Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 110983 |
| Name | oriT_pVPRE25a |
| Organism | Enterococcus faecalis strain RE25 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP128462 (1233..1282 [-], 50 nt) |
| oriT length | 50 nt |
| IRs (inverted repeats) | _ |
| Location of nic site | 34..35 |
| Conserved sequence flanking the nic site |
GCTTGCAAAA |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 50 nt
>oriT_pVPRE25a
AATATCGCAACATGGTACCATGTTGCTCCGCTTGCAAAAAGAAAGCCTAC
AATATCGCAACATGGTACCATGTTGCTCCGCTTGCAAAAAGAAAGCCTAC
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file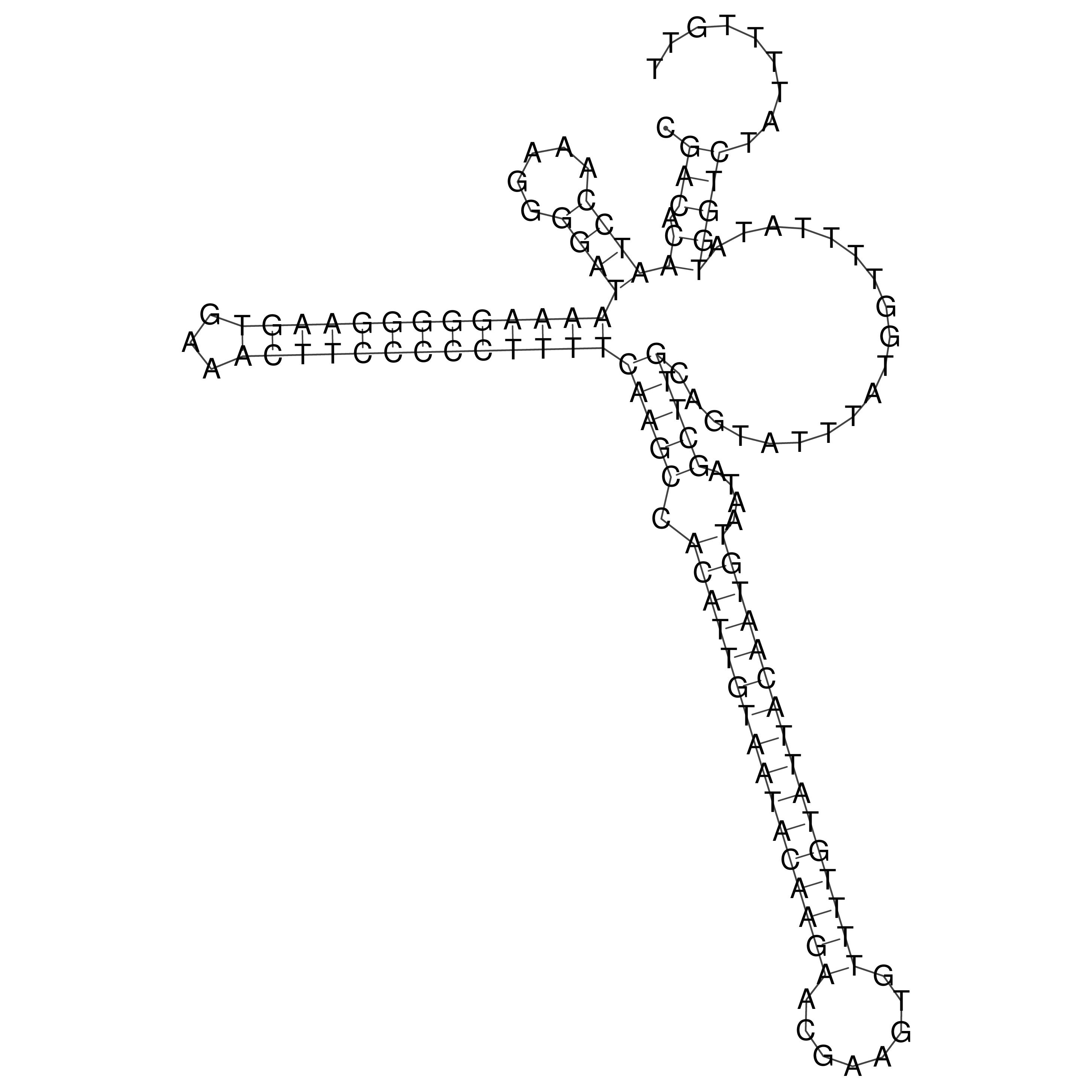
Relaxase
| ID | 7229 | GenBank | WP_289690431 |
| Name | mobV_QSV41_RS00040_pVPRE25a |
UniProt ID | _ |
| Length | 559 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 559 a.a. Molecular weight: 65101.57 Da Isoelectric Point: 8.6009
>WP_289690431.1 MobV family relaxase [Enterococcus faecalis]
MSKIVARMEKMKDGNLSGIQRHNQRETNNHSNPDIDIEKSHLNYDLVNPGSINYREKIKQIIESQRISKR
AVRKDAVLVNEWIITSDTAFFQKNTDTQAFFTDVVAYFSDRCGRQNVAYATVHLDETTPHMHLGIVPMYE
GRLSSKQVFSRQNLLEIQEELPTYLQERGYAIERGLRGSPQKHLSVKEYKDVQKELQVSKQELANQEQIL
AEKERELHSYEQAIFRIDRLLEEKNEEVKGKVQAMNQLETKRNEIINSLQTTVLPDLANVKEKKFSLNGL
KYVLSASDFSAIKSITKDFETLRTQNKHLKIENNNLAQKNKELTQVNHEQELQITELAYELTLKKEQLTA
FEEKQLNYDQSEQIKQELDTTTQTVSVLETENKRLKDKITDLTYDFSELNEKYQEIKLKFDGLTTYLHER
LGHAKETILFTIQQGVKYLQNAPKLKKSLERYSKDRTYLIDEKGTLFLPPDTGSKGVTMSLNPHFYPLKA
PFYNRETGCYEFVGEYMANGRSRTNTLQLSEEQVLEKKAFTFEQSTDKKAWQQSKKYSRDMSKGRGLSR
MSKIVARMEKMKDGNLSGIQRHNQRETNNHSNPDIDIEKSHLNYDLVNPGSINYREKIKQIIESQRISKR
AVRKDAVLVNEWIITSDTAFFQKNTDTQAFFTDVVAYFSDRCGRQNVAYATVHLDETTPHMHLGIVPMYE
GRLSSKQVFSRQNLLEIQEELPTYLQERGYAIERGLRGSPQKHLSVKEYKDVQKELQVSKQELANQEQIL
AEKERELHSYEQAIFRIDRLLEEKNEEVKGKVQAMNQLETKRNEIINSLQTTVLPDLANVKEKKFSLNGL
KYVLSASDFSAIKSITKDFETLRTQNKHLKIENNNLAQKNKELTQVNHEQELQITELAYELTLKKEQLTA
FEEKQLNYDQSEQIKQELDTTTQTVSVLETENKRLKDKITDLTYDFSELNEKYQEIKLKFDGLTTYLHER
LGHAKETILFTIQQGVKYLQNAPKLKKSLERYSKDRTYLIDEKGTLFLPPDTGSKGVTMSLNPHFYPLKA
PFYNRETGCYEFVGEYMANGRSRTNTLQLSEEQVLEKKAFTFEQSTDKKAWQQSKKYSRDMSKGRGLSR
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Host bacterium
| ID | 11418 | GenBank | NZ_CP128462 |
| Plasmid name | pVPRE25a | Incompatibility group | - |
| Plasmid size | 5141 bp | Coordinate of oriT [Strand] | 1233..1282 [-] |
| Host baterium | Enterococcus faecalis strain RE25 |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |