Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 114450 |
| Name | oriT_pG3L-1 |
| Organism | Leclercia sp. G3L |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP088263 (252403..252687 [+], 285 nt) |
| oriT length | 285 nt |
| IRs (inverted repeats) | 184..189, 191..196 (AAAAGT..ACTTTT) |
| Location of nic site | 109..110 |
| Conserved sequence flanking the nic site |
TTTGGTTAAA |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 285 nt
>oriT_pG3L-1
GGTTATTGCTACTTAATGCCGATAACGACTCAGGCTTTGAGGTTTTTTTATACGGTTCACATTTCGTTAGCAAGGTCAGGGTTTTTTGATAAAATTCTGGTTAGTTTGGTTAAAAAGTGTTACAAGTAAGGGTAATGGCTGAAAGGTTAGTTTTAAGGTTCAAAGCGGCAGTATTAAAATTCCAAAAGTTACTTTTCATCCTTCAGAATCCAGACCTTAATTTCATGTAGAAGATTCGTACAATTGTATTGGCGCAAGGACAATCCGCACATGTCAGAATCAGAT
GGTTATTGCTACTTAATGCCGATAACGACTCAGGCTTTGAGGTTTTTTTATACGGTTCACATTTCGTTAGCAAGGTCAGGGTTTTTTGATAAAATTCTGGTTAGTTTGGTTAAAAAGTGTTACAAGTAAGGGTAATGGCTGAAAGGTTAGTTTTAAGGTTCAAAGCGGCAGTATTAAAATTCCAAAAGTTACTTTTCATCCTTCAGAATCCAGACCTTAATTTCATGTAGAAGATTCGTACAATTGTATTGGCGCAAGGACAATCCGCACATGTCAGAATCAGAT
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file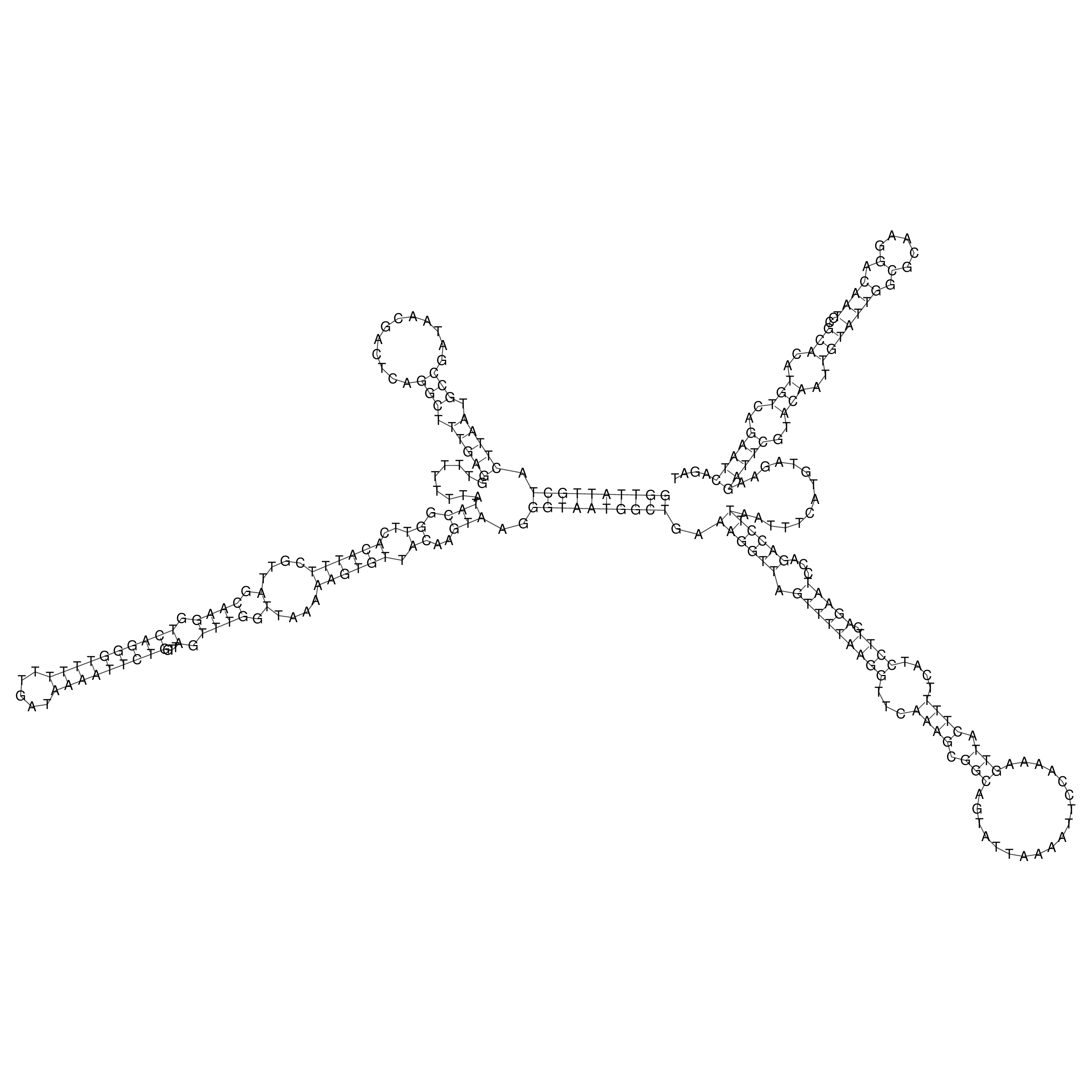
Relaxase
| ID | 9264 | GenBank | WP_039026431 |
| Name | mobH_LRS40_RS01335_pG3L-1 |
UniProt ID | A0A8E6NZY6 |
| Length | 1011 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1011 a.a. Molecular weight: 112997.99 Da Isoelectric Point: 4.5108
>WP_039026431.1 MULTISPECIES: MobH family relaxase [Enterobacterales]
MNFRALFLSMQRVFGIFSRRENDVSELMMKDAANFSPFAQIIGEQKYTVPDHPNPEVLKFIEYPTRPAGI
QTFNEQSILSLYRDKLHSISMMLAISDGDIREDAYTFTNLVLKPLIEYIRWIHLLPASENHHHNGIGGLL
SHSLEVAMISLKNANHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLSETLTWAPS
SQSLLDWARENNVVEYEIHWRKRIHNQHNIWSSVFLERILDPVCMSFLDRVKKERVYAKMVTALNVYNDG
NDFLSKCVRTSDYYSTGTDLNVLRDPIMGLRSNDAAARAIGTIKHNFTSININNYKTKPMHIIIVNGEVY
LNENAFLDFVLSDFAAHKFNFPQGDAGKTVLVESLVLRGYVEPYDDERVVHYFIPGTYSENEIASIFRNG
IGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRESVTRVTDTVN
DAIENTPQYGLQLVNGPDADSNNIISSENTESITDSLEESGADISNEIFETQVVTAIDTAETVNADEPEQ
VEEHDDRSQIHLVEQLHEMLLSAPLPHHAVINIDSVPYLDLDAAIALIPGIDEAAFCNGPFFQLTYRDGS
LDGMWIVRDVNNLRLIQLGDNCAGMQVSTSEPRNTSSLKSLFDTSMYQPLDIPEAPSVNEAASPPQTPLE
LPQPRLNAPVAEEASSVAEQTNAHSEPDSVIATEYEQYGHLLEETLDSDGEAYSDLIASDSTEAEYSATD
PQSSDFAQLPRETALSVAPGDLDYSEGAIKPPAPDATGKETILTSPEPAEDVRETVAAVEKASHLSPALA
RLFAVSTHAEKKHEKTQEPSPVKEVKNPTSSTTVKAPISIEPPGAEEKEAVEEFTLLNDGEVTELEYVEI
ATMLHQILTKLSGSFKRKRKNRFMVLTQNTFYLTQSCIEKYGTQLNAPELFNQLPQYQVTSGAVVNTKCI
AFNIPTLVAASDRAKVDIELIINKLKEVGNL
MNFRALFLSMQRVFGIFSRRENDVSELMMKDAANFSPFAQIIGEQKYTVPDHPNPEVLKFIEYPTRPAGI
QTFNEQSILSLYRDKLHSISMMLAISDGDIREDAYTFTNLVLKPLIEYIRWIHLLPASENHHHNGIGGLL
SHSLEVAMISLKNANHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLSETLTWAPS
SQSLLDWARENNVVEYEIHWRKRIHNQHNIWSSVFLERILDPVCMSFLDRVKKERVYAKMVTALNVYNDG
NDFLSKCVRTSDYYSTGTDLNVLRDPIMGLRSNDAAARAIGTIKHNFTSININNYKTKPMHIIIVNGEVY
LNENAFLDFVLSDFAAHKFNFPQGDAGKTVLVESLVLRGYVEPYDDERVVHYFIPGTYSENEIASIFRNG
IGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRESVTRVTDTVN
DAIENTPQYGLQLVNGPDADSNNIISSENTESITDSLEESGADISNEIFETQVVTAIDTAETVNADEPEQ
VEEHDDRSQIHLVEQLHEMLLSAPLPHHAVINIDSVPYLDLDAAIALIPGIDEAAFCNGPFFQLTYRDGS
LDGMWIVRDVNNLRLIQLGDNCAGMQVSTSEPRNTSSLKSLFDTSMYQPLDIPEAPSVNEAASPPQTPLE
LPQPRLNAPVAEEASSVAEQTNAHSEPDSVIATEYEQYGHLLEETLDSDGEAYSDLIASDSTEAEYSATD
PQSSDFAQLPRETALSVAPGDLDYSEGAIKPPAPDATGKETILTSPEPAEDVRETVAAVEKASHLSPALA
RLFAVSTHAEKKHEKTQEPSPVKEVKNPTSSTTVKAPISIEPPGAEEKEAVEEFTLLNDGEVTELEYVEI
ATMLHQILTKLSGSFKRKRKNRFMVLTQNTFYLTQSCIEKYGTQLNAPELFNQLPQYQVTSGAVVNTKCI
AFNIPTLVAASDRAKVDIELIINKLKEVGNL
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4CP
| ID | 10741 | GenBank | WP_000167418 |
| Name | traD_LRS40_RS01330_pG3L-1 |
UniProt ID | _ |
| Length | 694 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 694 a.a. Molecular weight: 77969.74 Da Isoelectric Point: 8.7275
>WP_000167418.1 MULTISPECIES: conjugative transfer system coupling protein TraD [Enterobacterales]
MTKSKRTNLHAQENFYRPILEYRSASVLLICAVIMLVMGFRSDGVNIAPIILYTAVFLLLVCLYRCKTAH
PYLMAHWRVFQRQIMFISLKSLRTINKSNFFSNERKYRQLVQEYKKNNRPVPDRKTYFCNGFEWGPEHAD
RAYQIANLSSDKREIALPFVLSPIARHFETMARSMGGNNAIFAVDRRAPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVIVVIDPKNDAEWRQSLMDEASELGLPFYKFHPAQPSSSVCIDVCNAYTNVSD
LTSRLLSLVSVPGEVNPFVQYAEALVSTVITGLSYTDKKPSIYLIHKNMKSHMSVVNLTIKVMECCFARH
YGPDVWMEKVKYASNDTLQVRFKRLTEWFNAHFLNYEGAEPIEWIDTVGRLVDYSMSDPEHMSKMTAGIM
PLFSRLTEQPLNELLSPSPNTLTSREIVTSDGMFSTGGVLYISLDGLSNPESARAISQLIMSDLTSCAGS
RYNADDGDMSSHSRISIFVDEAHSAINNSMINLLAQGRAAQIALFICTQTISDFIAAANAETANRITGLC
NNYISLRVNDTPTQTLVVENFGKSPISTNMVTYTTGSETTLPHNNFSGSISERKQTTLEESIPKELLGQV
PKFHIVARLQDGRKVVGQIPIAVSEKAMKPNTTLLEMFLKPAGKVTLRQNVGLSYLNKYLRKLH
MTKSKRTNLHAQENFYRPILEYRSASVLLICAVIMLVMGFRSDGVNIAPIILYTAVFLLLVCLYRCKTAH
PYLMAHWRVFQRQIMFISLKSLRTINKSNFFSNERKYRQLVQEYKKNNRPVPDRKTYFCNGFEWGPEHAD
RAYQIANLSSDKREIALPFVLSPIARHFETMARSMGGNNAIFAVDRRAPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVIVVIDPKNDAEWRQSLMDEASELGLPFYKFHPAQPSSSVCIDVCNAYTNVSD
LTSRLLSLVSVPGEVNPFVQYAEALVSTVITGLSYTDKKPSIYLIHKNMKSHMSVVNLTIKVMECCFARH
YGPDVWMEKVKYASNDTLQVRFKRLTEWFNAHFLNYEGAEPIEWIDTVGRLVDYSMSDPEHMSKMTAGIM
PLFSRLTEQPLNELLSPSPNTLTSREIVTSDGMFSTGGVLYISLDGLSNPESARAISQLIMSDLTSCAGS
RYNADDGDMSSHSRISIFVDEAHSAINNSMINLLAQGRAAQIALFICTQTISDFIAAANAETANRITGLC
NNYISLRVNDTPTQTLVVENFGKSPISTNMVTYTTGSETTLPHNNFSGSISERKQTTLEESIPKELLGQV
PKFHIVARLQDGRKVVGQIPIAVSEKAMKPNTTLLEMFLKPAGKVTLRQNVGLSYLNKYLRKLH
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 179595..188763
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| LRS40_RS00925 (LRS40_00925) | 175006..176259 | - | 1254 | WP_039026198 | ParA family protein | - |
| LRS40_RS00930 (LRS40_00930) | 179595..182276 | - | 2682 | WP_001255540 | IncHI-type conjugal transfer ATPase TrhC | virb4 |
| LRS40_RS00935 (LRS40_00935) | 182285..183235 | - | 951 | WP_001022266 | TraV family lipoprotein | traV |
| LRS40_RS00940 (LRS40_00940) | 183245..183997 | - | 753 | WP_010892304 | protein-disulfide isomerase HtdT | - |
| LRS40_RS00945 (LRS40_00945) | 184103..184615 | - | 513 | WP_000429933 | hypothetical protein | - |
| LRS40_RS00950 (LRS40_00950) | 184624..185982 | - | 1359 | WP_000351840 | IncHI-type conjugal transfer protein TrhB | traB |
| LRS40_RS00955 (LRS40_00955) | 185972..186412 | - | 441 | WP_001224094 | plasmid transfer protein HtdO | - |
| LRS40_RS00960 (LRS40_00960) | 186414..187646 | - | 1233 | WP_000202110 | type-F conjugative transfer system secretin TraK | traK |
| LRS40_RS00965 (LRS40_00965) | 187649..188434 | - | 786 | WP_000771920 | TraE/TraK family type IV conjugative transfer system protein | traE |
| LRS40_RS00970 (LRS40_00970) | 188446..188763 | - | 318 | WP_000203871 | type IV conjugative transfer system protein TraL | traL |
| LRS40_RS00975 (LRS40_00975) | 188815..189168 | - | 354 | WP_000424120 | pili assembly chaperone | - |
| LRS40_RS00980 (LRS40_00980) | 189731..190075 | + | 345 | WP_153253765 | hypothetical protein | - |
| LRS40_RS00985 (LRS40_00985) | 190522..191397 | - | 876 | WP_001282731 | RepB family plasmid replication initiator protein | - |
| LRS40_RS00990 (LRS40_00990) | 191932..193008 | + | 1077 | WP_001015069 | hypothetical protein | - |
| LRS40_RS00995 (LRS40_00995) | 193026..193226 | + | 201 | WP_000357614 | hypothetical protein | - |
Host bacterium
| ID | 14885 | GenBank | NZ_CP088263 |
| Plasmid name | pG3L-1 | Incompatibility group | IncR |
| Plasmid size | 253593 bp | Coordinate of oriT [Strand] | 252403..252687 [+] |
| Host baterium | Leclercia sp. G3L |
Cargo genes
| Drug resistance gene | ant(3'')-Ia, blaTEM-1B, qnrS1, floR, blaLAP-2, tet(A), dfrA1, qacE, sul1, tet(X4), lnu(G) |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |