Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 108522 |
| Name | oriT_p1818-b |
| Organism | Enterococcus faecium strain 1818 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP091208 (18219..18256 [+], 38 nt) |
| oriT length | 38 nt |
| IRs (inverted repeats) | 14..22, 29..37 (TAAAGTATA..TATACTTTA) 2..8, 13..19 (ACTTTAT..ATAAAGT) |
| Location of nic site | 27..28 |
| Conserved sequence flanking the nic site |
GTGTGTTATA |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 38 nt
>oriT_p1818-b
CACTTTATGAATATAAAGTATAGTGTGTTATACTTTAC
CACTTTATGAATATAAAGTATAGTGTGTTATACTTTAC
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file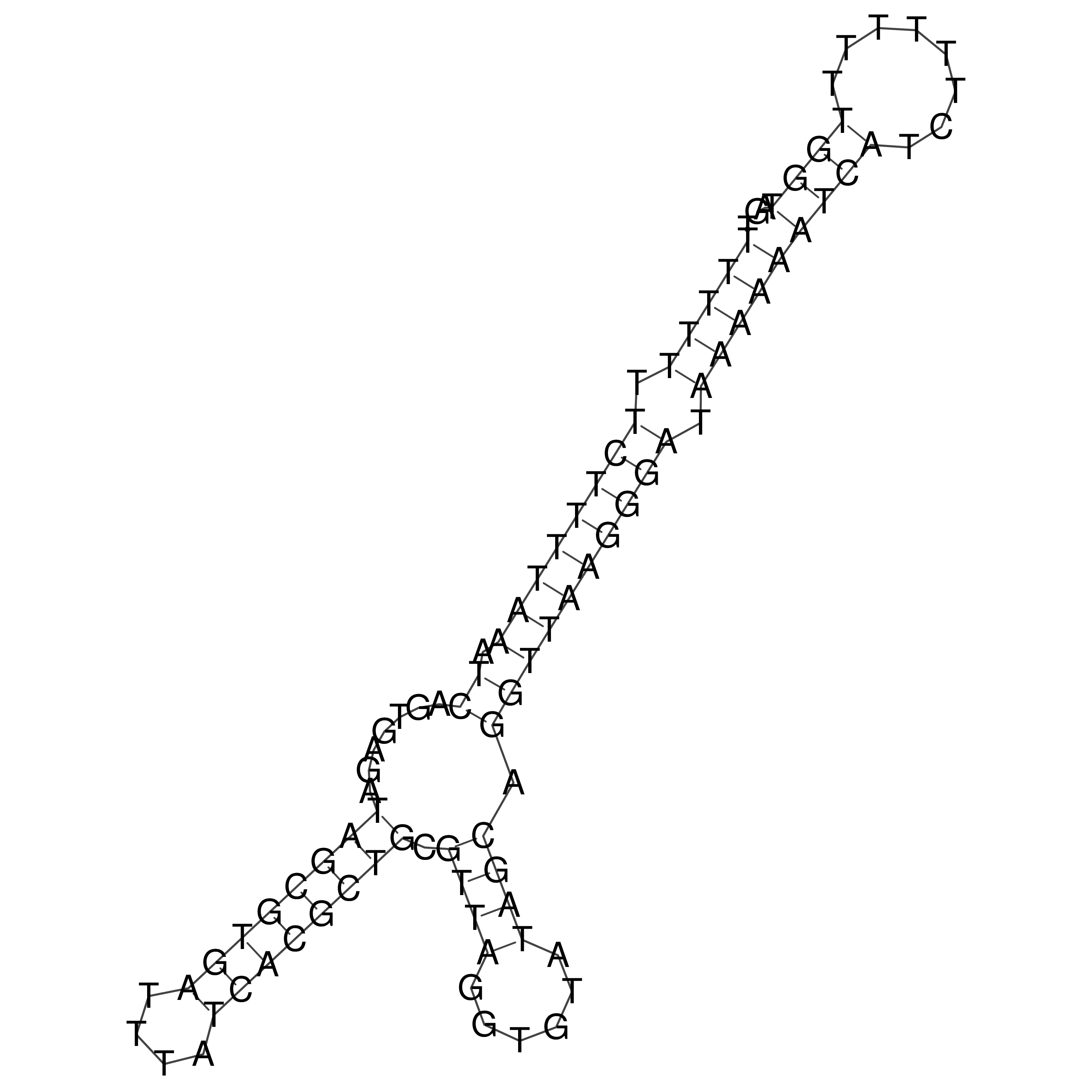
Relaxase
| ID | 5695 | GenBank | WP_000119405 |
| Name | mobV_LQ061_RS12590_p1818-b |
UniProt ID | F5X340 |
| Length | 420 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 420 a.a. Molecular weight: 49660.33 Da Isoelectric Point: 9.9693
>WP_000119405.1 MULTISPECIES: MobV family relaxase [Bacteria]
MSYAVCRMQKVKSAGLKGMQFHNQRERKSRTNDDIDHERTRENYDLKNDKNIDYNERVKEIIESQKTGTR
KTRKDAVLVNELLVTSDRDFFEQLDPGEQKRFFEESYKLFSERYGKQNIAYATVHNDEQTPHMHLGVVPM
RDGKLQGKNVFNRQELLWLQDKFPEHMKKQGFELKRGERGSDRKHIETAKFKKQTLEKEIDFLEKNLAVK
KDEWTAYSDKVKSDLEVPAKRHMKSVEVPTGEKSMFGLGKEIMKTEKKPTKNVVISERDYKNLVTAARDN
DRLKQHVRNLMSTDMAREYKKLSKEHGQVKEKYSGLVERFNENVNDYNELLEENKSLKSKISDLKRDVSL
IYESTKEFLKERTDGLKAFKNVFKGFVDKVKDKTAQFQEKHDLEPKKNEFELTHNREVKKERSRDQGMSL
MSYAVCRMQKVKSAGLKGMQFHNQRERKSRTNDDIDHERTRENYDLKNDKNIDYNERVKEIIESQKTGTR
KTRKDAVLVNELLVTSDRDFFEQLDPGEQKRFFEESYKLFSERYGKQNIAYATVHNDEQTPHMHLGVVPM
RDGKLQGKNVFNRQELLWLQDKFPEHMKKQGFELKRGERGSDRKHIETAKFKKQTLEKEIDFLEKNLAVK
KDEWTAYSDKVKSDLEVPAKRHMKSVEVPTGEKSMFGLGKEIMKTEKKPTKNVVISERDYKNLVTAARDN
DRLKQHVRNLMSTDMAREYKKLSKEHGQVKEKYSGLVERFNENVNDYNELLEENKSLKSKISDLKRDVSL
IYESTKEFLKERTDGLKAFKNVFKGFVDKVKDKTAQFQEKHDLEPKKNEFELTHNREVKKERSRDQGMSL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | F5X340 |
T4CP
| ID | 6072 | GenBank | WP_002325630 |
| Name | t4cp2_LQ061_RS12485_p1818-b |
UniProt ID | _ |
| Length | 551 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 551 a.a. Molecular weight: 63114.14 Da Isoelectric Point: 4.7040
>WP_002325630.1 MULTISPECIES: VirD4-like conjugal transfer protein, CD1115 family [Bacilli]
MSSLLAESDGLILGKFQSGNVVIQPEDSKISNRNIFVVGGPGSFKTQSYVLPNVVNNRSTSIVVTDPKGE
IYELTNEIKKAQGFKTVVINFKDFLLSSRYNPLLYIRKSNDTNKIANVIVSAKNDPKRKDFWFNAQLNLL
NTLIKYVYFEYEPSARTIEGILDFLEEFDPRYNEEGVSELDEQFERLPDGHEAKRSYYLGFRQAQSEARP
NIVISLLTTLQDFVDKEVAEFTSTNDFFFEELGTSKICLYVLISPLDRTWDGLVNLFFQQMFTELYFLGD
KHNAKLPVPLVMLLDEFVNLGYFPTYENFLATCRGYRISVSTILQSLPQGFELYGDKKFKAIIGNHAIKI
CLGGVEETTAEYFSRQVNDTTIKVYTGGTSESKTSAKTTNRSGSKSESYGYQKRRLITEGEVINLQQEEN
GRKSIVLIDGKPYMLRKTPQFELFGNLLKKHEISQQDYISSQTEFAKEYIEELEKQHKQRKVQLASAPIF
HEKKQEDDLNKKEINLPEQPIESEVTEEEEEKELLMSDILSSFDEGIEENTTNNDNSELPF
MSSLLAESDGLILGKFQSGNVVIQPEDSKISNRNIFVVGGPGSFKTQSYVLPNVVNNRSTSIVVTDPKGE
IYELTNEIKKAQGFKTVVINFKDFLLSSRYNPLLYIRKSNDTNKIANVIVSAKNDPKRKDFWFNAQLNLL
NTLIKYVYFEYEPSARTIEGILDFLEEFDPRYNEEGVSELDEQFERLPDGHEAKRSYYLGFRQAQSEARP
NIVISLLTTLQDFVDKEVAEFTSTNDFFFEELGTSKICLYVLISPLDRTWDGLVNLFFQQMFTELYFLGD
KHNAKLPVPLVMLLDEFVNLGYFPTYENFLATCRGYRISVSTILQSLPQGFELYGDKKFKAIIGNHAIKI
CLGGVEETTAEYFSRQVNDTTIKVYTGGTSESKTSAKTTNRSGSKSESYGYQKRRLITEGEVINLQQEEN
GRKSIVLIDGKPYMLRKTPQFELFGNLLKKHEISQQDYISSQTEFAKEYIEELEKQHKQRKVQLASAPIF
HEKKQEDDLNKKEINLPEQPIESEVTEEEEEKELLMSDILSSFDEGIEENTTNNDNSELPF
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Host bacterium
| ID | 8957 | GenBank | NZ_CP091208 |
| Plasmid name | p1818-b | Incompatibility group | - |
| Plasmid size | 36787 bp | Coordinate of oriT [Strand] | 18219..18256 [+] |
| Host baterium | Enterococcus faecium strain 1818 |
Cargo genes
| Drug resistance gene | tet(S), tet(L) |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |