Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 107601 |
| Name | oriT_pRLN1 |
| Organism | Rhizobium leguminosarum strain Norway |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP025013 (702400..702428 [+], 29 nt) |
| oriT length | 29 nt |
| IRs (inverted repeats) | 14..19, 24..29 (CGTCGC..GCGACG) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 29 nt
>oriT_pRLN1
AGGGCGCAATATACGTCGCTGTCGCGACG
AGGGCGCAATATACGTCGCTGTCGCGACG
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file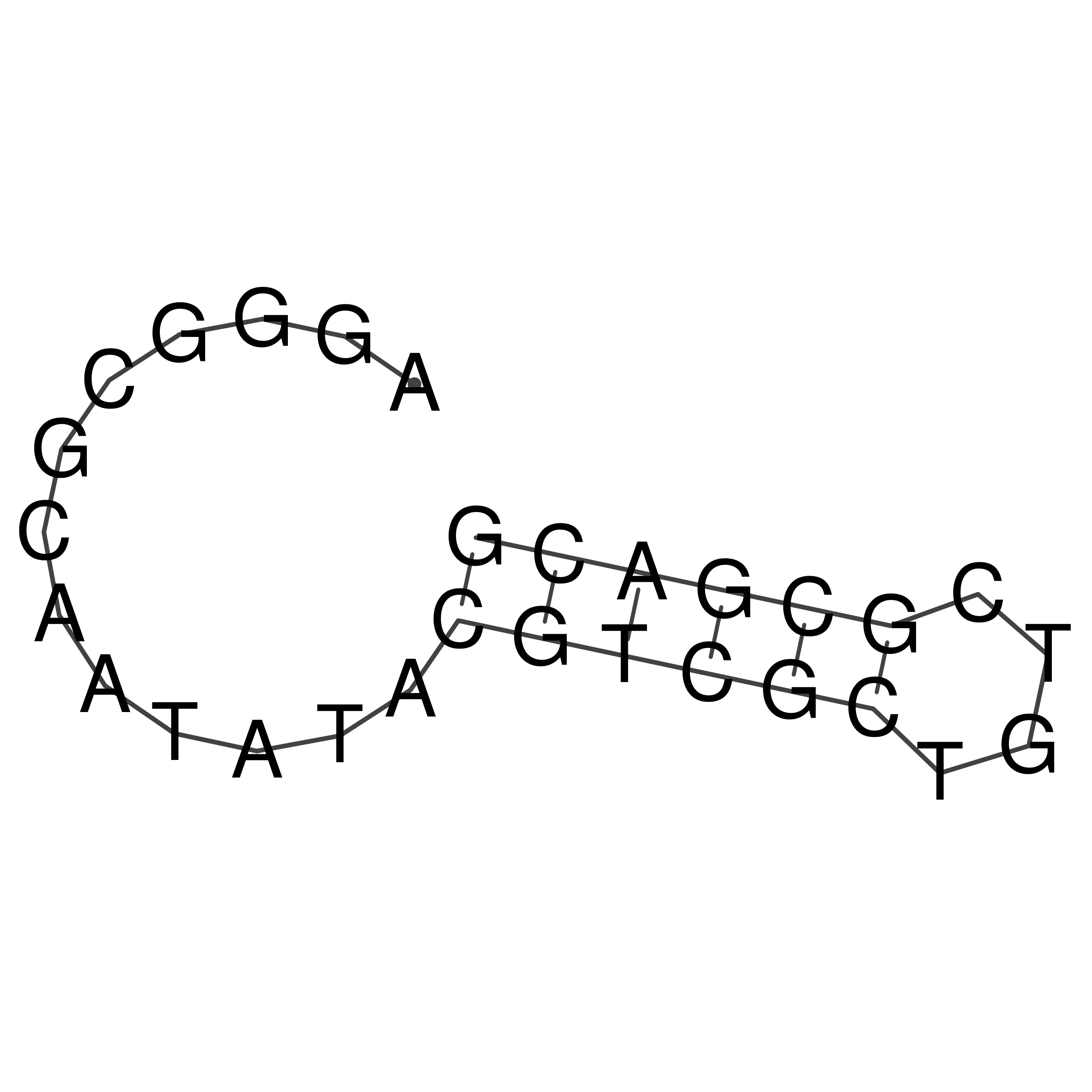
Relaxase
| ID | 5110 | GenBank | WP_105008846 |
| Name | traA_CUJ84_RS27785_pRLN1 |
UniProt ID | _ |
| Length | 1543 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1543 a.a. Molecular weight: 170626.73 Da Isoelectric Point: 7.0475
>WP_105008846.1 Ti-type conjugative transfer relaxase TraA [Rhizobium leguminosarum]
MAIMFVRAQVISRGAGRSIVSAAAYRHRTRMMDEQAGTSFSYRGGASELVHEELALPDQMPTWLRTAIDG
RSVAGASEALWNAVDAFEKRADAQLARELIIALPEELTRAENIALVREFVRDNLTSNGMVADWVYHDRDG
NPHIHLMTTLRPLTEEGFGQKKVAVTGDDGAPLRVVTPDRPKGRIVYRLWAGDKETMKAWKIAWAETANR
HLALAGHEIRLDGRSYAEQGLDGIAQTHLGPEKAALVRKGRELHFAPADLARRQHMADLLLADPALLLKQ
LSNERSTFDERDIAKALHRYVDDPAVFANIRARLMASDELVMLKPQQIDAETGKASEPAVFTTRAILRIE
YDMAHSAQLLAERRGFAVSDRVVAAAIRNVEARDPEKPFKLDPEQVDAVRHVTGDNGIAAVVGLAGAGKS
TLLAAARVAWESDGRRVIGAALAGKAAEGLEDSSGIRSRTLASWELAWANGRELLERGNVLVIDEAGMVS
SQQLARVLKIAEQAQAKVVLVGDAMQLQPIQAGAAFRAITERIGFAELAGVRRQHQQWAREASRLFARGE
VEQGLDVYARHGHLVEAGTRAETIDRIVADWSQARKLAIETSVREGNDGRLRGDELLVLAHTNQDVKRLN
EALRSVMTGEGALSETRAFQTERGAREFAVGDRIIFLENARFLEPRAPRLGPQHVKNGMLGTVVSTGDKR
GDTLLSVRLDNGRDVVISEDSYRNVDHGYAATIHKSQGATVERTFVLATGMMDQHLTYVSMTRHRDRADL
YAAKEDFEAKPEWGRKPHVDHAAGVTGALVETGEAKFRPNDEDADDSPYADVGTDDGTIHRLWGVSLPKA
LEDAGVSEGDTVTLRKDGVERVTIKVATVDEQTGQKRFEERDVERNVWTAKQIETAEERQERIEQESHRP
DLFRQLVERLSRSGAKTTTLDFESEAGYRAHADDFARRRGLDHLSLATAGVEGSLSRRWAGIAERREQLA
KLWERAGVAFGFTIERERQVAYSEERAETQTVLDADVVRGRYLIPPVTVFARSVEEDARLAQLASPAWKE
REAILRPLLETIYRDPDAALVALNALSSDTSIEPRRLADDLAKAPERLGRLRGSDLIVDGRAARDERNAA
ITALTDLLPLARAHATEFRRNSERFDLGEQQRRTHMSLPIPALSEKAIARLTEIEAVRRQGGDDAYKTAF
AIATEDRSLVQEVKAVSEALTARFGWSAFTSKADAVAERNMIERMPEDLASIRGEKLTRLFEAVKRFADE
QHLVERRDRSKIVAAASADQGMEADKESAAVLPMLAAVTEFKTPVDEEARSRVLSDPLYRQQRAALATVA
STIWRDPAGAVGKIEKLLQKSFAGERIATAVTNDPAAYGALRGSDRLMDRLLASGRERRGAVQAVPEAGA
RLRALGSAYVNVLAAERQAVVEERRRMAVAIPGLSKAAEDVLIRLTVEVKNNGRKLSASAASLDHSIRRE
FAAVSTALDERFGRNAVIRGEKDLINRIPPAQRRAFEAMQERLKVLQQTVRMESSEKIISERRQRTTNRS
IDL
MAIMFVRAQVISRGAGRSIVSAAAYRHRTRMMDEQAGTSFSYRGGASELVHEELALPDQMPTWLRTAIDG
RSVAGASEALWNAVDAFEKRADAQLARELIIALPEELTRAENIALVREFVRDNLTSNGMVADWVYHDRDG
NPHIHLMTTLRPLTEEGFGQKKVAVTGDDGAPLRVVTPDRPKGRIVYRLWAGDKETMKAWKIAWAETANR
HLALAGHEIRLDGRSYAEQGLDGIAQTHLGPEKAALVRKGRELHFAPADLARRQHMADLLLADPALLLKQ
LSNERSTFDERDIAKALHRYVDDPAVFANIRARLMASDELVMLKPQQIDAETGKASEPAVFTTRAILRIE
YDMAHSAQLLAERRGFAVSDRVVAAAIRNVEARDPEKPFKLDPEQVDAVRHVTGDNGIAAVVGLAGAGKS
TLLAAARVAWESDGRRVIGAALAGKAAEGLEDSSGIRSRTLASWELAWANGRELLERGNVLVIDEAGMVS
SQQLARVLKIAEQAQAKVVLVGDAMQLQPIQAGAAFRAITERIGFAELAGVRRQHQQWAREASRLFARGE
VEQGLDVYARHGHLVEAGTRAETIDRIVADWSQARKLAIETSVREGNDGRLRGDELLVLAHTNQDVKRLN
EALRSVMTGEGALSETRAFQTERGAREFAVGDRIIFLENARFLEPRAPRLGPQHVKNGMLGTVVSTGDKR
GDTLLSVRLDNGRDVVISEDSYRNVDHGYAATIHKSQGATVERTFVLATGMMDQHLTYVSMTRHRDRADL
YAAKEDFEAKPEWGRKPHVDHAAGVTGALVETGEAKFRPNDEDADDSPYADVGTDDGTIHRLWGVSLPKA
LEDAGVSEGDTVTLRKDGVERVTIKVATVDEQTGQKRFEERDVERNVWTAKQIETAEERQERIEQESHRP
DLFRQLVERLSRSGAKTTTLDFESEAGYRAHADDFARRRGLDHLSLATAGVEGSLSRRWAGIAERREQLA
KLWERAGVAFGFTIERERQVAYSEERAETQTVLDADVVRGRYLIPPVTVFARSVEEDARLAQLASPAWKE
REAILRPLLETIYRDPDAALVALNALSSDTSIEPRRLADDLAKAPERLGRLRGSDLIVDGRAARDERNAA
ITALTDLLPLARAHATEFRRNSERFDLGEQQRRTHMSLPIPALSEKAIARLTEIEAVRRQGGDDAYKTAF
AIATEDRSLVQEVKAVSEALTARFGWSAFTSKADAVAERNMIERMPEDLASIRGEKLTRLFEAVKRFADE
QHLVERRDRSKIVAAASADQGMEADKESAAVLPMLAAVTEFKTPVDEEARSRVLSDPLYRQQRAALATVA
STIWRDPAGAVGKIEKLLQKSFAGERIATAVTNDPAAYGALRGSDRLMDRLLASGRERRGAVQAVPEAGA
RLRALGSAYVNVLAAERQAVVEERRRMAVAIPGLSKAAEDVLIRLTVEVKNNGRKLSASAASLDHSIRRE
FAAVSTALDERFGRNAVIRGEKDLINRIPPAQRRAFEAMQERLKVLQQTVRMESSEKIISERRQRTTNRS
IDL
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4CP
| ID | 5334 | GenBank | WP_105008843 |
| Name | traG_CUJ84_RS27770_pRLN1 |
UniProt ID | _ |
| Length | 637 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 637 a.a. Molecular weight: 70419.20 Da Isoelectric Point: 9.2560
>WP_105008843.1 Ti-type conjugative transfer system protein TraG [Rhizobium leguminosarum]
MALKSRSHPSLLLVFVPITVTAITVYITGWRWTALAHGLSGKTEYWFLRAAPVPALLFGPLAGLLTVWAL
PLHRRRPIAIASLLCFLVVAGFYALREYGRLVPSVERGVVTWNQALSYLDLIAVIGALAGFMGVAVAARI
SVVVPEAVKRARGGTFGDADWLSMAAAAKLFPPDGEIVVGERYRVDKETVHQLPFDPKDRSTWGQGGQAP
LLTYSQDFDSTHMLFFAGSGGYKTTSNVLPTALRYTGPLICLDPSTEVAPMVVEHRTRILGREVMVLDPT
NPIMGFNVLDGIEHSKRKEEDIVGIAHMLLSESLRFESSTGSYFQNQAHNLLTGLLAHVMLSPEYAGRRN
LRSLRQIVSEPEPSVLAMLRDIQEHSETVFIRETLGVFTNMTEQTFSGVYSTASKDTQWLSLDSYAALVC
GNAFRSSDIVTAKTDVFLNIPASILRSYPGIGRVIIGSLINAMVQADGGFSRRALFMLDEVDLLGYMRVL
EEARDRGRKYGISMMLLYQSVGQLERHFGKDGATSWIDGCAFASYAAIKALDTARNVSSQCGEMTVEVQG
RSRNIGWDTKGGGTRKSESVNFQRRPLIMPHEITQSMRKDEQIIIVQGHSPIRCGRAIYFRRKEMDAQAK
ANRFVRT
MALKSRSHPSLLLVFVPITVTAITVYITGWRWTALAHGLSGKTEYWFLRAAPVPALLFGPLAGLLTVWAL
PLHRRRPIAIASLLCFLVVAGFYALREYGRLVPSVERGVVTWNQALSYLDLIAVIGALAGFMGVAVAARI
SVVVPEAVKRARGGTFGDADWLSMAAAAKLFPPDGEIVVGERYRVDKETVHQLPFDPKDRSTWGQGGQAP
LLTYSQDFDSTHMLFFAGSGGYKTTSNVLPTALRYTGPLICLDPSTEVAPMVVEHRTRILGREVMVLDPT
NPIMGFNVLDGIEHSKRKEEDIVGIAHMLLSESLRFESSTGSYFQNQAHNLLTGLLAHVMLSPEYAGRRN
LRSLRQIVSEPEPSVLAMLRDIQEHSETVFIRETLGVFTNMTEQTFSGVYSTASKDTQWLSLDSYAALVC
GNAFRSSDIVTAKTDVFLNIPASILRSYPGIGRVIIGSLINAMVQADGGFSRRALFMLDEVDLLGYMRVL
EEARDRGRKYGISMMLLYQSVGQLERHFGKDGATSWIDGCAFASYAAIKALDTARNVSSQCGEMTVEVQG
RSRNIGWDTKGGGTRKSESVNFQRRPLIMPHEITQSMRKDEQIIIVQGHSPIRCGRAIYFRRKEMDAQAK
ANRFVRT
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 399854..409459
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| CUJ84_RS26145 (CUJ84_pRLN1000386) | 395936..396934 | + | 999 | WP_105008599 | ABC transporter substrate-binding protein | - |
| CUJ84_RS26150 (CUJ84_pRLN1000387) | 396942..398693 | + | 1752 | WP_105008600 | adenylate/guanylate cyclase domain-containing protein | - |
| CUJ84_RS26155 (CUJ84_pRLN1000388) | 398866..399219 | - | 354 | WP_105008601 | MarR family transcriptional regulator | - |
| CUJ84_RS26160 (CUJ84_pRLN1000389) | 399311..399850 | + | 540 | WP_105008602 | hypothetical protein | - |
| CUJ84_RS26165 (CUJ84_pRLN1000390) | 399854..400516 | + | 663 | WP_105008603 | lytic transglycosylase domain-containing protein | virB1 |
| CUJ84_RS26170 (CUJ84_pRLN1000391) | 400513..400812 | + | 300 | WP_105008604 | TrbC/VirB2 family protein | virB2 |
| CUJ84_RS26175 (CUJ84_pRLN1000392) | 400818..401156 | + | 339 | WP_105008605 | type IV secretion system protein VirB3 | virB3 |
| CUJ84_RS26180 (CUJ84_pRLN1000393) | 401149..403515 | + | 2367 | WP_105008606 | VirB4 family type IV secretion system protein | virb4 |
| CUJ84_RS26185 (CUJ84_pRLN1000394) | 403551..404213 | + | 663 | WP_245457601 | P-type DNA transfer protein VirB5 | virB5 |
| CUJ84_RS26190 (CUJ84_pRLN1000395) | 404210..404443 | + | 234 | WP_105008608 | EexN family lipoprotein | - |
| CUJ84_RS26195 (CUJ84_pRLN1000396) | 404447..405367 | + | 921 | WP_105008609 | type IV secretion system protein | virB6 |
| CUJ84_RS26200 (CUJ84_pRLN1000397) | 405420..405701 | + | 282 | WP_105008610 | hypothetical protein | - |
| CUJ84_RS26205 (CUJ84_pRLN1000398) | 405703..406374 | + | 672 | WP_105008611 | virB8 family protein | virB8 |
| CUJ84_RS26210 (CUJ84_pRLN1000399) | 406371..407228 | + | 858 | WP_105008612 | P-type conjugative transfer protein VirB9 | virB9 |
| CUJ84_RS26215 (CUJ84_pRLN1000400) | 407238..408410 | + | 1173 | WP_105008613 | type IV secretion system protein VirB10 | virB10 |
| CUJ84_RS26220 (CUJ84_pRLN1000401) | 408419..409459 | + | 1041 | WP_105008614 | P-type DNA transfer ATPase VirB11 | virB11 |
| CUJ84_RS26230 (CUJ84_pRLN1000403) | 409647..410678 | - | 1032 | WP_105008616 | hypothetical protein | - |
| CUJ84_RS26235 (CUJ84_pRLN1000404) | 410668..412332 | - | 1665 | WP_105008617 | N-6 DNA methylase | - |
| CUJ84_RS26240 (CUJ84_pRLN1000405) | 412618..413148 | + | 531 | Protein_382 | ATPase, T2SS/T4P/T4SS family | - |
| CUJ84_RS26245 (CUJ84_pRLN1000406) | 413290..414216 | - | 927 | WP_105008618 | nucleotidyltransferase and HEPN domain-containing protein | - |
Host bacterium
| ID | 8036 | GenBank | NZ_CP025013 |
| Plasmid name | pRLN1 | Incompatibility group | - |
| Plasmid size | 1098158 bp | Coordinate of oriT [Strand] | 702400..702428 [+] |
| Host baterium | Rhizobium leguminosarum strain Norway |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | bphC |
| Symbiosis gene | fixA, fixB |
| Anti-CRISPR | AcrIIA7, AcrIIA9 |