Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 108474 |
| Name | oriT_pSAU071 |
| Organism | Staphylococcus aureus strain Guangzhou-SAU071 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP053184 (20561..20800 [-], 240 nt) |
| oriT length | 240 nt |
| IRs (inverted repeats) | 217..222, 232..237 (ATTTTA..TAAAAT) 171..178, 183..190 (CTATCATT..AATGATAG) 154..160, 164..170 (GTCTGGC..GCCAGAC) 26..32, 44..50 (TTTTTTA..TAAAAAA) 37..42, 45..50 (TTTTTT..AAAAAA) 1..7, 20..26 (AAGACAT..ATGTCTT) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 240 nt
>oriT_pSAU071
AAGACATTAGTGATAACTGATGTCTTTTTTTATTGATTTTTTGTAAAAAAGTACTGTCTTAGTTTTGTGACAAATACTGTATATAGTGTCACAAAAGTGTGACACTACAGCTTTTTATGATATCACTTTAAAATTACTTAAAACCCTTGGAATGTCTGGCTTTGCCAGACCTATCATTTTTGAATGATAGCAAATTCCCCTTATGCTCTTACGGAGATTTTAGAGTTAAATTAAAATTTC
AAGACATTAGTGATAACTGATGTCTTTTTTTATTGATTTTTTGTAAAAAAGTACTGTCTTAGTTTTGTGACAAATACTGTATATAGTGTCACAAAAGTGTGACACTACAGCTTTTTATGATATCACTTTAAAATTACTTAAAACCCTTGGAATGTCTGGCTTTGCCAGACCTATCATTTTTGAATGATAGCAAATTCCCCTTATGCTCTTACGGAGATTTTAGAGTTAAATTAAAATTTC
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file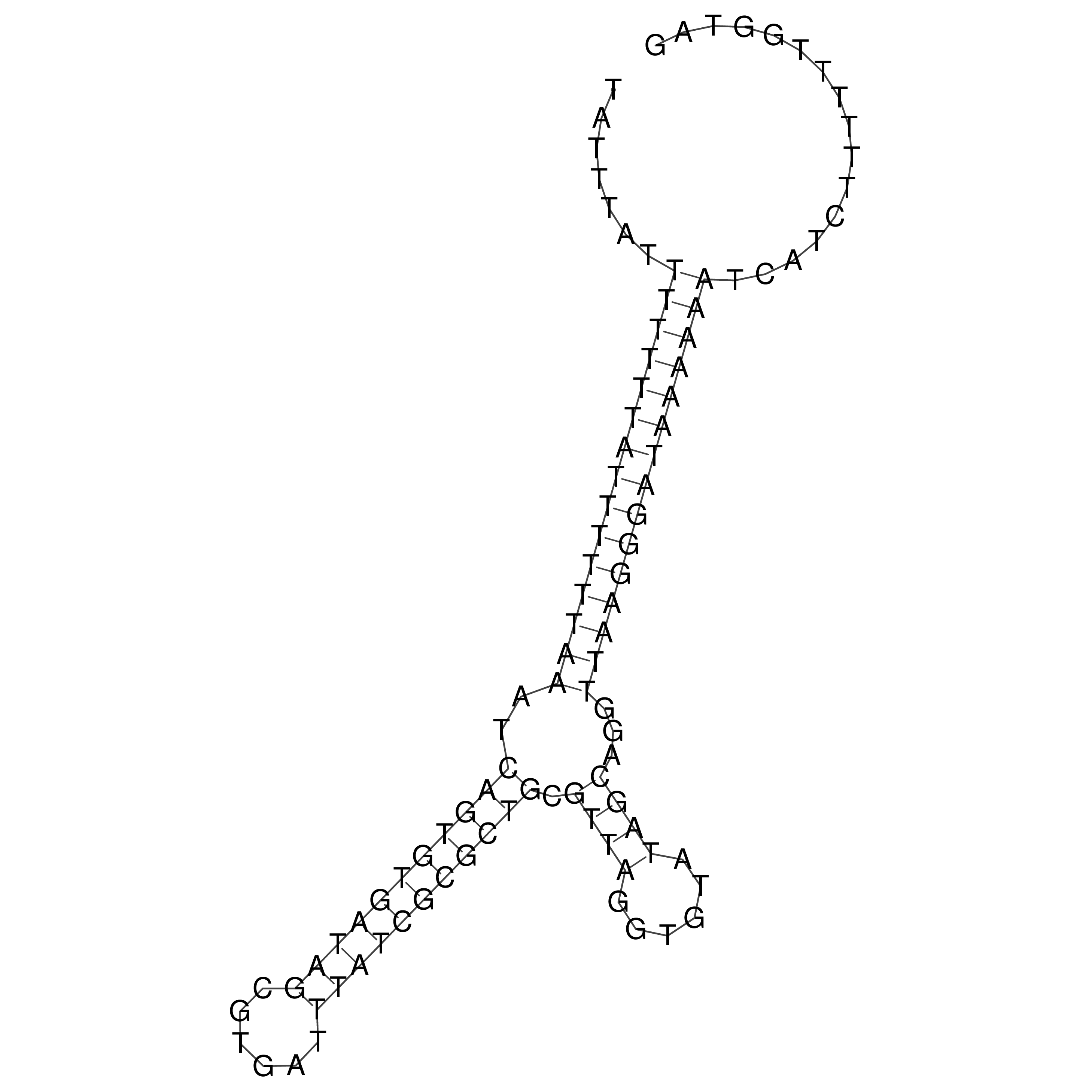
Relaxase
| ID | 5669 | GenBank | WP_000122374 |
| Name | mobV_HLB24_RS13750_pSAU071 |
UniProt ID | A0A2R4A9A7 |
| Length | 413 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 413 a.a. Molecular weight: 48526.00 Da Isoelectric Point: 6.1679
>WP_000122374.1 MULTISPECIES: MobV family relaxase [Bacteria]
MSYSIVRVSKVKSGTNTTGIQKHVQRENNNYENEDIDHSKTYLNYDLVNANKQNFNNLIDEKIEQNYTGK
RKIRTDAIKHIDGLITSDNDFFDNQTPEDTKQFFEYAKEFLEQEYGKDNLLYATVHMDEKTPHMHYGVVP
ITDDGRLSAKEVVGNKKALTAFQDRFNEHVKQRGYDLERGQSRQVTNAKHEQISQYKQKTEYHKQEYERE
SQKTDHIKQKNDKLMQEYQKSLNTLKKPINVPYEQETEKVGGLFSKEIQETGNVVISQKDFNEFQKQIKA
AQDISEDYEYIKSGRALDDKDKEIREKDDLLNKAVERIENADDNFNQLYENAKPLKENIEIALKLLKILL
KELERVLGRNTFAERVNKLTEDEPKLNGLAGNLDKKMNPELYSEQEQQQEQQKNQKRDRGMHL
MSYSIVRVSKVKSGTNTTGIQKHVQRENNNYENEDIDHSKTYLNYDLVNANKQNFNNLIDEKIEQNYTGK
RKIRTDAIKHIDGLITSDNDFFDNQTPEDTKQFFEYAKEFLEQEYGKDNLLYATVHMDEKTPHMHYGVVP
ITDDGRLSAKEVVGNKKALTAFQDRFNEHVKQRGYDLERGQSRQVTNAKHEQISQYKQKTEYHKQEYERE
SQKTDHIKQKNDKLMQEYQKSLNTLKKPINVPYEQETEKVGGLFSKEIQETGNVVISQKDFNEFQKQIKA
AQDISEDYEYIKSGRALDDKDKEIREKDDLLNKAVERIENADDNFNQLYENAKPLKENIEIALKLLKILL
KELERVLGRNTFAERVNKLTEDEPKLNGLAGNLDKKMNPELYSEQEQQQEQQKNQKRDRGMHL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A2R4A9A7 |
Host bacterium
| ID | 8909 | GenBank | NZ_CP053184 |
| Plasmid name | pSAU071 | Incompatibility group | - |
| Plasmid size | 25960 bp | Coordinate of oriT [Strand] | 20561..20800 [-] |
| Host baterium | Staphylococcus aureus strain Guangzhou-SAU071 |
Cargo genes
| Drug resistance gene | tet(K), blaZ |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | AcrIIA21 |