Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 106776 |
| Name | oriT_pCF1807-4 |
| Organism | Citrobacter freundii strain CF1807 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP110890 (11379..11468 [-], 90 nt) |
| oriT length | 90 nt |
| IRs (inverted repeats) | 34..41, 44..51 (AGTGCGAT..ATCGCACT) |
| Location of nic site | 62..63 |
| Conserved sequence flanking the nic site |
GGTGTATAGC |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 90 nt
>oriT_pCF1807-4
TATTTCATTTTTTGTTTTTTTAATCAGTTAGTTAGTGCGATTCATCGCACTGCGTTAGGTGTATAGCAGGTTAAGGGCTAAAAAATAATC
TATTTCATTTTTTGTTTTTTTAATCAGTTAGTTAGTGCGATTCATCGCACTGCGTTAGGTGTATAGCAGGTTAAGGGCTAAAAAATAATC
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file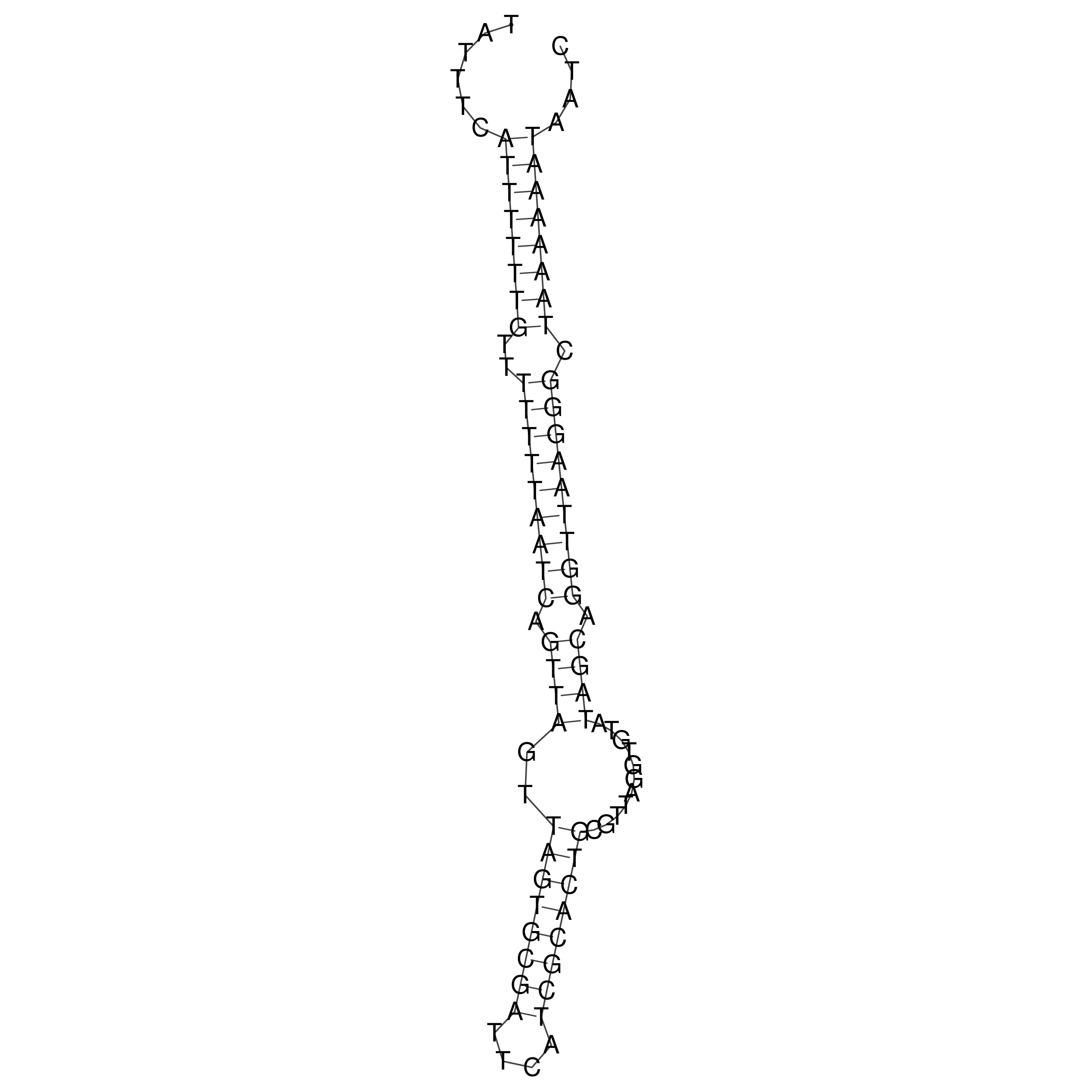
Relaxase
| ID | 4467 | GenBank | WP_024561907 |
| Name | mobF_OQW59_RS00110_pCF1807-4 |
UniProt ID | H1ZMV6 |
| Length | 1072 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1072 a.a. Molecular weight: 118896.59 Da Isoelectric Point: 6.4636
>WP_024561907.1 MULTISPECIES: MobF family relaxase [Enterobacterales]
MLDITTITRIDIASVVGYYADAKDDYYSKDSSFTAWHGQGAEALGLSGEVSSQRFKELLVGEIDPFTQMK
RSSGDATKERLGYDLTFSAPKGVSMQALIHGDKAIIAAHEKAVAAAVQEAEKLAQARTTHAGKTITQNTG
NLVVASFRHETSRALDPELHTHAFVMNMTQRGDGQWRALKNDELMRAKMHLGDVYKQELALELTKAGYEL
RYNNKNNTFDMAHFTDEQIRGFSRRSAQIEEGLAAMGLTRETADAQTKSRVSMATRDKKTDDYSREDIHK
EWVNRANELGIDFNDLSWAGKGKDGPQTSMNTVPNFTSPEVKADRAIQFALKSLSERDASFDQSRLLAVA
NKKAIGQASKQDIEAAYYRAVAKGAIIEGEARYTTTTDQRSKDAPPLPAMTKAEWQAQLTDKKGMNNDQA
KALVAEGIKSGRLKKTSHRVTTVEGIRVEKAVLNIEARGRGKVIAALTSEQVTAQLVGKTLKAEQQKSVV
DICTTTDRFVAAHGFAGTGKSYMTMSAKAVLESQGFNVTALAPYGTQKKSLEDEGMPARTVAAFVKAKDK
KIDEKSVVFIDEAGVIPARQMKVLMETIEKAGARAVFLGDTSQTKAVEAGKPFEQLISAGMQTSYMKDIQ
RQKNDVLLEAVKLAAEGHISGSLARLSNISAEKDQDKRLEAVANRWLSFSPEQREGTLIISGTNESRVIL
NSQIREALQLQGKGVEVNFLERVDSTQAERRDSKYYQVGQIIIPEKDYKNGLQRGESYRVLDTGPGNLLT
VAGSDGQKISFSPRTHKNLSVYTSVAAELAVGDKVMVTRNDKTLDVANGDRFTVTGTTEKGTFTLTNAKG
REIELDQKQAAYLSYAYATTVHKAQGLTCDRVLFNIDTRSLTTSKDVFYVGISRARHEVEIFTDDKKRLL
SSASRNSPKTTASEIDRFLGMEERYKDVNRSYGHSDGQQEKSYGHDAGHQTEEKENTMKNATEAAHDGPD
TSGTENTSQDLRQREQEEVNAAIRYEQRQEATPEMEANPYHDYERYADEGPDYGDMQDYSTYEEYERARD
VELPQHEHENHQHDHDKEGYGL
MLDITTITRIDIASVVGYYADAKDDYYSKDSSFTAWHGQGAEALGLSGEVSSQRFKELLVGEIDPFTQMK
RSSGDATKERLGYDLTFSAPKGVSMQALIHGDKAIIAAHEKAVAAAVQEAEKLAQARTTHAGKTITQNTG
NLVVASFRHETSRALDPELHTHAFVMNMTQRGDGQWRALKNDELMRAKMHLGDVYKQELALELTKAGYEL
RYNNKNNTFDMAHFTDEQIRGFSRRSAQIEEGLAAMGLTRETADAQTKSRVSMATRDKKTDDYSREDIHK
EWVNRANELGIDFNDLSWAGKGKDGPQTSMNTVPNFTSPEVKADRAIQFALKSLSERDASFDQSRLLAVA
NKKAIGQASKQDIEAAYYRAVAKGAIIEGEARYTTTTDQRSKDAPPLPAMTKAEWQAQLTDKKGMNNDQA
KALVAEGIKSGRLKKTSHRVTTVEGIRVEKAVLNIEARGRGKVIAALTSEQVTAQLVGKTLKAEQQKSVV
DICTTTDRFVAAHGFAGTGKSYMTMSAKAVLESQGFNVTALAPYGTQKKSLEDEGMPARTVAAFVKAKDK
KIDEKSVVFIDEAGVIPARQMKVLMETIEKAGARAVFLGDTSQTKAVEAGKPFEQLISAGMQTSYMKDIQ
RQKNDVLLEAVKLAAEGHISGSLARLSNISAEKDQDKRLEAVANRWLSFSPEQREGTLIISGTNESRVIL
NSQIREALQLQGKGVEVNFLERVDSTQAERRDSKYYQVGQIIIPEKDYKNGLQRGESYRVLDTGPGNLLT
VAGSDGQKISFSPRTHKNLSVYTSVAAELAVGDKVMVTRNDKTLDVANGDRFTVTGTTEKGTFTLTNAKG
REIELDQKQAAYLSYAYATTVHKAQGLTCDRVLFNIDTRSLTTSKDVFYVGISRARHEVEIFTDDKKRLL
SSASRNSPKTTASEIDRFLGMEERYKDVNRSYGHSDGQQEKSYGHDAGHQTEEKENTMKNATEAAHDGPD
TSGTENTSQDLRQREQEEVNAAIRYEQRQEATPEMEANPYHDYERYADEGPDYGDMQDYSTYEEYERARD
VELPQHEHENHQHDHDKEGYGL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | H1ZMV6 |
T4CP
| ID | 4514 | GenBank | WP_032493179 |
| Name | t4cp2_OQW59_RS00105_pCF1807-4 |
UniProt ID | _ |
| Length | 510 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 510 a.a. Molecular weight: 57912.96 Da Isoelectric Point: 9.5127
>WP_032493179.1 MULTISPECIES: type IV secretion system DNA-binding domain-containing protein [Enterobacterales]
MTEQERGLIFLCSLLGPIPFVWFITAKFVYGINSETAHYLVPYLLKNTFNLWPLWVAIIAGAVIGLCGFI
GFIIYDKGRVYKGPRFKRVLRGTKLVRASSLAEKTKERDHKQLTVAGVPVPTEYETLHFSIGGATGTGKS
TIIKEAIYSSLKRGTDKRIVLDPNGEYVETFYRPGDVILSPYDKRSPGWSMFNEIRDDFDYDLYAVSIIQ
ESKERSAEEWYRYGRLIYREVSKKLLTLKRSPTMEEVMHWCTIVENEKLKEFVKGTPAEALFIKGAERAT
GSARFVLGDKLAPHLKMPAGDFSLRDWLEDDKPGTLFITWQEQMKKSLNPLISCWLDIMFSSLLGMGKRD
QRTLFFIDELESLEYLPNLQDLLTKGRKSGASVWAGYQTYSQLIERYGINIAETLLGSMRSGIVMGSSRL
GKETLEQMSRALGEIEGEVEHKQGNPFQAGIGNRPRIETRTVRAVTPTEISELKNLTGYVYFPGDLPVGK
FKTKHVQYTRKNPVPGIIMR
MTEQERGLIFLCSLLGPIPFVWFITAKFVYGINSETAHYLVPYLLKNTFNLWPLWVAIIAGAVIGLCGFI
GFIIYDKGRVYKGPRFKRVLRGTKLVRASSLAEKTKERDHKQLTVAGVPVPTEYETLHFSIGGATGTGKS
TIIKEAIYSSLKRGTDKRIVLDPNGEYVETFYRPGDVILSPYDKRSPGWSMFNEIRDDFDYDLYAVSIIQ
ESKERSAEEWYRYGRLIYREVSKKLLTLKRSPTMEEVMHWCTIVENEKLKEFVKGTPAEALFIKGAERAT
GSARFVLGDKLAPHLKMPAGDFSLRDWLEDDKPGTLFITWQEQMKKSLNPLISCWLDIMFSSLLGMGKRD
QRTLFFIDELESLEYLPNLQDLLTKGRKSGASVWAGYQTYSQLIERYGINIAETLLGSMRSGIVMGSSRL
GKETLEQMSRALGEIEGEVEHKQGNPFQAGIGNRPRIETRTVRAVTPTEISELKNLTGYVYFPGDLPVGK
FKTKHVQYTRKNPVPGIIMR
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 12039..29539
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| OQW59_RS00055 (OQW59_00055) | 7154..7354 | + | 201 | WP_015060066 | hypothetical protein | - |
| OQW59_RS00060 (OQW59_00060) | 7409..7906 | + | 498 | WP_040115102 | MucR family transcriptional regulator | - |
| OQW59_RS00065 (OQW59_00065) | 8279..8536 | - | 258 | WP_157676234 | hypothetical protein | - |
| OQW59_RS00070 (OQW59_00070) | 8686..8892 | + | 207 | WP_032493183 | DUF905 family protein | - |
| OQW59_RS00075 (OQW59_00075) | 8967..9149 | + | 183 | WP_032493182 | hypothetical protein | - |
| OQW59_RS00080 (OQW59_00080) | 9208..9558 | - | 351 | WP_014014961 | hypothetical protein | - |
| OQW59_RS00085 (OQW59_00085) | 9620..9988 | - | 369 | WP_000360828 | plasmid stabilization protein StbC | - |
| OQW59_RS00090 (OQW59_00090) | 9991..10707 | - | 717 | WP_000861579 | StbB family protein | - |
| OQW59_RS00095 (OQW59_00095) | 10716..11147 | - | 432 | WP_029697170 | plasmid stabilization protein StbA | - |
| OQW59_RS00100 (OQW59_00100) | 11614..12039 | + | 426 | WP_225247941 | traK protein | - |
| OQW59_RS00105 (OQW59_00105) | 12039..13571 | + | 1533 | WP_032493179 | type IV secretion system DNA-binding domain-containing protein | virb4 |
| OQW59_RS00110 (OQW59_00110) | 13576..16794 | + | 3219 | WP_024561907 | MobF family relaxase | - |
| OQW59_RS00115 (OQW59_00115) | 16791..17456 | + | 666 | WP_000971716 | DUF6710 family protein | - |
| OQW59_RS00120 (OQW59_00120) | 17449..17811 | + | 363 | WP_024561906 | hypothetical protein | - |
| OQW59_RS00125 (OQW59_00125) | 18739..19251 | - | 513 | WP_029697418 | phospholipase D family protein | - |
| OQW59_RS00130 (OQW59_00130) | 19263..20264 | - | 1002 | WP_001076634 | ATPase, T2SS/T4P/T4SS family | virB11 |
| OQW59_RS00135 (OQW59_00135) | 20315..21448 | - | 1134 | WP_000101920 | type IV secretion system protein VirB10 | virB10 |
| OQW59_RS00140 (OQW59_00140) | 21448..22335 | - | 888 | WP_000758230 | TrbG/VirB9 family P-type conjugative transfer protein | virB9 |
| OQW59_RS00145 (OQW59_00145) | 22346..23041 | - | 696 | WP_001208352 | type IV secretion system protein | virB8 |
| OQW59_RS28915 | 23052..23183 | - | 132 | WP_032493177 | hypothetical protein | - |
| OQW59_RS00150 (OQW59_00150) | 23260..24297 | - | 1038 | WP_014014948 | type IV secretion system protein | virB6 |
| OQW59_RS00155 (OQW59_00155) | 24314..24556 | - | 243 | WP_000858958 | EexN family lipoprotein | - |
| OQW59_RS00160 (OQW59_00160) | 24566..25267 | - | 702 | WP_014014949 | type IV secretion system protein | virB5 |
| OQW59_RS00165 (OQW59_00165) | 25283..27856 | - | 2574 | WP_032493176 | VirB4 family type IV secretion/conjugal transfer ATPase | virb4 |
| OQW59_RS00170 (OQW59_00170) | 27857..28174 | - | 318 | WP_000462629 | VirB3 family type IV secretion system protein | virB3 |
| OQW59_RS00175 (OQW59_00175) | 28214..28510 | - | 297 | WP_015060075 | TrbC/VirB2 family protein | virB2 |
| OQW59_RS00185 (OQW59_00180) | 28757..29539 | - | 783 | WP_024562119 | lytic transglycosylase domain-containing protein | virB1 |
| OQW59_RS00190 (OQW59_00185) | 29639..29956 | + | 318 | WP_000960955 | H-NS family nucleoid-associated regulatory protein | - |
| OQW59_RS00195 (OQW59_00190) | 29960..30310 | + | 351 | WP_024562120 | hypothetical protein | - |
| OQW59_RS00200 (OQW59_00195) | 30307..30645 | + | 339 | WP_032493175 | hypothetical protein | - |
| OQW59_RS00205 (OQW59_00200) | 30649..30957 | + | 309 | WP_000733764 | hypothetical protein | - |
| OQW59_RS00210 (OQW59_00205) | 31242..31427 | + | 186 | WP_024562121 | ribbon-helix-helix domain-containing protein | - |
| OQW59_RS00215 (OQW59_00210) | 31424..32032 | - | 609 | WP_000275218 | recombinase family protein | - |
| OQW59_RS00220 (OQW59_00215) | 32219..32551 | - | 333 | WP_045723726 | hypothetical protein | - |
| OQW59_RS00225 (OQW59_00220) | 32623..32817 | - | 195 | WP_050136673 | hypothetical protein | - |
| OQW59_RS00230 (OQW59_00225) | 32904..33533 | - | 630 | WP_064754415 | hypothetical protein | - |
Host bacterium
| ID | 7213 | GenBank | NZ_CP110890 |
| Plasmid name | pCF1807-4 | Incompatibility group | IncN3 |
| Plasmid size | 33719 bp | Coordinate of oriT [Strand] | 11379..11468 [-] |
| Host baterium | Citrobacter freundii strain CF1807 |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |