Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 114373 |
| Name | oriT_pYps.R2 |
| Organism | Yersinia pseudotuberculosis strain Yps.R2 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_LT221038 (24644..24744 [+], 101 nt) |
| oriT length | 101 nt |
| IRs (inverted repeats) | 80..85, 91..96 (AAAAAA..TTTTTT) 20..26, 38..44 (TAAATCA..TGATTTA) |
| Location of nic site | 62..63 |
| Conserved sequence flanking the nic site |
GGTGTATAGC |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 101 nt
>oriT_pYps.R2
TATTTATTTTTTTATCTTTTAAATCAGTATGATAGCGTGATTTATCGCGCTGCGTTAGGTGTATAGCAGGTTAAGGGATAAAAAATCATCTTTTTTGGTAG
TATTTATTTTTTTATCTTTTAAATCAGTATGATAGCGTGATTTATCGCGCTGCGTTAGGTGTATAGCAGGTTAAGGGATAAAAAATCATCTTTTTTGGTAG
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file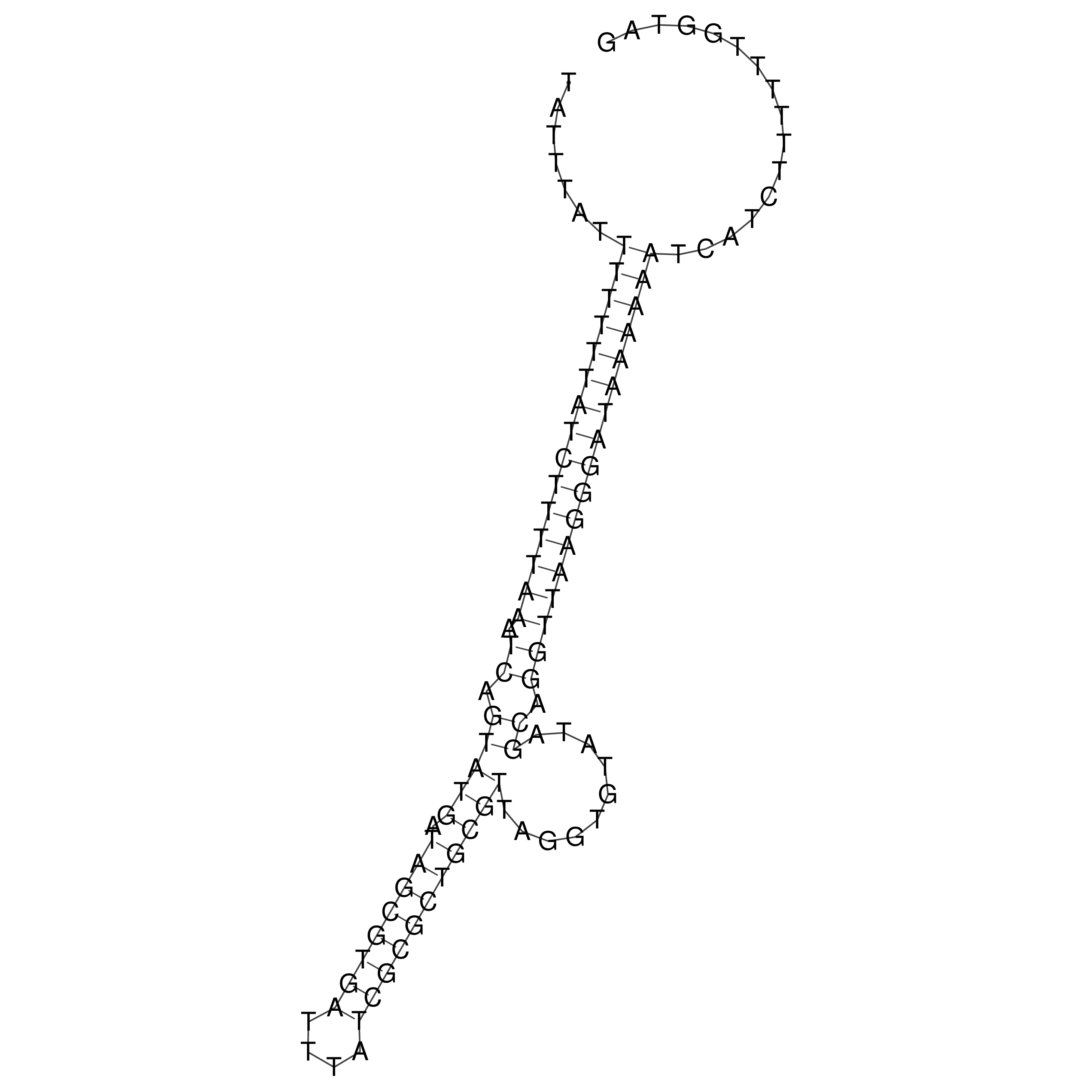
Relaxase
| ID | 9214 | GenBank | WP_012561166 |
| Name | mobF_BN866_RS00090_pYps.R2 |
UniProt ID | D7RU03 |
| Length | 1078 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1078 a.a. Molecular weight: 120141.30 Da Isoelectric Point: 6.5229
>WP_012561166.1 MULTISPECIES: MobF family relaxase [Gammaproteobacteria]
MLDITTITRQNVTSVVGYYSDAKDDYYSKDSSFTSWQGTGAEALGLSGDVESARFKELLVGEIDTFTHMQ
RHVGDAKKERLGYDLTFSAPKGVSMQALIHGDKTIIEAHEKAVAAAVREAEKLAQARTTRQGKSVTQNTN
NLVVATFRHETSRALDPDLHTHAFVMNMTQREDGQWRALKNDELMRNKMHLGDVYKQELALELTKAGYEL
RYNSKNNTFDMAHFSDEQIRAFSRRSEQIEKGLAAMGLTRETADAQTKSRVSMATREKKTEHSREEIHQE
WASRAKTLGIDFDNREWQGHGKPLEADIARNMAPDFTSPEVKADRAIQFAVKSLSERDASFERQKLIQIA
NKQVLGHATMADVEKAYLKAVQKGAIIEGEARYQSTLKVGASVMAETLTRKEWIDSLTNSGMRADKARFA
VDDGIKNGRLKKTSHRVTTVEGIRLERSILTIESRGRGQMPRQLTAEIAGQLLAGKTLKNEQMRAVTEIV
TSKDRFVAAHGYAGTGKSYMTMAAKELLESQGLKVTALAPYGTQKKALEDDGLPARTVAAFLKAKDKKLD
EKSVVFIDEAGVIPARQMKQLMEVIEKHNARAVFLGDTSQTKAVEAGKPFEQLIKAGMQTSYMKDIQRQK
NEVLLEAVKYAAEGNAARALKNITGVNELKEEAPRLAQLADRYLSLSSEQQDATLIISGTNASRKTLNDY
IRGNLGLAGTGETFTLLDRVDSTQAERRDSRYFSKGQIIIPEQDYKNGMKRGESYQVLDTGPGNKLTVES
SSGEQIAFSPRTHTKLSVYQAVSAELAPGDKVMVTRNDKTLDVANGDRFTVKTVEGEKLTLEDKKGRTVE
LDKKQASYLSYAYATTVHKSQGLTCDRVLFNIDTKSLTTSKDVFYVGISRARHEVEIFTDDKKSLASSVS
RDSPKTTAAEIDRFFGLEARFKDIGRDTSLETRSAEKGLPEATGESMAFNQKPDEHNMTTGTDYQPVSNA
EDAFHLKQNPMDDSVGLRRHEAQQNDAELAHDYAAADDQQWSAQEYADYEHYAEASDYDFDSSIYDDYAM
PQTSQAEQSHTGKEHTHEHEHEEGGHEI
MLDITTITRQNVTSVVGYYSDAKDDYYSKDSSFTSWQGTGAEALGLSGDVESARFKELLVGEIDTFTHMQ
RHVGDAKKERLGYDLTFSAPKGVSMQALIHGDKTIIEAHEKAVAAAVREAEKLAQARTTRQGKSVTQNTN
NLVVATFRHETSRALDPDLHTHAFVMNMTQREDGQWRALKNDELMRNKMHLGDVYKQELALELTKAGYEL
RYNSKNNTFDMAHFSDEQIRAFSRRSEQIEKGLAAMGLTRETADAQTKSRVSMATREKKTEHSREEIHQE
WASRAKTLGIDFDNREWQGHGKPLEADIARNMAPDFTSPEVKADRAIQFAVKSLSERDASFERQKLIQIA
NKQVLGHATMADVEKAYLKAVQKGAIIEGEARYQSTLKVGASVMAETLTRKEWIDSLTNSGMRADKARFA
VDDGIKNGRLKKTSHRVTTVEGIRLERSILTIESRGRGQMPRQLTAEIAGQLLAGKTLKNEQMRAVTEIV
TSKDRFVAAHGYAGTGKSYMTMAAKELLESQGLKVTALAPYGTQKKALEDDGLPARTVAAFLKAKDKKLD
EKSVVFIDEAGVIPARQMKQLMEVIEKHNARAVFLGDTSQTKAVEAGKPFEQLIKAGMQTSYMKDIQRQK
NEVLLEAVKYAAEGNAARALKNITGVNELKEEAPRLAQLADRYLSLSSEQQDATLIISGTNASRKTLNDY
IRGNLGLAGTGETFTLLDRVDSTQAERRDSRYFSKGQIIIPEQDYKNGMKRGESYQVLDTGPGNKLTVES
SSGEQIAFSPRTHTKLSVYQAVSAELAPGDKVMVTRNDKTLDVANGDRFTVKTVEGEKLTLEDKKGRTVE
LDKKQASYLSYAYATTVHKSQGLTCDRVLFNIDTKSLTTSKDVFYVGISRARHEVEIFTDDKKSLASSVS
RDSPKTTAAEIDRFFGLEARFKDIGRDTSLETRSAEKGLPEATGESMAFNQKPDEHNMTTGTDYQPVSNA
EDAFHLKQNPMDDSVGLRRHEAQQNDAELAHDYAAADDQQWSAQEYADYEHYAEASDYDFDSSIYDDYAM
PQTSQAEQSHTGKEHTHEHEHEEGGHEI
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | D7RU03 |
Auxiliary protein
| ID | 5400 | GenBank | WP_001749975 |
| Name | WP_001749975_pYps.R2 |
UniProt ID | A0A5C2CVS6 |
| Length | 138 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
Auxiliary protein sequence
Download Length: 138 a.a. Molecular weight: 15332.60 Da Isoelectric Point: 5.9670
>WP_001749975.1 MULTISPECIES: hypothetical protein [Gammaproteobacteria]
MPIITAKVSDELLAYIDLVSGGNRSDYLRRCIEAGPGDRESGLKIVADRLSDVNRKLDYLFDRASDADFG
PLRDELKAITETLSGVKFPPAGQMMLHESLAIETLILLRSIAEPGKTKAAKAEVERNGYKVWEPKKER
MPIITAKVSDELLAYIDLVSGGNRSDYLRRCIEAGPGDRESGLKIVADRLSDVNRKLDYLFDRASDADFG
PLRDELKAITETLSGVKFPPAGQMMLHESLAIETLILLRSIAEPGKTKAAKAEVERNGYKVWEPKKER
Protein domains
No domain identified.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A5C2CVS6 |
T4CP
| ID | 10680 | GenBank | WP_000342688 |
| Name | t4cp2_BN866_RS00095_pYps.R2 |
UniProt ID | _ |
| Length | 509 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 509 a.a. Molecular weight: 57762.87 Da Isoelectric Point: 9.6551
>WP_000342688.1 MULTISPECIES: type IV secretion system DNA-binding domain-containing protein [Enterobacterales]
MDDRERGLAFLFAITLPPVMVWFLVAKFTYGIDPSTAKYLIPYLVKNTFSLWPLWSALIAGWFIGVGGLI
AFIIYDKSRVFKGERFKKIYRGTELVRARTLADKTRERGVNQLTVANIPIPTYAENLHFSIAGTTGTGKT
TIFNELLFKSIIRGGKNIALDPNGGFLKNFYRPGDVILNAYDKRTEGWVFFNEIRRSYDYERLVNSIVQE
SPDMATEEWFGYGRLIFSEVSKKLHSLYSTVTMEEVIHWACNVDQKKLKEFLMGTPAEAIFSGSEKAVGS
ARFVLSKNLAPHLKMPEGNFSLRDWLDDGKPGTLFITWQEEMKRSLNPLISCWLDSIFSIVLGMGEKESR
INVFIDELESLQFLPNLNDALTKGRKSGLCVYAGYQTYSQLVKVYGRDMAQTILANMRSNIVLGGSRLGD
ETLDQMSRSLGEIEGEVERKESDPQKPWIVRKRRDVKVVRAVTPTEISMLPNLTGYLALPGDMPVAKFKA
KHVKYHRKNPVPGIELREI
MDDRERGLAFLFAITLPPVMVWFLVAKFTYGIDPSTAKYLIPYLVKNTFSLWPLWSALIAGWFIGVGGLI
AFIIYDKSRVFKGERFKKIYRGTELVRARTLADKTRERGVNQLTVANIPIPTYAENLHFSIAGTTGTGKT
TIFNELLFKSIIRGGKNIALDPNGGFLKNFYRPGDVILNAYDKRTEGWVFFNEIRRSYDYERLVNSIVQE
SPDMATEEWFGYGRLIFSEVSKKLHSLYSTVTMEEVIHWACNVDQKKLKEFLMGTPAEAIFSGSEKAVGS
ARFVLSKNLAPHLKMPEGNFSLRDWLDDGKPGTLFITWQEEMKRSLNPLISCWLDSIFSIVLGMGEKESR
INVFIDELESLQFLPNLNDALTKGRKSGLCVYAGYQTYSQLVKVYGRDMAQTILANMRSNIVLGGSRLGD
ETLDQMSRSLGEIEGEVERKESDPQKPWIVRKRRDVKVVRAVTPTEISMLPNLTGYLALPGDMPVAKFKA
KHVKYHRKNPVPGIELREI
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 60738..71067
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| BN866_RS00295 | 56930..58039 | + | 1110 | WP_000842134 | S-(hydroxymethyl)glutathione dehydrogenase/class III alcohol dehydrogenase | - |
| BN866_RS00300 | 58061..58483 | + | 423 | Protein_58 | alpha/beta hydrolase-fold protein | - |
| BN866_RS00305 | 58529..59233 | - | 705 | WP_001067855 | IS6-like element IS26 family transposase | - |
| BN866_RS00310 | 59298..59477 | - | 180 | Protein_60 | ArsR/SmtB family transcription factor | - |
| BN866_RS00445 | 59744..60031 | + | 288 | Protein_61 | DUF6710 family protein | - |
| BN866_RS00325 | 60205..60738 | - | 534 | WP_000792636 | phospholipase D family protein | - |
| BN866_RS00330 | 60738..61733 | - | 996 | WP_000128596 | ATPase, T2SS/T4P/T4SS family | virB11 |
| BN866_RS00335 | 61775..62935 | - | 1161 | WP_000101710 | type IV secretion system protein VirB10 | virB10 |
| BN866_RS00340 | 62935..63819 | - | 885 | WP_000735066 | TrbG/VirB9 family P-type conjugative transfer protein | virB9 |
| BN866_RS00345 | 63830..64528 | - | 699 | WP_000646594 | virB8 family protein | virB8 |
| BN866_RS00350 | 64518..64676 | - | 159 | WP_012561180 | hypothetical protein | - |
| BN866_RS00355 | 64747..65787 | - | 1041 | WP_001749958 | type IV secretion system protein | virB6 |
| BN866_RS00360 | 65803..66030 | - | 228 | WP_001749959 | IncN-type entry exclusion lipoprotein EexN | - |
| BN866_RS00365 | 66038..66802 | - | 765 | WP_013149461 | type IV secretion system protein | virB5 |
| BN866_RS00370 | 66769..69369 | - | 2601 | WP_012561149 | VirB4 family type IV secretion/conjugal transfer ATPase | virb4 |
| BN866_RS00375 | 69369..69686 | - | 318 | WP_000496058 | VirB3 family type IV secretion system protein | virB3 |
| BN866_RS00380 | 69737..70030 | - | 294 | WP_001749962 | hypothetical protein | virB2 |
| BN866_RS00385 | 70040..70321 | - | 282 | WP_016338364 | transcriptional repressor KorA | - |
| BN866_RS00390 | 70330..71067 | - | 738 | WP_013279384 | lytic transglycosylase domain-containing protein | virB1 |
| BN866_RS00395 | 71104..71481 | + | 378 | WP_193567606 | H-NS family nucleoid-associated regulatory protein | - |
| BN866_RS00400 | 71497..71841 | + | 345 | WP_012561155 | hypothetical protein | - |
| BN866_RS00405 | 71838..72152 | + | 315 | WP_016338363 | TrbM/KikA/MpfK family conjugal transfer protein | - |
| BN866_RS00410 | 72188..72499 | + | 312 | WP_013279382 | hypothetical protein | - |
| BN866_RS00415 | 72555..73196 | + | 642 | WP_012561158 | restriction endonuclease | - |
| BN866_RS00420 | 73201..73407 | + | 207 | WP_001749967 | hypothetical protein | - |
| BN866_RS00425 | 73418..73690 | - | 273 | Protein_82 | IS1 family transposase | - |
| BN866_RS00430 | 73789..75222 | + | 1434 | WP_001288432 | DNA cytosine methyltransferase | - |
Host bacterium
| ID | 14808 | GenBank | NZ_LT221038 |
| Plasmid name | pYps.R2 | Incompatibility group | IncN |
| Plasmid size | 76709 bp | Coordinate of oriT [Strand] | 24644..24744 [+] |
| Host baterium | Yersinia pseudotuberculosis strain Yps.R2 |
Cargo genes
| Drug resistance gene | aph(3'')-Ib, aph(6)-Id, sul2, tet(D) |
| Virulence gene | - |
| Metal resistance gene | arsR, arsD, arsA, arsB, arsC |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |