Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 115035 |
| Name | oriT_pPE52IMP |
| Organism | Pseudomonas aeruginosa strain PE52 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP102481 (9054..9164 [-], 111 nt) |
| oriT length | 111 nt |
| IRs (inverted repeats) | 71..78, 81..88 (CACCATGC..GCATGGTG) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 111 nt
>oriT_pPE52IMP
CCGGCTGGCCCCGCAGGGCAGGATGCCCCGTTGAGCGCCGCAGGCGCGAATAAGGGGAAGTGAAGAGGATCACCATGCTTGCATGGTGGGCCTACTTCACACATCCTGCCC
CCGGCTGGCCCCGCAGGGCAGGATGCCCCGTTGAGCGCCGCAGGCGCGAATAAGGGGAAGTGAAGAGGATCACCATGCTTGCATGGTGGGCCTACTTCACACATCCTGCCC
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file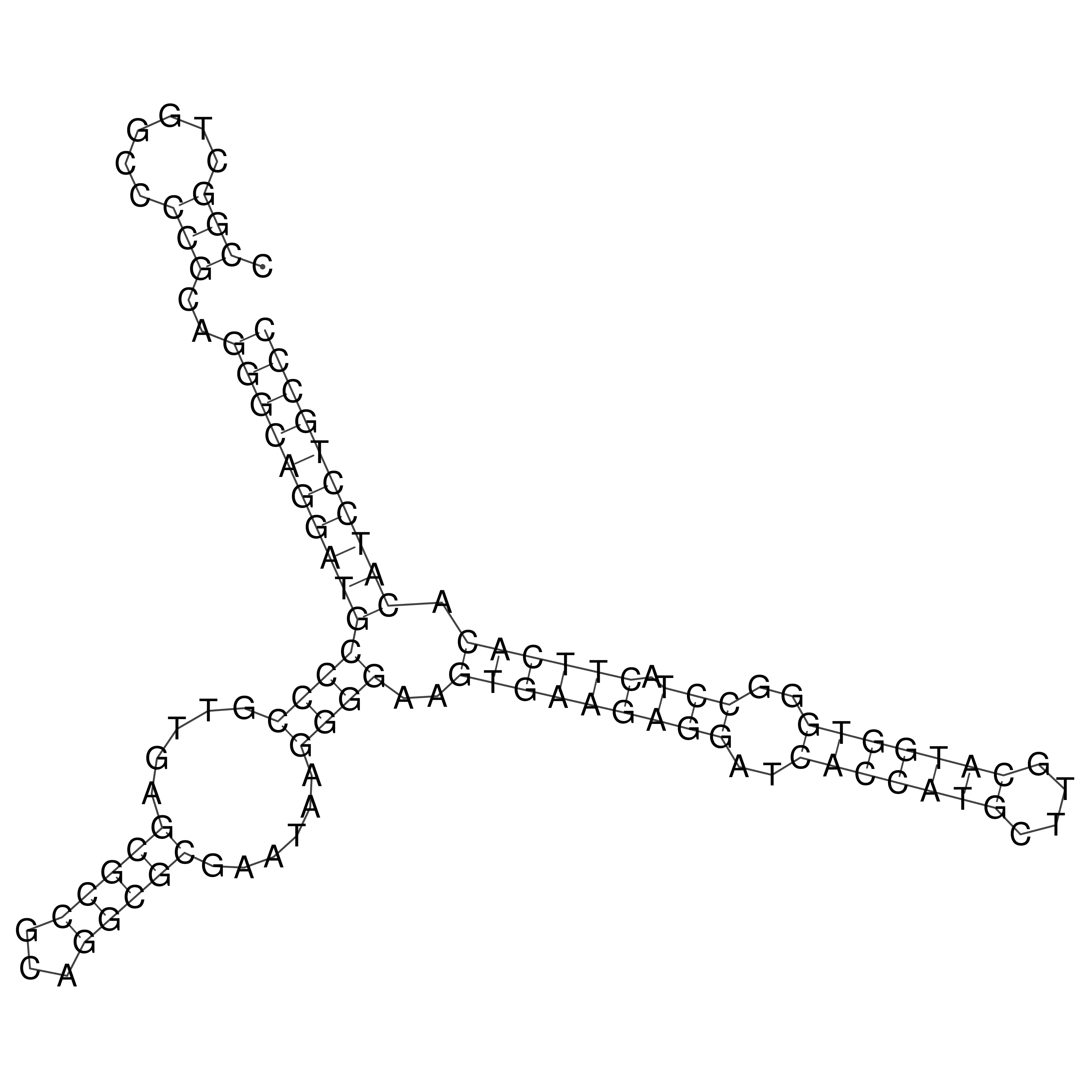
Relaxase
| ID | 9627 | GenBank | WP_033945278 |
| Name | traI_NUJ51_RS00070_pPE52IMP |
UniProt ID | A0A482F307 |
| Length | 609 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 609 a.a. Molecular weight: 66706.47 Da Isoelectric Point: 10.6661
>WP_033945278.1 MULTISPECIES: TraI/MobA(P) family conjugative relaxase [Pseudomonadota]
MIAKHVPMRSLGKSDFAGLVRYVTDAQSKDHRLGQVQVTNCAADTVRDAITEVLATQQANTRAKGDKTYH
LIVSFRAGEQPSADTLRQIEARICAGLGFAEHQRVSAVHTDTDNLHIHIAINKIHPTRGTMQEPYYSHRA
LADLCATLERDYGLERDNHEPRQRGAAARAGDLEQHAGVESLVGWIKRECLDEIKGAASWAELHQVMRDN
GLALRARGNGLVIEAGDGTIIKASTLARDLSKPALEKRLGTFEASPERQATPARRSYQKGRPMRLRGGLD
TTELYARYKGAQQNLTAARAGVLAEAKRRKDRAIENAKKAAATRRAAIKHLTDGRLTKKLLYSQASAALR
ADLEAIHKRHDQERAAIYNEYRRRPWADWLKQQATAGDGQALDALRAREAAHGLKGDTIQGDGPAKPGHA
PVIDNITKKGTIIYRAGSSAVRDDGERLAVSSQADRAGVAAALRLAMERYGERITVSGSVEFKAAVIRAA
VDAQLPITFADPALESRRQAEATKAQAKRRSGVPPVGQEPPPHRRNGLHTLGQLDRLHIEGEARRPAQAP
TQATPTPPAAKVQQPARPAPSELEQREARLRAQMEAKKAKRQEGKKRGR
MIAKHVPMRSLGKSDFAGLVRYVTDAQSKDHRLGQVQVTNCAADTVRDAITEVLATQQANTRAKGDKTYH
LIVSFRAGEQPSADTLRQIEARICAGLGFAEHQRVSAVHTDTDNLHIHIAINKIHPTRGTMQEPYYSHRA
LADLCATLERDYGLERDNHEPRQRGAAARAGDLEQHAGVESLVGWIKRECLDEIKGAASWAELHQVMRDN
GLALRARGNGLVIEAGDGTIIKASTLARDLSKPALEKRLGTFEASPERQATPARRSYQKGRPMRLRGGLD
TTELYARYKGAQQNLTAARAGVLAEAKRRKDRAIENAKKAAATRRAAIKHLTDGRLTKKLLYSQASAALR
ADLEAIHKRHDQERAAIYNEYRRRPWADWLKQQATAGDGQALDALRAREAAHGLKGDTIQGDGPAKPGHA
PVIDNITKKGTIIYRAGSSAVRDDGERLAVSSQADRAGVAAALRLAMERYGERITVSGSVEFKAAVIRAA
VDAQLPITFADPALESRRQAEATKAQAKRRSGVPPVGQEPPPHRRNGLHTLGQLDRLHIEGEARRPAQAP
TQATPTPPAAKVQQPARPAPSELEQREARLRAQMEAKKAKRQEGKKRGR
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A482F307 |
Auxiliary protein
| ID | 5662 | GenBank | WP_011997489 |
| Name | WP_011997489_pPE52IMP |
UniProt ID | A0A482F300 |
| Length | 125 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
Auxiliary protein sequence
Download Length: 125 a.a. Molecular weight: 13548.38 Da Isoelectric Point: 8.9596
>WP_011997489.1 MULTISPECIES: conjugal transfer transcriptional regulator TraJ [Pseudomonadota]
MEANENKVTRKGSTPIKVYCLPDERAAIEANAQAAGMSTSAYLLAVGQGYQVKGVTDFENVRDLVRVNGD
LGRLGGLLKLWLTDDPRTARFGDDTIRALLARIEATQDKISTIAESVVRPRGGRS
MEANENKVTRKGSTPIKVYCLPDERAAIEANAQAAGMSTSAYLLAVGQGYQVKGVTDFENVRDLVRVNGD
LGRLGGLLKLWLTDDPRTARFGDDTIRALLARIEATQDKISTIAESVVRPRGGRS
Protein domains
No domain identified.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A482F300 |
Host bacterium
| ID | 15470 | GenBank | NZ_CP102481 |
| Plasmid name | pPE52IMP | Incompatibility group | - |
| Plasmid size | 27635 bp | Coordinate of oriT [Strand] | 9054..9164 [-] |
| Host baterium | Pseudomonas aeruginosa strain PE52 |
Cargo genes
| Drug resistance gene | blaIMP-56, ant(3'')-Ia, blaOXA-2, qacE, sul1 |
| Virulence gene | - |
| Metal resistance gene | merE, merD, merA, merP, merT, merR |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |