Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 114804 |
| Name | oriT_253_2_EW_B|plas1 |
| Organism | Escherichia albertii strain 253_2_EW_B |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP099872 (56052..56175 [+], 124 nt) |
| oriT length | 124 nt |
| IRs (inverted repeats) | 101..106, 119..124 (TTTAAT..ATTAAA) 91..99, 113..121 (AATAATGTA..TACATTATT) 90..95, 107..112 (AAATAA..TTATTT) 39..46, 49..56 (GCAAAAAC..GTTTTTGC) 3..10, 15..22 (TTGGTGGT..ACCACCAA) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 124 nt
GGTTGGTGGTTCTCACCACCAAAAGCACCACACCCCACGCAAAAACAAGTTTTTGCTGATTTGCTTTTTGAATCATTAGCTTATGTTTTAAATAATGTATTTTAATTTATTTTACATTATTAAA
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file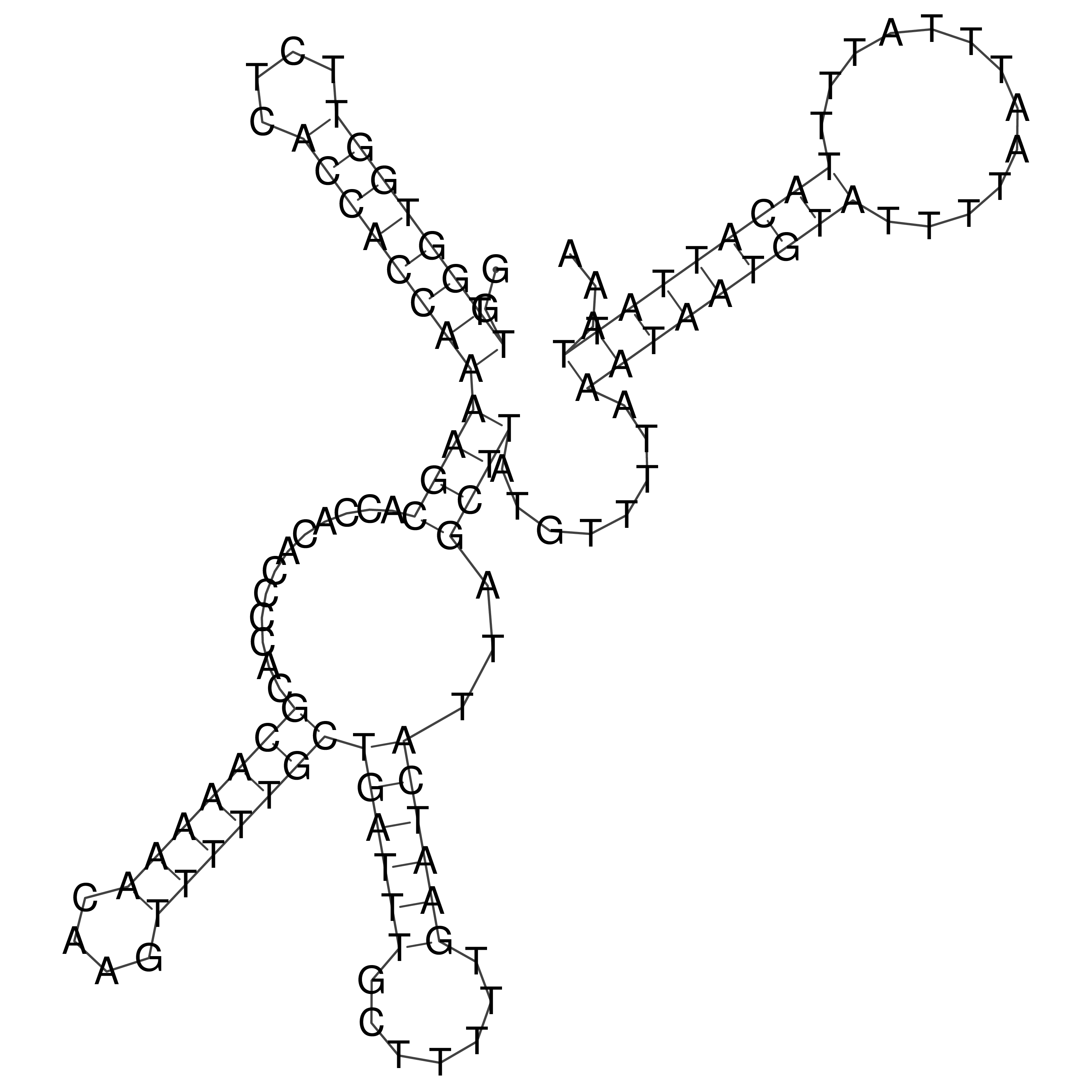
Relaxase
| ID | 9488 | GenBank | WP_256878393 |
| Name | traI_NIZ24_RS23270_253_2_EW_B|plas1 |
UniProt ID | _ |
| Length | 1756 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1756 a.a. Molecular weight: 191896.68 Da Isoelectric Point: 5.7978
MMSIAQVRSAGSAGNYYTDKDNYYVLGSMGERWAGRGAEQLGLQGSVDKDVFTRLLEGKLPDGADLSRMQ
DGSNRHRPGYDLTFSAPKSVSMMAMLGGDKRLIDAHNQAVDFAVRQVEALASTRVMTDGQSETVLTGNLV
MALFNHDTSRDQEPQLHTHAVVANVTQHNGEWKTLSSDKVGKTGFIENVYANQIAFGRLYREKLKEQVEA
LGYETEVVGKHGMWEMPGVPVEAFSGRSQAIREAVGEDASLKSRDVAALDTRKSKQHVDPEVRMAEWMQT
LKETGFDIRAYRDAADQRAETRTQAPGAVSQEGPDVQQAVTQAIAGLSERKVQFTYTDVLARTVGILPPE
NGVIERARAGIDEAISREQLIPLDREKGLFTSGIYVLDELSVRALSRDIMKQNRVTVHPEKSVPRTAGYS
DAVSVLAQDRPSLAIVSGQGGAAGQRERVAELVMMAREQGREVQIIAADRRSQMNLKQDERLSGELITGR
RQLLEGMAFTPGSTVIVDQGEKLSLKETLTLLDGAARHNVQVLITDSGQRTGTGSALMAMKDAGVNTYRW
QGGEQRPATIISEPDRNVRYARLAGDFAASVKAGEESVAQVSGVREQAILTQAIRSELKTQGVLGHPEMT
MTALSPVWLDSRSRYLRDMYRPGMVMEQWNPETRSHDRYVIDRVTAQSHSLTLRDAQGETQVVRISSLDS
SWSLFRPEKMPVADGERLRVTGKIPGLRVSGGDRLQVASVSEDAMTVVVPGRAEPASLPVSDSPFTALKL
ENGWVETPGHSVSDSAKVFASVTQMAMDNATLNGLARSGRDVRLYSSLDETRTAEKLARHPSFTVVSEQI
KARAGETLLETAISLQKAGLHTPAQQAIHLALPVLESKNLAFSMVDLLTEAKSFAAEGTSFIDLGGEINA
QIKRGDLLYVDVAKGYGTGLLVSRASYEAEKSILRHILEGKEAVTPLMERVPGELMEKLTSGQRAATRMI
LETSDRFTVVQGYAGVGKTTQFRAVMSAVNMLPESERPRVVGLGPTHRAVGEMRSAGVDAQTLASFLHDT
QLQQRSGETPDFSNTLFLLDESSMVGNTDMARAYALIAAGGGRAVASGDTDQLQAIAPGQPFRLQQTRSA
ADVVIMKEIVRQTPELREAVYSLINRDVERALSGLESVKPSQVPRQEGAWAPEHSVTEFSHSQEAKLAEA
QQKAMLKGEAFPDIPMTLYEAIVRDYTGRTPEAREQTLIVTHLNEDRRVLNSMIHDAREKAGELGKEQVM
VPVLNTANIRDGELRRLSTWENNPDALALVDSVYHRIAGISKDDGLITLQDAEGNTRLISPREAVAEGVT
LYTPDTIRVGTGDRMRFTKSDRERGYVANSVWTVTAVSGDSVTLSDGQQTRVIHPGQERAEQHLDLAYAI
TAHGAQGASETFAIALEGTEGNRKQMAGFESAYVALSRMKQHVQVYTDNRQGWTDAINNAVQKGTAHDVF
EPKPDREVMNAERLFSTARELRDVAAGRAVLRQAGLAGGDSPARFIAPGRKYPQPYVALPAFDRNGKSAG
IWLNSLTTDDGNGLRGFSGEGRVKGSGDAQFVALQGSRNGESLLADNMQDGVRIARDNPDSGVVVRIAGE
GRPWNPGAITGGRVWGDIPDNSVQPGAGNGEPVTAEVLAQRQAEEAIRRETERRADEIVRKMAENKPDLP
DGKTELAVREIAGQERDRAAITEREAALPESVLREPQREREAVREVARENLLQERLQQMERDMVRDLQKE
KTLGGD
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Auxiliary protein
| ID | 5569 | GenBank | WP_000124981 |
| Name | WP_000124981_253_2_EW_B|plas1 |
UniProt ID | A0A630FTW9 |
| Length | 127 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
Auxiliary protein sequence
Download Length: 127 a.a. Molecular weight: 14542.59 Da Isoelectric Point: 5.0278
MARVNLYISNEVHEKINMIVEKRRQEGARDKDISLSGTASMLLELGLRVYDAQMERKESAFNQTEFNKLL
LECVVKTQSTVAKILGIESLSPHVSGNPKFEYASMVDDIREKVSVEMDRFFPKNDDE
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A630FTW9 |
| ID | 5570 | GenBank | WP_001254386 |
| Name | WP_001254386_253_2_EW_B|plas1 |
UniProt ID | A0A3Z6KJB5 |
| Length | 75 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
Auxiliary protein sequence
Download Length: 75 a.a. Molecular weight: 9005.11 Da Isoelectric Point: 10.1422
MRRRNARGGISRTVSVYLDEDTNNRLIKAKDRSGRSKTIEVQIRLRDHLKRFPDFYNEEIFREVTEESES
TFKEL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A3Z6KJB5 |
T4CP
| ID | 11045 | GenBank | WP_256878435 |
| Name | traC_NIZ24_RS23175_253_2_EW_B|plas1 |
UniProt ID | _ |
| Length | 876 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 876 a.a. Molecular weight: 99347.12 Da Isoelectric Point: 6.3032
MSNNPLEAVTQAVNSLVTALKLPDESAKANEVLGEMSFPQFSRLLPYRDYNQESGLFMNDTTMGFMLEAI
PINGANESIVEALDHMLRTKLPRGIPLCIHLMSSQLVGDRIEYGLREFSWSGEQAERFNAITRAYYMKAA
ATQFPLPEGMNLPLTLRHYRVFISYCSPSKKKSRADILEMENLVKIIRASLQGASITTQTVDAQAFIDIV
GEMINHNPDSLYPKRRQLDPYSDLNYQCVEDSFDLKVRADYLTLGLRESGRNSTARILNFHLARNPEIAF
LWNMADNYSNLLNPELSISCPFILTLTLVVEDQVKTHSEANLKYMDLEKKSKTSYAKWFPSVEKEAKEWG
ELRQRLGSGQSSVVSYFLNITAFCKDNNETALEVEQDILNSFRKNGFELISPRFNHMRNFLTCLPFMAGK
GLFKQLKEAGVVQRAESFNVANLMPLVADNPLTPAGLLAPTYRNQLAFIDIFFRGMNNTNYNMAVCGTSG
AGKTGLIQPLIRSVLDSGGFAVVFDMGDGYKSLCENMGGVYLDGETLRFNPFANITDIDQSAERVRDQLS
VMASPNGNLDEVHEGLLLQAVRASWLAKKNKARIDDVVDFLKNARDNDQYVESPTIRSRLDEMIVLLDQY
TAGGTYGRYFNSDEPSLRDDAKMVVLELGGLEDRPSLLVAVMFSLIIYIENRMYRTPRNLKKLNVIDEGW
RLLDFKNHKVGEFIEKGYRTARRHTGAYITITQNIVDFDSDKASSAARAAWGNSSYKIILKQSAKEFAKY
NQLYPDQFQPLQRDMIGKFGAARDQWFSSFLLQVENHSSWHRLFVDPLSRAMYSSDGPDFEFVQQKRKEG
LSIHEAVWQLAWKKSGPEMASLEAWLEEHEKYRSVA
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
| ID | 11046 | GenBank | WP_256878392 |
| Name | traD_NIZ24_RS23265_253_2_EW_B|plas1 |
UniProt ID | _ |
| Length | 717 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 717 a.a. Molecular weight: 81468.13 Da Isoelectric Point: 5.1711
MSFNAKDMTQGGQIASMRIRMFSQIANIMLYCLFIFFWILVGLVLWVKISWQTFVNGCIYWWCTTLEGMR
DLIKSQPVYEIQYYGKTFRMNAAQVLHDKYMIWCGEQLWSAFVLATVVALVICLITFFVVSWILGRQGKQ
QSENEVTGGRQLTDNPKDVARMLKKDGKDSDIRIGDLPIIRDSEIQNFCLHGTVSTGKSEVIRRLANYAR
KRGDMVVIYDRSCEFVKSYYDPSIDKILNPLDARCAAWDLWKECLTQPDFDNVANTLIPMGTKEDPFWQG
SGRTIFAEAAYLMRNDPNRSYSKLVDTLLSIKIEKLRTYLRNSPAANLVEEKIEKTAISIRAVLTNYVKA
IRYLQGIEHNGESFTIRDWMRGVREDKKNGWLFISSNADTHGSLKPVVSMWLSIAIRGLLAMGENRNRRV
WFFCDELPTLHKLPDLVEILPEARKFGGCYVFGIQSYAQLEDIYGEKAAATLFDVLNTRAFFRSPSHQIA
EFAAGEIGEKEHLKASLQYSYGADPVRDGISTGKEMERQTLVSYSDIQSLPDLTCYVTLPGPYPAVKLSL
KYQARPKIAPEFIPREINPEMENRLSAVLAAREAEGRQVASLFEPDVPEVVSGEDVTQAEQPQQPVSPAI
NDKKSDAGVNVPAGGIEPELKMKPEEEMEQQLPPGISESGEVVDMAAYEAWQQENHPDIQQQMQRREEVN
INVHRERGEDVEPGDDF
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 55480..79591
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| NIZ24_RS23040 (NIZ24_23025) | 51448..51882 | + | 435 | WP_032217500 | conjugation system SOS inhibitor PsiB | - |
| NIZ24_RS23045 (NIZ24_23030) | 51879..52641 | + | 763 | Protein_53 | plasmid SOS inhibition protein A | - |
| NIZ24_RS23050 (NIZ24_23035) | 52610..52798 | - | 189 | WP_001336239 | hypothetical protein | - |
| NIZ24_RS23055 (NIZ24_23040) | 52820..52969 | + | 150 | Protein_55 | plasmid maintenance protein Mok | - |
| NIZ24_RS23060 (NIZ24_23045) | 52911..53036 | + | 126 | WP_001372321 | type I toxin-antitoxin system Hok family toxin | - |
| NIZ24_RS23065 (NIZ24_23050) | 53256..53486 | + | 231 | WP_001426396 | hypothetical protein | - |
| NIZ24_RS23070 (NIZ24_23055) | 53484..53657 | - | 174 | Protein_58 | hypothetical protein | - |
| NIZ24_RS23075 (NIZ24_23060) | 53727..53933 | + | 207 | WP_000547971 | hypothetical protein | - |
| NIZ24_RS23080 (NIZ24_23065) | 53958..54244 | + | 287 | Protein_60 | hypothetical protein | - |
| NIZ24_RS23085 (NIZ24_23070) | 54362..55183 | + | 822 | WP_227479334 | DUF932 domain-containing protein | - |
| NIZ24_RS23090 (NIZ24_23075) | 55480..56127 | - | 648 | WP_000614935 | transglycosylase SLT domain-containing protein | virB1 |
| NIZ24_RS23095 (NIZ24_23080) | 56404..56787 | + | 384 | WP_000124981 | conjugal transfer relaxosome DNA-binding protein TraM | - |
| NIZ24_RS23100 (NIZ24_23085) | 56978..57664 | + | 687 | WP_256878428 | PAS domain-containing protein | - |
| NIZ24_RS23105 (NIZ24_23090) | 57759..57986 | + | 228 | WP_001254386 | conjugal transfer relaxosome protein TraY | - |
| NIZ24_RS23110 (NIZ24_23095) | 58020..58385 | + | 366 | WP_000340270 | type IV conjugative transfer system pilin TraA | - |
| NIZ24_RS23115 (NIZ24_23100) | 58400..58711 | + | 312 | WP_000012106 | type IV conjugative transfer system protein TraL | traL |
| NIZ24_RS23120 (NIZ24_23105) | 58733..59299 | + | 567 | WP_000399792 | type IV conjugative transfer system protein TraE | traE |
| NIZ24_RS23125 (NIZ24_23110) | 59286..60014 | + | 729 | WP_001230787 | type-F conjugative transfer system secretin TraK | traK |
| NIZ24_RS23130 (NIZ24_23115) | 60014..61441 | + | 1428 | WP_256878430 | F-type conjugal transfer pilus assembly protein TraB | traB |
| NIZ24_RS23135 (NIZ24_23120) | 61431..62021 | + | 591 | WP_000002784 | conjugal transfer pilus-stabilizing protein TraP | - |
| NIZ24_RS23140 (NIZ24_23125) | 62008..62328 | + | 321 | WP_001057307 | conjugal transfer protein TrbD | - |
| NIZ24_RS23145 (NIZ24_23130) | 62325..62840 | + | 516 | WP_256878431 | type IV conjugative transfer system lipoprotein TraV | traV |
| NIZ24_RS23150 (NIZ24_23135) | 62975..63196 | + | 222 | WP_256878432 | conjugal transfer protein TraR | - |
| NIZ24_RS23155 (NIZ24_23140) | 63189..63602 | + | 414 | WP_256878433 | hypothetical protein | - |
| NIZ24_RS23160 (NIZ24_23145) | 63595..64068 | + | 474 | WP_256878434 | hypothetical protein | - |
| NIZ24_RS23165 (NIZ24_23150) | 64148..64393 | + | 246 | WP_032083161 | hypothetical protein | - |
| NIZ24_RS23170 (NIZ24_23155) | 64390..64752 | + | 363 | WP_042972088 | hypothetical protein | - |
| NIZ24_RS23175 (NIZ24_23160) | 64878..67508 | + | 2631 | WP_256878435 | type IV secretion system protein TraC | virb4 |
| NIZ24_RS23180 (NIZ24_23165) | 67505..67891 | + | 387 | WP_256878436 | type-F conjugative transfer system protein TrbI | - |
| NIZ24_RS23185 (NIZ24_23170) | 67888..68520 | + | 633 | WP_001203720 | type-F conjugative transfer system protein TraW | traW |
| NIZ24_RS23190 (NIZ24_23175) | 68517..69509 | + | 993 | WP_201477276 | conjugal transfer pilus assembly protein TraU | traU |
| NIZ24_RS23195 (NIZ24_23180) | 69538..69873 | + | 336 | WP_256878437 | hypothetical protein | - |
| NIZ24_RS23200 (NIZ24_23185) | 69866..70318 | + | 453 | WP_256878438 | hypothetical protein | - |
| NIZ24_RS23205 (NIZ24_23190) | 70346..70984 | + | 639 | WP_256878439 | type-F conjugative transfer system pilin assembly protein TrbC | trbC |
| NIZ24_RS23210 (NIZ24_23195) | 70981..72789 | + | 1809 | WP_256878440 | type-F conjugative transfer system mating-pair stabilization protein TraN | traN |
| NIZ24_RS23215 (NIZ24_23200) | 72816..73073 | + | 258 | WP_000864325 | conjugal transfer protein TrbE | - |
| NIZ24_RS23220 (NIZ24_23205) | 73066..73809 | + | 744 | WP_256878390 | type-F conjugative transfer system pilin assembly protein TraF | traF |
| NIZ24_RS23225 (NIZ24_23210) | 73825..74172 | + | 348 | WP_001287677 | conjugal transfer protein TrbA | - |
| NIZ24_RS23230 (NIZ24_23215) | 74291..74575 | + | 285 | WP_000624108 | type-F conjugative transfer system pilin chaperone TraQ | - |
| NIZ24_RS23235 (NIZ24_23220) | 74562..75107 | + | 546 | WP_256878391 | type-F conjugative transfer system pilin assembly thiol-disulfide isomerase TrbB | traF |
| NIZ24_RS23240 (NIZ24_23225) | 75037..75399 | + | 363 | WP_001443292 | P-type conjugative transfer protein TrbJ | - |
| NIZ24_RS23245 (NIZ24_23230) | 75399..76772 | + | 1374 | WP_001137358 | conjugal transfer pilus assembly protein TraH | traH |
| NIZ24_RS23250 (NIZ24_23235) | 76769..79591 | + | 2823 | WP_001007026 | conjugal transfer mating-pair stabilization protein TraG | traG |
| NIZ24_RS23255 (NIZ24_23240) | 79606..80109 | + | 504 | WP_000605510 | hypothetical protein | - |
| NIZ24_RS23260 (NIZ24_23245) | 80141..80872 | + | 732 | WP_000850424 | conjugal transfer complement resistance protein TraT | - |
| NIZ24_RS23265 (NIZ24_23250) | 81124..83277 | + | 2154 | WP_256878392 | type IV conjugative transfer system coupling protein TraD | - |
Host bacterium
| ID | 15239 | GenBank | NZ_CP099872 |
| Plasmid name | 253_2_EW_B|plas1 | Incompatibility group | IncFII |
| Plasmid size | 145531 bp | Coordinate of oriT [Strand] | 56052..56175 [+] |
| Host baterium | Escherichia albertii strain 253_2_EW_B |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | iucA, iucB, iucC, iucD, iutA, gspG, gspH, vat |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |