Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 107056 |
| Name | oriT_pRIVM_C009363_4 |
| Organism | Enterobacter kobei strain RIVM_C009363 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP119532 (5536..5580 [+], 45 nt) |
| oriT length | 45 nt |
| IRs (inverted repeats) | 7..13, 25..31 (CGTTTCT..AGAAACG) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 45 nt
>oriT_pRIVM_C009363_4
GCGCACCGTTTCTGAACGAAGTGAAGAAACGTCTAAGTGCGCCCT
GCGCACCGTTTCTGAACGAAGTGAAGAAACGTCTAAGTGCGCCCT
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file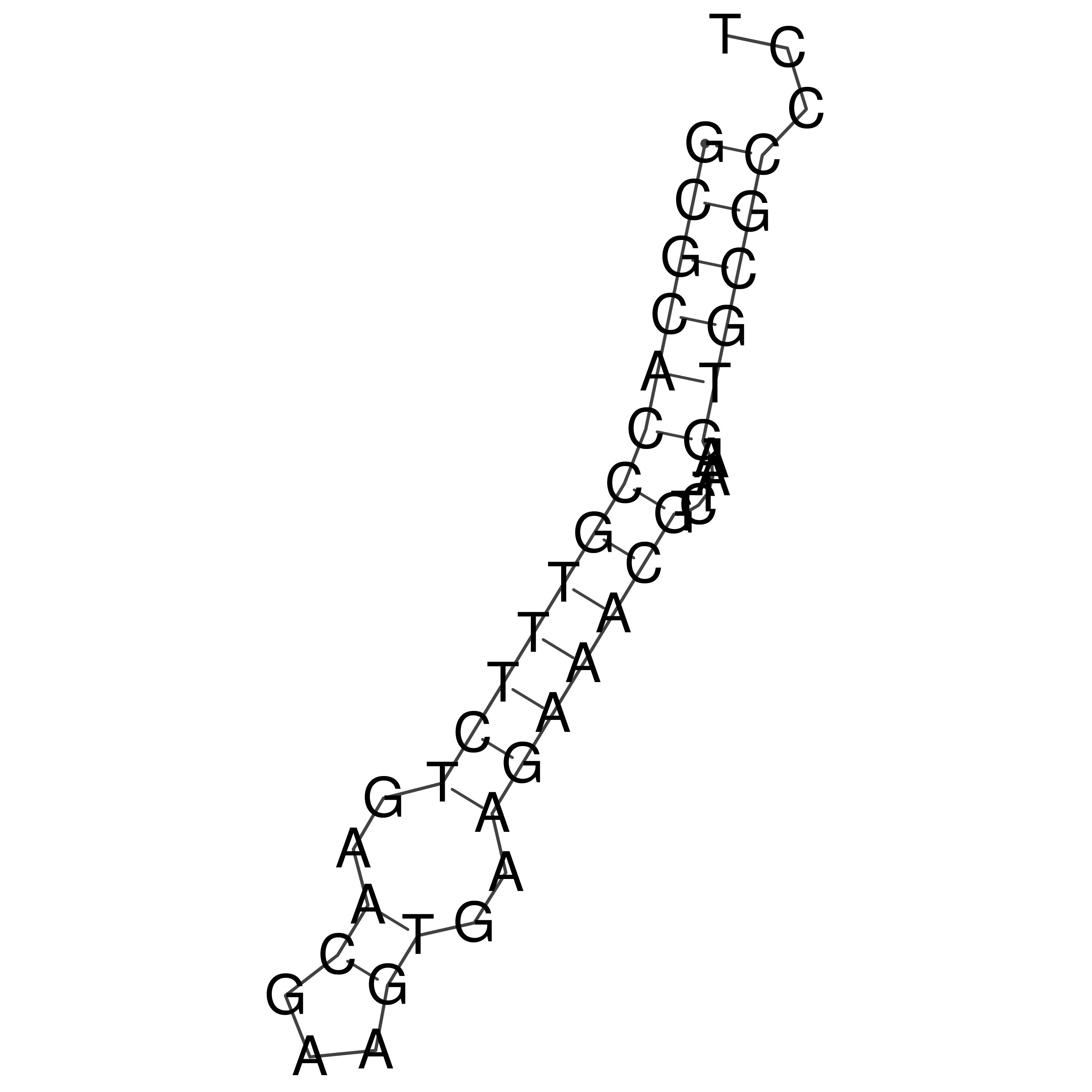
Relaxase
| ID | 4666 | GenBank | WP_040113349 |
| Name | mobA_P2D94_RS00025_pRIVM_C009363_4 |
UniProt ID | A0A734ICP3 |
| Length | 365 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 365 a.a. Molecular weight: 42217.70 Da Isoelectric Point: 6.3344
>WP_040113349.1 MULTISPECIES: mobilization protein MobA [Enterobacterales]
MASYHLSVKTGGKGSASPHADYISREGKYAREKDSDLEHKESGNMPAWAAHKPTEFWKAADTFERANGCT
YREIEIALPRELKPEQRLELVRDFVRQEIGDRHAYQFAIHNPKAAIAGGEQPHAHIMFSERMNDGIDRDP
QQYFKRANSKAPERGGAKKARFGETPTERKEHLVAQRERWADLQNKHLERYQHTDRVDARSLKAQGIERE
PERHLGAGQVQRFDTAQLQAILERRDAERQAEQSRDERDSVIDVTTSLREAISERDNLTIKQEQKPEPAQ
EDTSGRVFDFEKEPEKLNALVSDALKDVQDEIDLQSLVNETMAEFQGIHQEMERLAEKQRQQEKEQQRLA
EQTRQKPDKGWSFSR
MASYHLSVKTGGKGSASPHADYISREGKYAREKDSDLEHKESGNMPAWAAHKPTEFWKAADTFERANGCT
YREIEIALPRELKPEQRLELVRDFVRQEIGDRHAYQFAIHNPKAAIAGGEQPHAHIMFSERMNDGIDRDP
QQYFKRANSKAPERGGAKKARFGETPTERKEHLVAQRERWADLQNKHLERYQHTDRVDARSLKAQGIERE
PERHLGAGQVQRFDTAQLQAILERRDAERQAEQSRDERDSVIDVTTSLREAISERDNLTIKQEQKPEPAQ
EDTSGRVFDFEKEPEKLNALVSDALKDVQDEIDLQSLVNETMAEFQGIHQEMERLAEKQRQQEKEQQRLA
EQTRQKPDKGWSFSR
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A734ICP3 |
Host bacterium
| ID | 7492 | GenBank | NZ_CP119532 |
| Plasmid name | pRIVM_C009363_4 | Incompatibility group | ColE10 |
| Plasmid size | 12808 bp | Coordinate of oriT [Strand] | 5536..5580 [+] |
| Host baterium | Enterobacter kobei strain RIVM_C009363 |
Cargo genes
| Drug resistance gene | mcr-4.3 |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |