Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 113340 |
| Name | oriT_S1_VPH_Chula|unnamed1 |
| Organism | Salmonella enterica strain S1_VPH_Chula |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP121416 (215881..216134 [-], 254 nt) |
| oriT length | 254 nt |
| IRs (inverted repeats) | 152..157, 159..164 (AAAAGT..ACTTTT) 121..129, 134..142 (TTAAGGCTT..AAGCCTTAA) 50..55, 59..64 (AATTTT..AAAATT) |
| Location of nic site | 77..78 |
| Conserved sequence flanking the nic site |
TTTGGTTAAA |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 254 nt
>oriT_S1_VPH_Chula|unnamed1
GGATTTAGGTTTTTTTTTAATCGCTTCACATTTCGTTAGCATGCGAAGAAATTTTGATAAAATTCTGGTCAGTTTGGTTAAAAAGTGTTACAAGTAAGCGGTGTGGTTGAAGGGATAGATTTAAGGCTTATTCAAGCCTTAAGAAAATACTAAAAGTTACTTTTCACCCTACCGAACACCTAACAAAAAATCCATGTTGAAGATTTGAACAATTGTAATGGCGCAAGGACAATCAGCACATGTCAGAATCTGAT
GGATTTAGGTTTTTTTTTAATCGCTTCACATTTCGTTAGCATGCGAAGAAATTTTGATAAAATTCTGGTCAGTTTGGTTAAAAAGTGTTACAAGTAAGCGGTGTGGTTGAAGGGATAGATTTAAGGCTTATTCAAGCCTTAAGAAAATACTAAAAGTTACTTTTCACCCTACCGAACACCTAACAAAAAATCCATGTTGAAGATTTGAACAATTGTAATGGCGCAAGGACAATCAGCACATGTCAGAATCTGAT
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file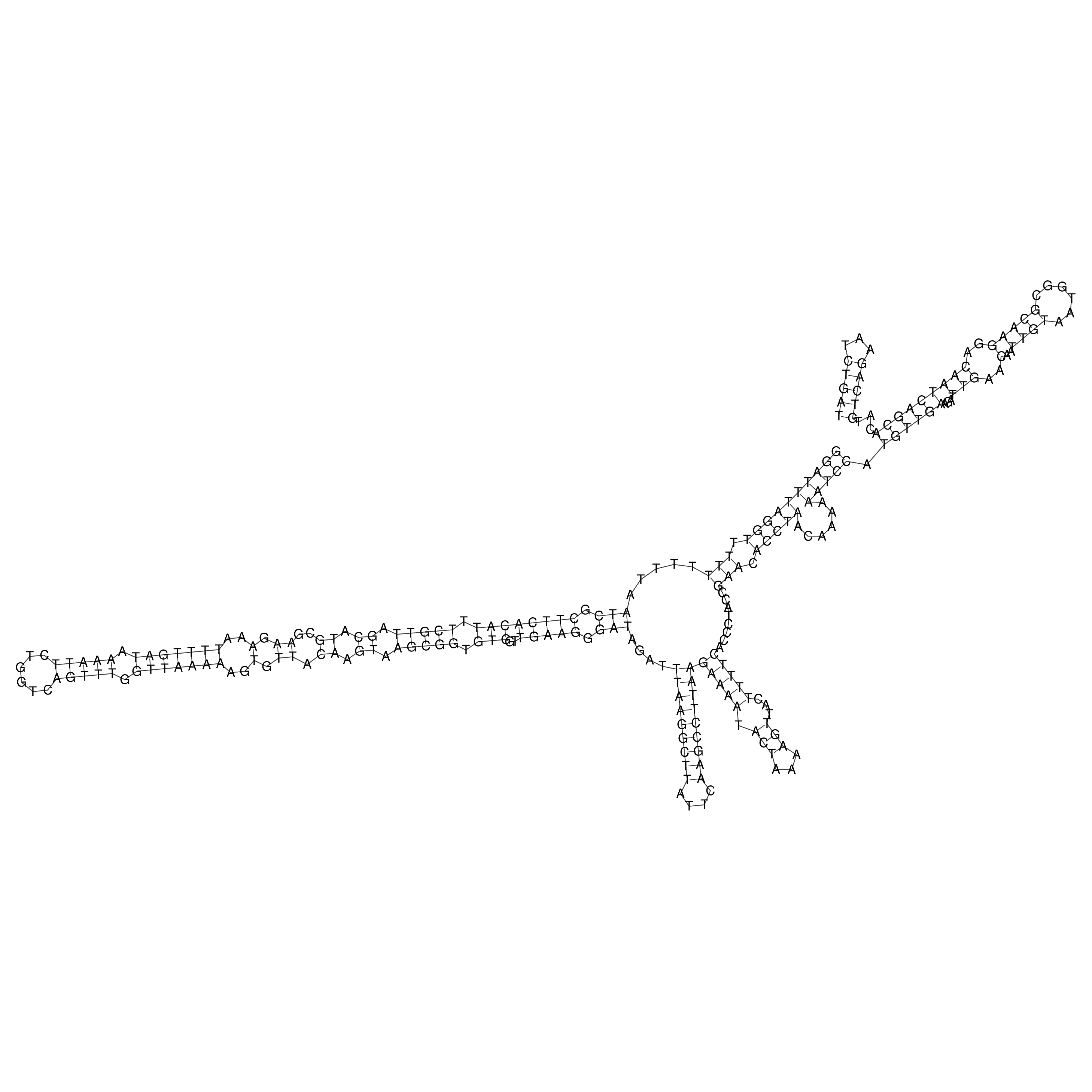
Relaxase
| ID | 8553 | GenBank | WP_000980119 |
| Name | mobH_QAT86_RS24395_S1_VPH_Chula|unnamed1 |
UniProt ID | A0A0H3VVI8 |
| Length | 1050 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1050 a.a. Molecular weight: 118035.42 Da Isoelectric Point: 4.6054
>WP_000980119.1 MULTISPECIES: MobH family relaxase [Enterobacteriaceae]
MMMNFRALYLCIKRILGIFSSQENDATSVMIEDISSLSPFAQILGDQKYTVPDHPNPEVLKFIEYPTRPT
GIQTFNEQSILSLYREKLHSISMMLAISDSDIRDDAYTFTNLVLKPLVEYVRWIHLLPASENHHHNGIGG
LLSHSLEVAILSLKNAHHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLASPIIWT
PSSQSLLDWARENDVVEYEIHWRKRIHNQHNIWSSVFLERILNPVCLAFLDRVNKERVYSKMITALNVYT
DGNDFLSKCVRTADFYSTGTDLNVLRDPIMGLRSNDAAARAISTIKHNFTSININNYNAKPMHIIIVNGE
VYLNENAFLDFVLNDFELHKYNFPQGEAGKTVLVESLVQRGYVEPYDDERVVHYFIPGIYSENEISNIFR
NGIGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRDAITKITDT
VETAVLKVNDLGRSTASIDVDIHSKKNEGSSDDFEKKAESDNEIDNDTQIVKSEGEEAADPVIPDIEESE
DESAKDTESHVLVNQLHELLLSAPLSNDYIVCVDAVPYLNIDATMALLPGLDEKAFSEEPYFQLTFREGS
LDGMWIVRDIDDLRLVQLGDNCAGFQLTYHEPRRPTTLKSLFNTSMYQALVINDESSVENSAPRPKQTLE
LPPPRVNAVEEHSGDVEYHGTDSASATGPLKTEAVEYEHYQHLFEEEDEEHEIIDYTDFTQLSVSRPEVG
SCATSSSVHNEKLLPEPSELPELNREQNADPQGTIERSMDVSVGQENSEPDTEGNCPPPAEVVYSQTEAA
ATSVMASEEPALPPVLEESNGEHAPTDAKGHHLSPALARLFAPTAPVEKQNPKRNRNKSSDKAEVQKPAS
PVSGHNLNSKVFATTESDQNGEFSLISEGDVTELEFVEIASVLHQILSKMEVAFKRKRKNRFMVSTPNTL
YLTQSCVEKFGSQLEAQDLFNKLPQYLVNSGAVINTKCHAFNMPTLLAASDRAKVDIERIINNLKEAGNL
MMMNFRALYLCIKRILGIFSSQENDATSVMIEDISSLSPFAQILGDQKYTVPDHPNPEVLKFIEYPTRPT
GIQTFNEQSILSLYREKLHSISMMLAISDSDIRDDAYTFTNLVLKPLVEYVRWIHLLPASENHHHNGIGG
LLSHSLEVAILSLKNAHHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLASPIIWT
PSSQSLLDWARENDVVEYEIHWRKRIHNQHNIWSSVFLERILNPVCLAFLDRVNKERVYSKMITALNVYT
DGNDFLSKCVRTADFYSTGTDLNVLRDPIMGLRSNDAAARAISTIKHNFTSININNYNAKPMHIIIVNGE
VYLNENAFLDFVLNDFELHKYNFPQGEAGKTVLVESLVQRGYVEPYDDERVVHYFIPGIYSENEISNIFR
NGIGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRDAITKITDT
VETAVLKVNDLGRSTASIDVDIHSKKNEGSSDDFEKKAESDNEIDNDTQIVKSEGEEAADPVIPDIEESE
DESAKDTESHVLVNQLHELLLSAPLSNDYIVCVDAVPYLNIDATMALLPGLDEKAFSEEPYFQLTFREGS
LDGMWIVRDIDDLRLVQLGDNCAGFQLTYHEPRRPTTLKSLFNTSMYQALVINDESSVENSAPRPKQTLE
LPPPRVNAVEEHSGDVEYHGTDSASATGPLKTEAVEYEHYQHLFEEEDEEHEIIDYTDFTQLSVSRPEVG
SCATSSSVHNEKLLPEPSELPELNREQNADPQGTIERSMDVSVGQENSEPDTEGNCPPPAEVVYSQTEAA
ATSVMASEEPALPPVLEESNGEHAPTDAKGHHLSPALARLFAPTAPVEKQNPKRNRNKSSDKAEVQKPAS
PVSGHNLNSKVFATTESDQNGEFSLISEGDVTELEFVEIASVLHQILSKMEVAFKRKRKNRFMVSTPNTL
YLTQSCVEKFGSQLEAQDLFNKLPQYLVNSGAVINTKCHAFNMPTLLAASDRAKVDIERIINNLKEAGNL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A0H3VVI8 |
T4CP
| ID | 9876 | GenBank | WP_001284076 |
| Name | traD_QAT86_RS24400_S1_VPH_Chula|unnamed1 |
UniProt ID | _ |
| Length | 694 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 694 a.a. Molecular weight: 78137.93 Da Isoelectric Point: 8.0177
>WP_001284076.1 MULTISPECIES: conjugative transfer system coupling protein TraD [Enterobacterales]
MSDSKRTNLHAQENFYRPILEYRSASILLICSVSMLYMGLSSDGLDIAPIVLFTSILLFLLCLYRCKTAA
PFLMAHWRVFKRHFMFVSLDSLRVINKSNFFSNERKYRQLVQDYQNKNKDIPERKSYFCDGFEWGPEHAD
RAYQIANLSSDKREIELPFVFNPIKRHFDAMARKMGGSNAIFAVERREPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVVVVIDPKNDAEWRESLMEEAKTLGLPFYKFHPGQPASSVCIDVCNTYTNVSD
LTSRLLSLVTVPGEVNPFVQYAKALVSNVISGLSYIEKKPSIYLIHKNMKSHMSIVNLTVKVMESCYARY
YGYDVWTEKVKYVANETLPVRFKRLAEWFTAHFMNYEGSEQIDWLDTVSQLIDYSMSDPEHMAKMTAGIM
PVFDMLIEKPLNELLSPNPNSVSSREIVTSEGMFSTGGVLYISLDGLSNPDTAAAISQLIMSDLTSCAGS
RYNAQDGDMSANSRISIFVDEAHSAINNPMINLLAQGRAAKIALFICTQTISDFIAAASVETANRITGLC
NNYISLRVNDTPTQTLVVENFGKSAISTNMVTYTTGSETSLPHNNFSGSISERKQTTLEESIPKDLLGQV
PMFHIVARLQDGRKVVGQIPIAVAEKQMKPNTTLSEMLFKKAGKVTLRQNLDIKNLNKFLRKLH
MSDSKRTNLHAQENFYRPILEYRSASILLICSVSMLYMGLSSDGLDIAPIVLFTSILLFLLCLYRCKTAA
PFLMAHWRVFKRHFMFVSLDSLRVINKSNFFSNERKYRQLVQDYQNKNKDIPERKSYFCDGFEWGPEHAD
RAYQIANLSSDKREIELPFVFNPIKRHFDAMARKMGGSNAIFAVERREPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVVVVIDPKNDAEWRESLMEEAKTLGLPFYKFHPGQPASSVCIDVCNTYTNVSD
LTSRLLSLVTVPGEVNPFVQYAKALVSNVISGLSYIEKKPSIYLIHKNMKSHMSIVNLTVKVMESCYARY
YGYDVWTEKVKYVANETLPVRFKRLAEWFTAHFMNYEGSEQIDWLDTVSQLIDYSMSDPEHMAKMTAGIM
PVFDMLIEKPLNELLSPNPNSVSSREIVTSEGMFSTGGVLYISLDGLSNPDTAAAISQLIMSDLTSCAGS
RYNAQDGDMSANSRISIFVDEAHSAINNPMINLLAQGRAAKIALFICTQTISDFIAAASVETANRITGLC
NNYISLRVNDTPTQTLVVENFGKSAISTNMVTYTTGSETSLPHNNFSGSISERKQTTLEESIPKDLLGQV
PMFHIVARLQDGRKVVGQIPIAVAEKQMKPNTTLSEMLFKKAGKVTLRQNLDIKNLNKFLRKLH
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 49028..58235
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| QAT86_RS23485 (QAT86_23490) | 49028..51709 | - | 2682 | WP_000387412 | TraC family protein | virb4 |
| QAT86_RS23490 (QAT86_23495) | 51718..52668 | - | 951 | WP_001022587 | IncHI-type conjugal transfer lipoprotein TrhV | traV |
| QAT86_RS23495 (QAT86_23500) | 52678..53430 | - | 753 | WP_183079335 | protein-disulfide isomerase HtdT | - |
| QAT86_RS23500 (QAT86_23505) | 53549..54031 | - | 483 | WP_000377633 | hypothetical protein | - |
| QAT86_RS23505 (QAT86_23510) | 54039..55394 | - | 1356 | WP_000351841 | IncHI-type conjugal transfer protein TrhB | traB |
| QAT86_RS23510 (QAT86_23515) | 55384..55845 | - | 462 | WP_000521240 | plasmid transfer protein HtdO | - |
| QAT86_RS23515 (QAT86_23520) | 55847..57118 | - | 1272 | WP_000592090 | type-F conjugative transfer system secretin TraK | traK |
| QAT86_RS23520 (QAT86_23525) | 57118..57906 | - | 789 | WP_000783153 | TraE/TraK family type IV conjugative transfer system protein | traE |
| QAT86_RS23525 (QAT86_23530) | 57918..58235 | - | 318 | WP_000043356 | type IV conjugative transfer system protein TraL | traL |
| QAT86_RS23530 (QAT86_23535) | 58286..58639 | - | 354 | WP_000423602 | hypothetical protein | - |
| QAT86_RS23535 (QAT86_23540) | 59788..60054 | - | 267 | Protein_54 | protein RepA | - |
| QAT86_RS23540 (QAT86_23545) | 60147..61169 | + | 1023 | WP_000255946 | IS21-like element IS100 family transposase | - |
| QAT86_RS23545 (QAT86_23550) | 61166..61948 | + | 783 | WP_001317493 | IS21-like element IS100kyp family helper ATPase IstB | - |
| QAT86_RS23550 (QAT86_23555) | 61987..62619 | + | 633 | WP_225519953 | hypothetical protein | - |
Host bacterium
| ID | 13775 | GenBank | NZ_CP121416 |
| Plasmid name | S1_VPH_Chula|unnamed1 | Incompatibility group | IncN |
| Plasmid size | 293619 bp | Coordinate of oriT [Strand] | 215881..216134 [-] |
| Host baterium | Salmonella enterica strain S1_VPH_Chula |
Cargo genes
| Drug resistance gene | tet(X4), floR, tet(A), aph(6)-Id, aph(3'')-Ib, sul1, qacE, aadA2, dfrA12, aac(3)-IId, blaCTX-M-55, blaTEM-1B, tet(M) |
| Virulence gene | - |
| Metal resistance gene | terW, terZ, terA, terB, terC, terD, terE, silE, silS, silR, silC, silF, silB, silA, silP, pcoA, pcoB, pcoC, pcoE |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |