Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 106136 |
| Name | oriT_p733-MDR |
| Organism | Salmonella enterica strain 733 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP060847 (178959..179243 [-], 285 nt) |
| oriT length | 285 nt |
| IRs (inverted repeats) | 184..189, 191..196 (AAAAGT..ACTTTT) |
| Location of nic site | 109..110 |
| Conserved sequence flanking the nic site |
TTTGGTTAAA |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 285 nt
>oriT_p733-MDR
GGTTATTGCTACTTAATGCCGATAACGACTCAGGCTTTGAGGTTTTTTTATACGGTTCACATTTCGTTAGCAAGGCCAGGGTTTTTTGATAAAATTCTGGTTAGTTTGGTTAAAAAGTGTTACAAGTAAGGGTAATGGCTGAAAGGTTAGTTTTAAGGTTCAAAGCGGCAGTATTAAAATTCCAAAAGTTACTTTTCATCCTTCAGAATCCAGACCTTAATTTCATGTAGAAGATTCGTACAATTGTATTGGCGCAAGGACAATCCGCACATGTCAGAATCAGAT
GGTTATTGCTACTTAATGCCGATAACGACTCAGGCTTTGAGGTTTTTTTATACGGTTCACATTTCGTTAGCAAGGCCAGGGTTTTTTGATAAAATTCTGGTTAGTTTGGTTAAAAAGTGTTACAAGTAAGGGTAATGGCTGAAAGGTTAGTTTTAAGGTTCAAAGCGGCAGTATTAAAATTCCAAAAGTTACTTTTCATCCTTCAGAATCCAGACCTTAATTTCATGTAGAAGATTCGTACAATTGTATTGGCGCAAGGACAATCCGCACATGTCAGAATCAGAT
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file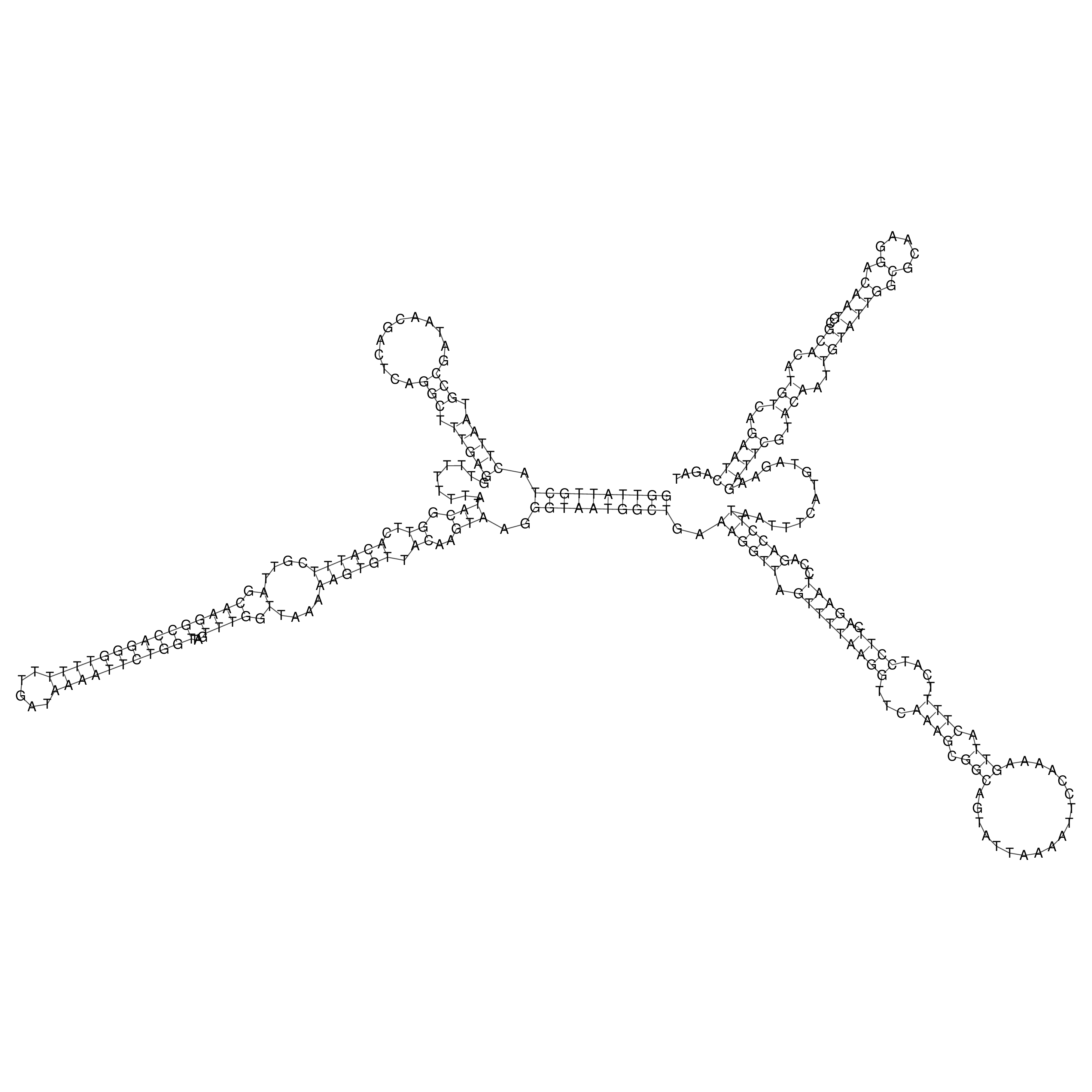
Relaxase
| ID | 4111 | GenBank | WP_038989148 |
| Name | mobH_IAR32_RS23380_p733-MDR |
UniProt ID | A0A7T0P871 |
| Length | 1011 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1011 a.a. Molecular weight: 113008.03 Da Isoelectric Point: 4.5108
>WP_038989148.1 MULTISPECIES: MobH family relaxase [Enterobacteriaceae]
MNFRALFLSMQRVFGIFSRRENDVSELMMKDAANFSPFAQIIGEQKYTVPDHPNPEVLKFIEYPTRPAGI
QTFNEQSILSLYRDKLHSISMMLAISDGDIREDAYTFTNLVLKPLIEYIRWIHLLPASENHHHNGIGGLL
SHSLEVAMISLKNANHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLSETLTWAPS
SQSLLDWARENNVVEYEIHWRKRIHNQHNIWSSVFLERILDPVCMSFLDRVKKERVYAKMVTALNVYNDG
NDFLSKCVRTSDYYSTGTDLNVLRDPIMGLRSNDAAARAIGTIKHNFTSININNYKTKPMHIIIVNGEVY
LNENAFLDFVLSDFAAHKFNFPQGDAGKTVLVESLVLRGYVEPYDDERVVHYFIPGTYSENEIASIFRNG
IGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRESVTRVTDTVN
DAIENTPQYGLQLVNGPDADSNNIISSENTESITDSLEESGADISNEIFETQVVTAIDTAETVNADEPEQ
VEEHDDRSQIHLVEQLHEMLLSAPLPHHAVINIDSVPYLDLDAAIALIPGIDEAAFCNGPFFQLTYRDGS
LDGMWIVRDVNNLRLIQLGDNCAGMQVSTSEPRNTSSLKSLFDTSMYQPLDIPEAPSVNEAASPPQTPLE
LPQPRLNAPVAEEASSVAEQTNAHSEPDSVIATEYEQYGHLLEETLDSDGEAYSDLIASDSTEAEYPATD
PQSSDFAQLPRETALSVAPGDLDYSEGAIKPPAPDATGKETILTSPEPAEDVRETVAAVEKASHLSPALA
RLFAVSTHAEKKHEKTQEPSPVKEVKNPTSSTTVKAPISIEPPGAEEKEAVEEFTLLNDGEVTELEYVEI
ATMLHQILTKLSGSFKRKRKNRFMVLTQNTFYLTQSCIEKYGTQLNAPELFNQLPQYQVTSGAVVNTKCI
AFNIPTLVAASDRAKVDIELIINKLKEVGNL
MNFRALFLSMQRVFGIFSRRENDVSELMMKDAANFSPFAQIIGEQKYTVPDHPNPEVLKFIEYPTRPAGI
QTFNEQSILSLYRDKLHSISMMLAISDGDIREDAYTFTNLVLKPLIEYIRWIHLLPASENHHHNGIGGLL
SHSLEVAMISLKNANHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLSETLTWAPS
SQSLLDWARENNVVEYEIHWRKRIHNQHNIWSSVFLERILDPVCMSFLDRVKKERVYAKMVTALNVYNDG
NDFLSKCVRTSDYYSTGTDLNVLRDPIMGLRSNDAAARAIGTIKHNFTSININNYKTKPMHIIIVNGEVY
LNENAFLDFVLSDFAAHKFNFPQGDAGKTVLVESLVLRGYVEPYDDERVVHYFIPGTYSENEIASIFRNG
IGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRESVTRVTDTVN
DAIENTPQYGLQLVNGPDADSNNIISSENTESITDSLEESGADISNEIFETQVVTAIDTAETVNADEPEQ
VEEHDDRSQIHLVEQLHEMLLSAPLPHHAVINIDSVPYLDLDAAIALIPGIDEAAFCNGPFFQLTYRDGS
LDGMWIVRDVNNLRLIQLGDNCAGMQVSTSEPRNTSSLKSLFDTSMYQPLDIPEAPSVNEAASPPQTPLE
LPQPRLNAPVAEEASSVAEQTNAHSEPDSVIATEYEQYGHLLEETLDSDGEAYSDLIASDSTEAEYPATD
PQSSDFAQLPRETALSVAPGDLDYSEGAIKPPAPDATGKETILTSPEPAEDVRETVAAVEKASHLSPALA
RLFAVSTHAEKKHEKTQEPSPVKEVKNPTSSTTVKAPISIEPPGAEEKEAVEEFTLLNDGEVTELEYVEI
ATMLHQILTKLSGSFKRKRKNRFMVLTQNTFYLTQSCIEKYGTQLNAPELFNQLPQYQVTSGAVVNTKCI
AFNIPTLVAASDRAKVDIELIINKLKEVGNL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A7T0P871 |
T4CP
| ID | 4126 | GenBank | WP_000167418 |
| Name | traD_IAR32_RS23385_p733-MDR |
UniProt ID | _ |
| Length | 694 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 694 a.a. Molecular weight: 77969.74 Da Isoelectric Point: 8.7275
>WP_000167418.1 MULTISPECIES: conjugative transfer system coupling protein TraD [Enterobacterales]
MTKSKRTNLHAQENFYRPILEYRSASVLLICAVIMLVMGFRSDGVNIAPIILYTAVFLLLVCLYRCKTAH
PYLMAHWRVFQRQIMFISLKSLRTINKSNFFSNERKYRQLVQEYKKNNRPVPDRKTYFCNGFEWGPEHAD
RAYQIANLSSDKREIALPFVLSPIARHFETMARSMGGNNAIFAVDRRAPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVIVVIDPKNDAEWRQSLMDEASELGLPFYKFHPAQPSSSVCIDVCNAYTNVSD
LTSRLLSLVSVPGEVNPFVQYAEALVSTVITGLSYTDKKPSIYLIHKNMKSHMSVVNLTIKVMECCFARH
YGPDVWMEKVKYASNDTLQVRFKRLTEWFNAHFLNYEGAEPIEWIDTVGRLVDYSMSDPEHMSKMTAGIM
PLFSRLTEQPLNELLSPSPNTLTSREIVTSDGMFSTGGVLYISLDGLSNPESARAISQLIMSDLTSCAGS
RYNADDGDMSSHSRISIFVDEAHSAINNSMINLLAQGRAAQIALFICTQTISDFIAAANAETANRITGLC
NNYISLRVNDTPTQTLVVENFGKSPISTNMVTYTTGSETTLPHNNFSGSISERKQTTLEESIPKELLGQV
PKFHIVARLQDGRKVVGQIPIAVSEKAMKPNTTLLEMFLKPAGKVTLRQNVGLSYLNKYLRKLH
MTKSKRTNLHAQENFYRPILEYRSASVLLICAVIMLVMGFRSDGVNIAPIILYTAVFLLLVCLYRCKTAH
PYLMAHWRVFQRQIMFISLKSLRTINKSNFFSNERKYRQLVQEYKKNNRPVPDRKTYFCNGFEWGPEHAD
RAYQIANLSSDKREIALPFVLSPIARHFETMARSMGGNNAIFAVDRRAPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVIVVIDPKNDAEWRQSLMDEASELGLPFYKFHPAQPSSSVCIDVCNAYTNVSD
LTSRLLSLVSVPGEVNPFVQYAEALVSTVITGLSYTDKKPSIYLIHKNMKSHMSVVNLTIKVMECCFARH
YGPDVWMEKVKYASNDTLQVRFKRLTEWFNAHFLNYEGAEPIEWIDTVGRLVDYSMSDPEHMSKMTAGIM
PLFSRLTEQPLNELLSPSPNTLTSREIVTSDGMFSTGGVLYISLDGLSNPESARAISQLIMSDLTSCAGS
RYNADDGDMSSHSRISIFVDEAHSAINNSMINLLAQGRAAQIALFICTQTISDFIAAANAETANRITGLC
NNYISLRVNDTPTQTLVVENFGKSPISTNMVTYTTGSETTLPHNNFSGSISERKQTTLEESIPKELLGQV
PKFHIVARLQDGRKVVGQIPIAVSEKAMKPNTTLLEMFLKPAGKVTLRQNVGLSYLNKYLRKLH
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 240056..249224
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| IAR32_RS23705 (IAR32_23530) | 235593..235793 | - | 201 | WP_000357613 | hypothetical protein | - |
| IAR32_RS23710 (IAR32_23535) | 235811..236887 | - | 1077 | WP_001015069 | hypothetical protein | - |
| IAR32_RS23715 (IAR32_23540) | 237422..238297 | + | 876 | WP_001282731 | RepB family plasmid replication initiator protein | - |
| IAR32_RS23720 (IAR32_23545) | 238744..239088 | - | 345 | WP_153253765 | hypothetical protein | - |
| IAR32_RS23725 (IAR32_23550) | 239651..240004 | + | 354 | WP_038988950 | hypothetical protein | - |
| IAR32_RS23730 (IAR32_23555) | 240056..240373 | + | 318 | WP_000203871 | type IV conjugative transfer system protein TraL | traL |
| IAR32_RS23735 (IAR32_23560) | 240385..241170 | + | 786 | WP_000771920 | TraE/TraK family type IV conjugative transfer system protein | traE |
| IAR32_RS23740 (IAR32_23565) | 241173..242405 | + | 1233 | WP_000202110 | type-F conjugative transfer system secretin TraK | traK |
| IAR32_RS23745 (IAR32_23570) | 242407..242847 | + | 441 | WP_001224093 | plasmid transfer protein HtdO | - |
| IAR32_RS23750 (IAR32_23575) | 242837..244195 | + | 1359 | WP_000351840 | IncHI-type conjugal transfer protein TrhB | traB |
| IAR32_RS23755 (IAR32_23580) | 244204..244716 | + | 513 | WP_000429933 | hypothetical protein | - |
| IAR32_RS23760 (IAR32_23585) | 244822..245574 | + | 753 | WP_010892304 | protein-disulfide isomerase HtdT | - |
| IAR32_RS23765 (IAR32_23590) | 245584..246534 | + | 951 | WP_001022266 | TraV family lipoprotein | traV |
| IAR32_RS23770 (IAR32_23595) | 246543..249224 | + | 2682 | WP_001255540 | IncHI-type conjugal transfer ATPase TrhC | virb4 |
| IAR32_RS23775 (IAR32_23600) | 249894..250322 | + | 429 | WP_124061973 | hypothetical protein | - |
| IAR32_RS23780 (IAR32_23605) | 251000..251173 | + | 174 | WP_005046574 | hypothetical protein | - |
| IAR32_RS23785 | 251311..251493 | + | 183 | WP_071925124 | hypothetical protein | - |
| IAR32_RS23790 (IAR32_23610) | 251524..251757 | + | 234 | WP_032489575 | hypothetical protein | - |
| IAR32_RS23795 (IAR32_23615) | 252071..252217 | + | 147 | WP_005046583 | hypothetical protein | - |
Host bacterium
| ID | 6573 | GenBank | NZ_CP060847 |
| Plasmid name | p733-MDR | Incompatibility group | IncFIA |
| Plasmid size | 253764 bp | Coordinate of oriT [Strand] | 178959..179243 [-] |
| Host baterium | Salmonella enterica strain 733 |
Cargo genes
| Drug resistance gene | qnrS1, blaTEM-1B, sul3, ant(3'')-Ia, cmlA1, aadA2, dfrA12, floR, sul2, tet(A), tet(M) |
| Virulence gene | - |
| Metal resistance gene | silE, silS, silR, silC, silF, silB, silA, silP, pcoA, pcoB, pcoC, pcoE |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |