Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 107638 |
| Name | oriT_pRL4b6 |
| Organism | Rhizobium leguminosarum bv. trifolii strain 4B |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP050105 (355440..355468 [+], 29 nt) |
| oriT length | 29 nt |
| IRs (inverted repeats) | 14..19, 24..29 (CGTCGC..GCGACG) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 29 nt
>oriT_pRL4b6
AGGGCGCAATATACGTCGCTGTCGCGACG
AGGGCGCAATATACGTCGCTGTCGCGACG
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file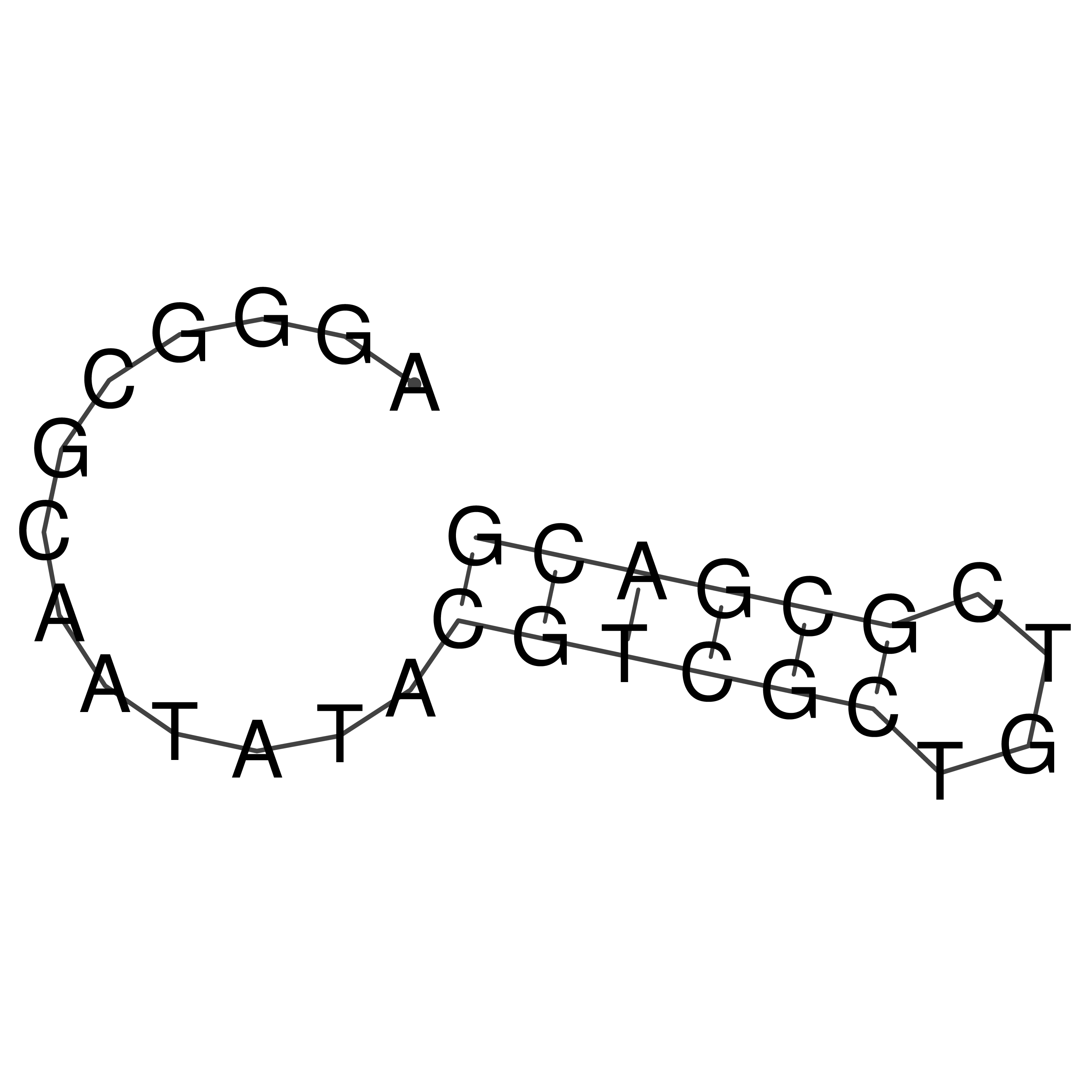
Relaxase
| ID | 5132 | GenBank | WP_166465927 |
| Name | traA_HA460_RS31090_pRL4b6 |
UniProt ID | _ |
| Length | 1538 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1538 a.a. Molecular weight: 169692.64 Da Isoelectric Point: 7.2399
>WP_166465927.1 Ti-type conjugative transfer relaxase TraA [Rhizobium leguminosarum]
MAIMFVRAQVIGRGAGRSVVSAAAYRHRARMMDEQAGTSFSYRGGASELVHEELALPDQIPAWLRSAISG
QSVAKASEALWNAVDAFEKRADAQLARELIIALPEELTRAENIALVREFVLDNLTAKGMVADWVYHDKDG
NPHIHLMTTLRPLAEEGFGPKKVPVCGEDGKPLRVVTPDRPNGKVVYKLWAGDKETLQAWKIAWAETASR
HLALAGHEIRLDGRSYAEQGLDGIAQKHLGPEKAALSRRDAEIYFAPADLARRQEMADRLLADPALLLKQ
LSNERSTFDERDIAKALHRTVDDPADFANIRARLMASDDLIMLKPQQIDAETGKASEPAVFTTREMLRIE
YDMGRSARVLAERRGFAVSDRVVAAAIRQVETQDPENSFRLDPEQVDAIHHITRDDGIAAVVGLAGAGKS
TLLAAARVAWESDGRRVIGAALAGKAAEGLEDSSGIRSRTLAAWELAWASGRELLERGNVLVIDEAGMVS
SQQMARLLDIAEQAEVKVVLVGDAMQLQPIQAGAAFRAITERIGFAELAGVRRQRQPWAREASRLFARGE
TGKGLDAYAQQGHLIEAGTRVETIDRIVADWSQARKQAIQNSTTAGNDGRLRGDELLVLAHTNEDVKYLN
EALRSVMVGEGALGEDRAFRTERGARAFAAGDRIIFLENARFVEPRATRLGPQHVKNGMLGTVVSTGDKR
GDAVLSIRLDSGRDVVISESSYRNVDHGYAATIHKSQGATVERTFVLATGMMDQHLTYVSMTRHRDRVDL
YAAKDDFEAKPEWGRKPHVDHAAGVTGALVETGQAKFRPNDEDADDSPYADVRTDDGTLHRLWGVSLPKA
LEEGGVSEGDTVTLRKDGVERVTIKVAIVDEQTGQKRFEEREVDRNVWTAKRIETAEARQQRIERESHRP
DVFKQLVERLSRSGAKTTTLDFESEAGYRAHADDFARRRGLDHLSLAAAGAEESLSRRWAGIAEKREQLA
KLWERASVALGFTIERERRIAYNEERAEPQTVLEADVARGRYLIPPTTGFARSAEEDAQLAQLSSPAWKE
REAILRPLLEKIYRDPDAALVALNALSSDTSVDPRRLADDLAKAPERLGRLRGSELIVDGRAARDERNAA
ITALTDLLPLVRAHATRFRRNAERFESHEQQRRAHMALSIPALSEKAMARLTEIEAVRRQGGDDAYKAAF
ALAAEDRSIVQEVKAVSEALTARFGWSAFTSKADAVGERNMVERMPEDLASSRGEKLTRLFKAVKRFADE
QHLAERRDRSKIVAGASADQGMENVPMLAAVTEFRTAIDEEARSRVLSDPLYRQQRAALAAVASAIWRDP
VGAVGKIEELLQKGFAGERIAAAVTNDPAAYGALRGSDRLMDRMLAAGRERKQAVQAVPEAGARLRSLGS
AYVNVLDAERQAIAEERRRMAVAIPALSTAAEDVLLRLTAEAKNNDRKLTASAAPLDPSIRREFAAVSRA
LDERFGRNAIIRGEKDLINRVPQAQRSAFEAMQDRLKVLQRTVRMESSEQIISERRQRAVSRGRGIEL
MAIMFVRAQVIGRGAGRSVVSAAAYRHRARMMDEQAGTSFSYRGGASELVHEELALPDQIPAWLRSAISG
QSVAKASEALWNAVDAFEKRADAQLARELIIALPEELTRAENIALVREFVLDNLTAKGMVADWVYHDKDG
NPHIHLMTTLRPLAEEGFGPKKVPVCGEDGKPLRVVTPDRPNGKVVYKLWAGDKETLQAWKIAWAETASR
HLALAGHEIRLDGRSYAEQGLDGIAQKHLGPEKAALSRRDAEIYFAPADLARRQEMADRLLADPALLLKQ
LSNERSTFDERDIAKALHRTVDDPADFANIRARLMASDDLIMLKPQQIDAETGKASEPAVFTTREMLRIE
YDMGRSARVLAERRGFAVSDRVVAAAIRQVETQDPENSFRLDPEQVDAIHHITRDDGIAAVVGLAGAGKS
TLLAAARVAWESDGRRVIGAALAGKAAEGLEDSSGIRSRTLAAWELAWASGRELLERGNVLVIDEAGMVS
SQQMARLLDIAEQAEVKVVLVGDAMQLQPIQAGAAFRAITERIGFAELAGVRRQRQPWAREASRLFARGE
TGKGLDAYAQQGHLIEAGTRVETIDRIVADWSQARKQAIQNSTTAGNDGRLRGDELLVLAHTNEDVKYLN
EALRSVMVGEGALGEDRAFRTERGARAFAAGDRIIFLENARFVEPRATRLGPQHVKNGMLGTVVSTGDKR
GDAVLSIRLDSGRDVVISESSYRNVDHGYAATIHKSQGATVERTFVLATGMMDQHLTYVSMTRHRDRVDL
YAAKDDFEAKPEWGRKPHVDHAAGVTGALVETGQAKFRPNDEDADDSPYADVRTDDGTLHRLWGVSLPKA
LEEGGVSEGDTVTLRKDGVERVTIKVAIVDEQTGQKRFEEREVDRNVWTAKRIETAEARQQRIERESHRP
DVFKQLVERLSRSGAKTTTLDFESEAGYRAHADDFARRRGLDHLSLAAAGAEESLSRRWAGIAEKREQLA
KLWERASVALGFTIERERRIAYNEERAEPQTVLEADVARGRYLIPPTTGFARSAEEDAQLAQLSSPAWKE
REAILRPLLEKIYRDPDAALVALNALSSDTSVDPRRLADDLAKAPERLGRLRGSELIVDGRAARDERNAA
ITALTDLLPLVRAHATRFRRNAERFESHEQQRRAHMALSIPALSEKAMARLTEIEAVRRQGGDDAYKAAF
ALAAEDRSIVQEVKAVSEALTARFGWSAFTSKADAVGERNMVERMPEDLASSRGEKLTRLFKAVKRFADE
QHLAERRDRSKIVAGASADQGMENVPMLAAVTEFRTAIDEEARSRVLSDPLYRQQRAALAAVASAIWRDP
VGAVGKIEELLQKGFAGERIAAAVTNDPAAYGALRGSDRLMDRMLAAGRERKQAVQAVPEAGARLRSLGS
AYVNVLDAERQAIAEERRRMAVAIPALSTAAEDVLLRLTAEAKNNDRKLTASAAPLDPSIRREFAAVSRA
LDERFGRNAIIRGEKDLINRVPQAQRSAFEAMQDRLKVLQRTVRMESSEQIISERRQRAVSRGRGIEL
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Host bacterium
| ID | 8073 | GenBank | NZ_CP050105 |
| Plasmid name | pRL4b6 | Incompatibility group | - |
| Plasmid size | 693075 bp | Coordinate of oriT [Strand] | 355440..355468 [+] |
| Host baterium | Rhizobium leguminosarum bv. trifolii strain 4B |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | actP |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | AcrIIA9 |