Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 106854 |
| Name | oriT_pSYMB |
| Organism | Sinorhizobium meliloti Rm41 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NC_018701 (992496..992524 [-], 29 nt) |
| oriT length | 29 nt |
| IRs (inverted repeats) | 14..19, 24..29 (CGTCGC..GCGACG) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 29 nt
>oriT_pSYMB
AGGGCGCAATATACGTCGCTGGCGCGACG
AGGGCGCAATATACGTCGCTGGCGCGACG
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file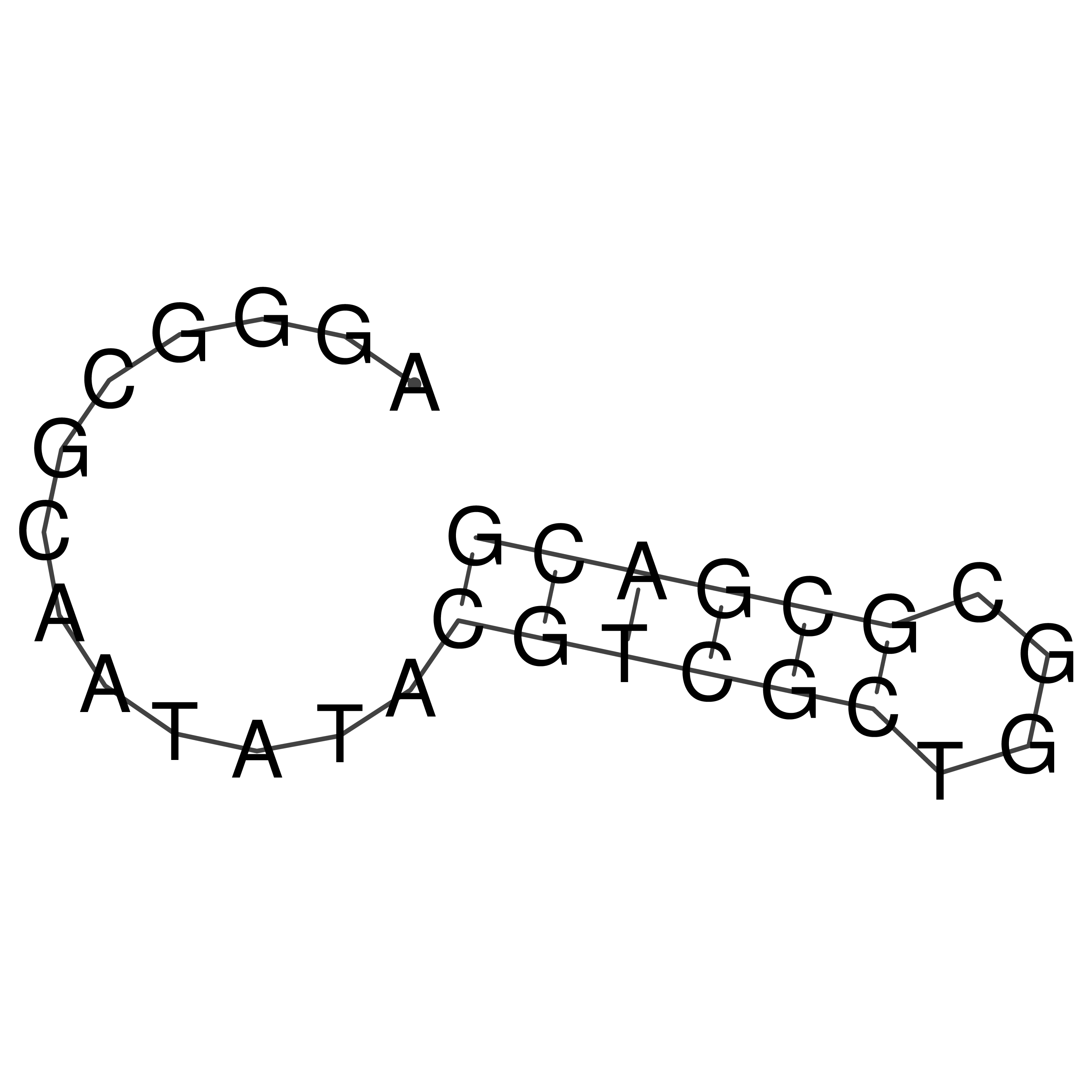
Relaxase
| ID | 4522 | GenBank | WP_015008383 |
| Name | traA_BN406_RS30740_pSYMB |
UniProt ID | _ |
| Length | 1539 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1539 a.a. Molecular weight: 169985.76 Da Isoelectric Point: 9.4869
>WP_015008383.1 Ti-type conjugative transfer relaxase TraA [Sinorhizobium meliloti]
MAIMFVRAQVIGRGAGRSIVSAAAYRHRTRMIDEQAGTSFSYRGGASELVHEELALPDDIPAWLKAAIDG
QSVAKASEALWNAVEAHETRADAQLARELIIALPEELTRAENIALVREFVRDNLTSKGMVADWVYHDKDG
NPHIHLMTALRPLTEQGFGPKKVPVLSEDGEPLRVVTPDRPNGKIVYKLWAGDKETIKAWKIAWAETANR
HLALAGHEIRLDGRSYAEQGLDGIAQKHLGPEKAALARKGIAMYFAPADLARRQEMADRLLAEPELLLKQ
LGNERSTFDERDIARALHRYVDDPVDFANIRARLMASDELVLLKPQQVDAEAGEAKQPAVFTTREMLRLE
YAMARSAEVLSRRKGFGVSNARAAAAVRSIETADTEKPFRLDPEQVDAVRHVTRDNAIAAVVGLAGAGKS
TLLAAARVAWEGEGRRVIGAALAGKAAEGLEDSSGIRSRTLASWELAWESGREQLNRGDVLVIDEAGMVS
SQQMARILKAVEDAGAKAVLVGDAMQLQPIEAGAAFRAITERIGFAELAGVRRQRDAWARDASRLFARGK
VEEGLDAYAQQGRIVETETRAEIVDRIVADWANARRDLLQKSADGEHPGRLRGDELLVLAHTNDDVRKLN
EALRQVMIGEGALAGAREFQTARGLREFAAGDRIIFLENARFVEPRARRLGPQYVKNGMLGTVVSTGDRR
GDTLLSVRLDSGRDVVISQDSYRNVDHGYAATIHKSQGSTVDRTFVLATGMMDQHLTYVAMTRHRDRADL
YAAKEDFEAKPEWGRKPRVDHAAVVTGELVKEGMAKFRPNDEDADESPYADIRTDDGTVNRLWGVSLPKA
LKDAGVAEGDTITLRKDGVERVKVQVPIVDEQTGEKRFEERQVDRNVWSASQLETAAARRERIERESHRP
QLFAALVERLSRSGAKTTTLDFEDEAGYQAQARDFARRRGLYHLSLVAAGMEAEVLRRWAGIAEKREQVA
KLWERASVALGFAIERERRVAYNEERTETLSTGIPSDGKYLVPPTTTFSRSVAEDARLAQLSSQRWKERE
AIVHPVLAKVYRDPDGALSALNALASDAAIEPRKLAEDLGLAPDRLGRLRGSELVVDGRAARDERTAATV
ALSELLPLARAHATEFRRNAERFGIREQQRRAHMALSVPALSKTAMARLVEIEAVRKQGGDDAYRTAFAY
AVEDRLLVQEVKAVNEALTARFGWSAFTAKADVIAERNIAERMPEDLAPERREKLKRLFAVIRRFAEEQH
LAERQDRSKIVAGASVELGKGTFAVLPMLAPVTEFKTTVDEEARERALAAPHYAHHRAALVETATRVWRD
PADAIGKIEDLIVKGFAAERIAAAVTNDPAAYGALRGSDRIMDKLLAVGRERKGALQAVPEAASRIRSLG
ASYASALDAETRSITEERRRMAVAIPGLSPAAEDALKRLAAQIKNKDGKLDVAAGSLDPRIAREFAKVSR
ALDERFGRNAILRGETDVINRVSPAQRRAFEAMRDRLTILQQAVRIQSSQKIVSERRQRAINQSRGIRM
MAIMFVRAQVIGRGAGRSIVSAAAYRHRTRMIDEQAGTSFSYRGGASELVHEELALPDDIPAWLKAAIDG
QSVAKASEALWNAVEAHETRADAQLARELIIALPEELTRAENIALVREFVRDNLTSKGMVADWVYHDKDG
NPHIHLMTALRPLTEQGFGPKKVPVLSEDGEPLRVVTPDRPNGKIVYKLWAGDKETIKAWKIAWAETANR
HLALAGHEIRLDGRSYAEQGLDGIAQKHLGPEKAALARKGIAMYFAPADLARRQEMADRLLAEPELLLKQ
LGNERSTFDERDIARALHRYVDDPVDFANIRARLMASDELVLLKPQQVDAEAGEAKQPAVFTTREMLRLE
YAMARSAEVLSRRKGFGVSNARAAAAVRSIETADTEKPFRLDPEQVDAVRHVTRDNAIAAVVGLAGAGKS
TLLAAARVAWEGEGRRVIGAALAGKAAEGLEDSSGIRSRTLASWELAWESGREQLNRGDVLVIDEAGMVS
SQQMARILKAVEDAGAKAVLVGDAMQLQPIEAGAAFRAITERIGFAELAGVRRQRDAWARDASRLFARGK
VEEGLDAYAQQGRIVETETRAEIVDRIVADWANARRDLLQKSADGEHPGRLRGDELLVLAHTNDDVRKLN
EALRQVMIGEGALAGAREFQTARGLREFAAGDRIIFLENARFVEPRARRLGPQYVKNGMLGTVVSTGDRR
GDTLLSVRLDSGRDVVISQDSYRNVDHGYAATIHKSQGSTVDRTFVLATGMMDQHLTYVAMTRHRDRADL
YAAKEDFEAKPEWGRKPRVDHAAVVTGELVKEGMAKFRPNDEDADESPYADIRTDDGTVNRLWGVSLPKA
LKDAGVAEGDTITLRKDGVERVKVQVPIVDEQTGEKRFEERQVDRNVWSASQLETAAARRERIERESHRP
QLFAALVERLSRSGAKTTTLDFEDEAGYQAQARDFARRRGLYHLSLVAAGMEAEVLRRWAGIAEKREQVA
KLWERASVALGFAIERERRVAYNEERTETLSTGIPSDGKYLVPPTTTFSRSVAEDARLAQLSSQRWKERE
AIVHPVLAKVYRDPDGALSALNALASDAAIEPRKLAEDLGLAPDRLGRLRGSELVVDGRAARDERTAATV
ALSELLPLARAHATEFRRNAERFGIREQQRRAHMALSVPALSKTAMARLVEIEAVRKQGGDDAYRTAFAY
AVEDRLLVQEVKAVNEALTARFGWSAFTAKADVIAERNIAERMPEDLAPERREKLKRLFAVIRRFAEEQH
LAERQDRSKIVAGASVELGKGTFAVLPMLAPVTEFKTTVDEEARERALAAPHYAHHRAALVETATRVWRD
PADAIGKIEDLIVKGFAAERIAAAVTNDPAAYGALRGSDRIMDKLLAVGRERKGALQAVPEAASRIRSLG
ASYASALDAETRSITEERRRMAVAIPGLSPAAEDALKRLAAQIKNKDGKLDVAAGSLDPRIAREFAKVSR
ALDERFGRNAILRGETDVINRVSPAQRRAFEAMRDRLTILQQAVRIQSSQKIVSERRQRAINQSRGIRM
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4CP
| ID | 4586 | GenBank | WP_013850291 |
| Name | tcpA_BN406_RS37775_pSYMB |
UniProt ID | _ |
| Length | 946 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 946 a.a. Molecular weight: 101616.47 Da Isoelectric Point: 4.6107
>WP_013850291.1 FtsK/SpoIIIE domain-containing protein [Sinorhizobium meliloti]
MRIPRTNFSPAALYDANHADDYEDPHPAAQGAAHVPAAPEAAAHDHLFEAPEAPRRAPGHRDEETGYAGR
AARLYGHGSAYAPAEVTVSTRDPLPTLAEITGLSGEPGWESHFFLSPNVRFTRTPERELMKRHPPAPEES
RIEADEAAAEASDAETVMDVVPAEPAPSVVETELPSYSPSELLRVLTQQLPSWSAARSQAPEASVTKPAI
TESVAVAEEKPATSETTALPVTDEVPVVPHALPVANLAPESAAVEEDVPHQADARLAYLSDFAFFEFMPL
EVAAVPPTVTEPVKEAARIPAPIAAAPKVSPPKIVAAMPVEIRPQPPTAISSLFRVVECRRPEPATAEPV
TAAPQDIAAEPAAAEAAALPAPRVVETPAPALAEPAIVPASEPAPEAPVTRAAITMPAVIQRSSPSLPPI
GAIEPLQGGDAYEFPSKELLQEPPQGQGFFMTQEQLEQNAGLLESVLEDFGVKGEIIHVRPGPVVTLYEF
EPAPGVKSSRVIGLADDIARSMSALSARVAVVPGRNVIGIELPNATRETVYFRELIESGDFQKTGCKLAL
CLGKTIGGEPVIAELAKMPHLLVAGTTGSGKSVAINTMILSLLYRLKPEECRLIMVDPKMLELSVYDGIP
HLLTPVVTDPKKAVMALKWAVREMEDRYRKMSRLGVRNIDGYNQRAAAAREKGAPILATVQTGFEKGTGE
PLFEQQEMDLSPMPYIVVIVDEMADLMMVAGKEIEGAIQRLAQMARAAGIHLIMATQRPSVDVITGTIKA
NFPTRISFQVTSKIDSRTILGEQGAEQLLGQGDMLHMAGGGRIARVHGPFVSDQEVEHVVAHLKTQGRPE
YLETVTADEEEEEVEEDQGAVFDKSAIAAEDGNELYDQAVKVVLRDKKCSTSYIQRRLGIGYNRAASLVE
RMEKDGLVGPANHVGKREIIYGNRDNAPKPESDDLD
MRIPRTNFSPAALYDANHADDYEDPHPAAQGAAHVPAAPEAAAHDHLFEAPEAPRRAPGHRDEETGYAGR
AARLYGHGSAYAPAEVTVSTRDPLPTLAEITGLSGEPGWESHFFLSPNVRFTRTPERELMKRHPPAPEES
RIEADEAAAEASDAETVMDVVPAEPAPSVVETELPSYSPSELLRVLTQQLPSWSAARSQAPEASVTKPAI
TESVAVAEEKPATSETTALPVTDEVPVVPHALPVANLAPESAAVEEDVPHQADARLAYLSDFAFFEFMPL
EVAAVPPTVTEPVKEAARIPAPIAAAPKVSPPKIVAAMPVEIRPQPPTAISSLFRVVECRRPEPATAEPV
TAAPQDIAAEPAAAEAAALPAPRVVETPAPALAEPAIVPASEPAPEAPVTRAAITMPAVIQRSSPSLPPI
GAIEPLQGGDAYEFPSKELLQEPPQGQGFFMTQEQLEQNAGLLESVLEDFGVKGEIIHVRPGPVVTLYEF
EPAPGVKSSRVIGLADDIARSMSALSARVAVVPGRNVIGIELPNATRETVYFRELIESGDFQKTGCKLAL
CLGKTIGGEPVIAELAKMPHLLVAGTTGSGKSVAINTMILSLLYRLKPEECRLIMVDPKMLELSVYDGIP
HLLTPVVTDPKKAVMALKWAVREMEDRYRKMSRLGVRNIDGYNQRAAAAREKGAPILATVQTGFEKGTGE
PLFEQQEMDLSPMPYIVVIVDEMADLMMVAGKEIEGAIQRLAQMARAAGIHLIMATQRPSVDVITGTIKA
NFPTRISFQVTSKIDSRTILGEQGAEQLLGQGDMLHMAGGGRIARVHGPFVSDQEVEHVVAHLKTQGRPE
YLETVTADEEEEEVEEDQGAVFDKSAIAAEDGNELYDQAVKVVLRDKKCSTSYIQRRLGIGYNRAASLVE
RMEKDGLVGPANHVGKREIIYGNRDNAPKPESDDLD
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Host bacterium
| ID | 7291 | GenBank | NC_018701 |
| Plasmid name | pSYMB | Incompatibility group | - |
| Plasmid size | 1664896 bp | Coordinate of oriT [Strand] | 992496..992524 [-] |
| Host baterium | Sinorhizobium meliloti Rm41 |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | htpB |
| Metal resistance gene | actP |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | AcrIIA7 |