYTNVSD
LTSRLLSLVTVPGEVNPFVQYAKALVSNVISGLSYIEKKPSIYLIHKNMKSHMSIVNLTVKVMESCYARY
YGYDVWTEKVKYVANDTLPVRFKRLAEWFTAHFMNYEGSEQIDWLDTVSQLIDYSMSDPEHMAKMTAGIM
PVFDMLIEKPLNELLSPNPNSVSSREIVTSEGMFSTGGVLYISLDGLSNPDTAAAISQLIMSDLTSCAGS
RYNAQDGDMSANSRISIFVDEAHSAINNPMINLLAQGRAAKIALFICTQTISDFIAAASVETANRITGLC
NNYISLRVNDTPTQTLVVENFGKSAISTNMVTYTTGSETSLPHNNFSGSISERKQTTLEESIPKDLLGQV
PMFHIVARLQDGRKVVGQIPIAVAEKQMKPNTTLSEMLFKKAGKVTLRQNLDIKNLNKFLRKLH
MSDSKRTNLHAQENFYRPILEYRSASILLICSVSMLYMGLSSDGLDIAPIVLFTSILLFLLCLYRCKTAA
PFLMAHWRVFKRHFMFVSLDSLRVINKSNFFSNERKYRQLVQDYQNKNKDIPERKSYFCDGFEWGPEHAD
RAYQIANLSSDKREIELPFVFNPIKRHFDAMARKMGGSNAIFAVERREPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVVVVIDPKNDAEWRESLMEEAKTLGLPFYKFHPGQPASSVCIDVCNTYTNVSD
LTSRLLSLVTVPGEVNPFVQYAKALVSNVISGLSYIEKKPSIYLIHKNMKSHMSIVNLTVKVMESCYARY
YGYDVWTEKVKYVANDTLPVRFKRLAEWFTAHFMNYEGSEQIDWLDTVSQLIDYSMSDPEHMAKMTAGIM
PVFDMLIEKPLNELLSPNPNSVSSREIVTSEGMFSTGGVLYISLDGLSNPDTAAAISQLIMSDLTSCAGS
RYNAQDGDMSANSRISIFVDEAHSAINNPMINLLAQGRAAKIALFICTQTISDFIAAASVETANRITGLC
NNYISLRVNDTPTQTLVVENFGKSAISTNMVTYTTGSETSLPHNNFSGSISERKQTTLEESIPKDLLGQV
PMFHIVARLQDGRKVVGQIPIAVAEKQMKPNTTLSEMLFKKAGKVTLRQNLDIKNLNKFLRKLH
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Host bacterium
| ID | 15855 | GenBank | NZ_MH909331 |
| Plasmid name | p707804-NDM | Incompatibility group | IncHI2A |
| Plasmid size | 306267 bp | Coordinate of oriT [Strand] | 157424..157678 [+] |
| Host baterium | Leclercia adecarboxylata |
Cargo genes
| Drug resistance gene | aac(6')-Ib-cr, blaOXA-1, catB3, ARR-3, qacE, sul1, blaNDM-1, qnrA1, msr(E), mph(E), mcr-9 |
| Virulence gene | - |
| Metal resistance gene | terW, terZ, terA, terB, terC, terD, terE, arsH, arsR, arsB, arsC, rcnR/yohL, ncrC, pcoE, pcoS, silP, silA, silB, silF, silC, silR, silS, silE, merR, merT, merP, merC, merE, merD, merA |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |