Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 106985 |
| Name | oriT_p48659_KPC |
| Organism | Morganella morganii strain MM48659 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP070535 (46259..46357 [+], 99 nt) |
| oriT length | 99 nt |
| IRs (inverted repeats) | 77..82, 89..94 (AAAAAA..TTTTTT) 77..82, 88..93 (AAAAAA..TTTTTT) 31..38, 41..48 (AGCGTGAT..ATCACGCT) 17..23, 35..41 (TAAATCA..TGATTTA) |
| Location of nic site | 59..60 |
| Conserved sequence flanking the nic site |
GGTGTATAGC |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 99 nt
>oriT_p48659_KPC
TTTGTTTTTTTTCTTTTAAATCAGTGCGATAGCGTGATTTATCACGCTGCGTTAGGTGTATAGCAGGTTAAGGGATAAAAAATCATCTTTTTTTGGTAG
TTTGTTTTTTTTCTTTTAAATCAGTGCGATAGCGTGATTTATCACGCTGCGTTAGGTGTATAGCAGGTTAAGGGATAAAAAATCATCTTTTTTTGGTAG
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file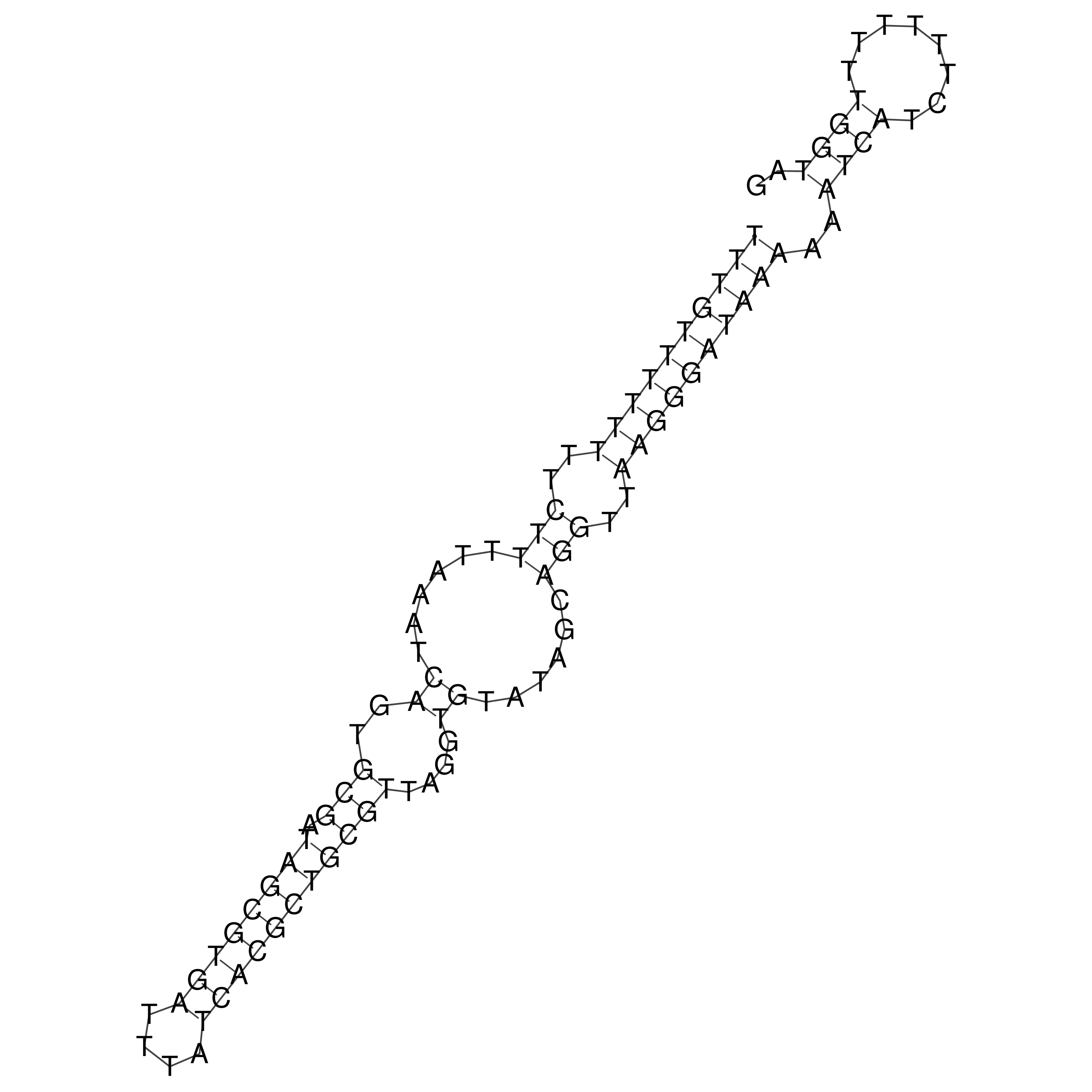
Relaxase
| ID | 4610 | GenBank | WP_000130000 |
| Name | Replic_Relax_JW294_RS00185_p48659_KPC |
UniProt ID | R4WML4 |
| Length | 101 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 101 a.a. Molecular weight: 11477.11 Da Isoelectric Point: 7.5204
>WP_000130000.1 MULTISPECIES: PadR family transcriptional regulator [Pseudomonadota]
MTDKDLYGGLIRLHILHHAAEEPVFGLGIIEELRRHGYEMSAGTVYPMLHGLEKKGYLTSRHERTGRRER
RVYDITEQGRTALADAKTKVKELFGELVEGG
MTDKDLYGGLIRLHILHHAAEEPVFGLGIIEELRRHGYEMSAGTVYPMLHGLEKKGYLTSRHERTGRRER
RVYDITEQGRTALADAKTKVKELFGELVEGG
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | R4WML4 |
Host bacterium
| ID | 7422 | GenBank | NZ_CP070535 |
| Plasmid name | p48659_KPC | Incompatibility group | IncP6 |
| Plasmid size | 62326 bp | Coordinate of oriT [Strand] | 46259..46357 [+] |
| Host baterium | Morganella morganii strain MM48659 |
Cargo genes
| Drug resistance gene | aac(6')-Ib, blaKPC-2, mph(A), sul1, qacE, ARR-3, catB3, blaOXA-1, aac(6')-Ib-cr |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |