Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 105528 |
| Name | oriT_pSHE-CTX-M |
| Organism | Shewanella bicestrii strain JAB-1 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP022359 (190763..190913 [+], 151 nt) |
| oriT length | 151 nt |
| IRs (inverted repeats) | 93..98, 106..111 (TGGCCT..AGGCCA) 12..17, 24..29 (AACCCT..AGGGTT) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 151 nt
>oriT_pSHE-CTX-M
TGGTCGATGTGAACCCTTTCGACAGGGTTATGAATGAATTGAAAAGTCGTGGCCGCAAGAACGCTCACATCCTGAGCATCCTCCAATTCGACTGGCCTGCATCGGAGGCCATCATCGAGAAGCTGAGCTGCTACATCACAGACGGGATTAA
TGGTCGATGTGAACCCTTTCGACAGGGTTATGAATGAATTGAAAAGTCGTGGCCGCAAGAACGCTCACATCCTGAGCATCCTCCAATTCGACTGGCCTGCATCGGAGGCCATCATCGAGAAGCTGAGCTGCTACATCACAGACGGGATTAA
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file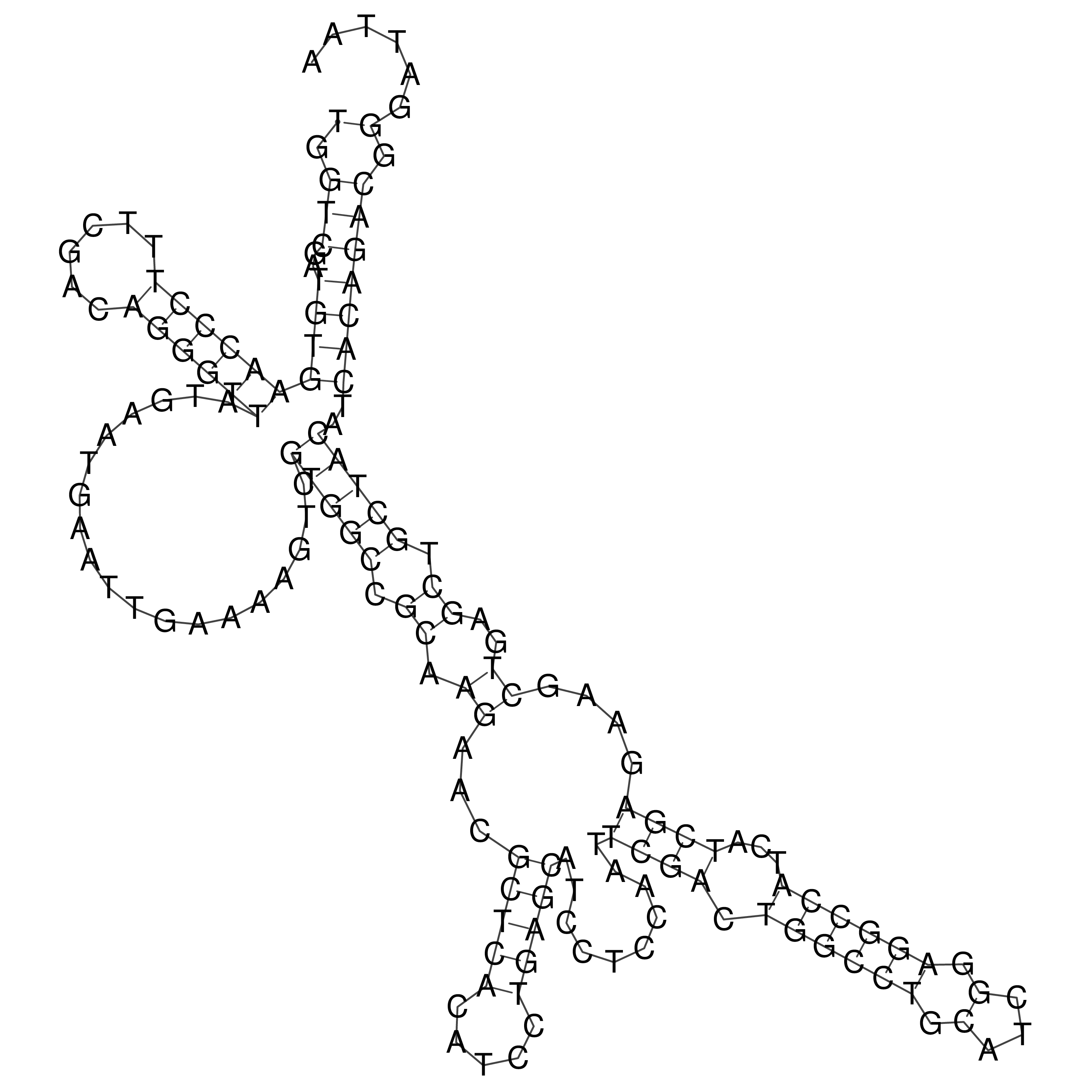
Relaxase
| ID | 3768 | GenBank | WP_001337757 |
| Name | mobH_CF168_RS22160_pSHE-CTX-M |
UniProt ID | A0A709FRI2 |
| Length | 992 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 992 a.a. Molecular weight: 109581.85 Da Isoelectric Point: 4.8956
>WP_001337757.1 MULTISPECIES: MobH family relaxase [Gammaproteobacteria]
MLKALNKLFGGRSGVIETAPSARVLPLKDVEDEEIPRYPPFAKGLPVAPLDKILATQAELIEKVRNSLGF
TVDDFNRLVLPVIQRYAAFVHLLPASESHHHRGAGGLFRHGLEVAFWAAQASESVIFSIEGTPRERRDNE
PRWRLASCFSGLLHDVGKPLSDVSITDKDGSITWNPYSESLHDWAHRHEIDRYFIRWRDKRHKRHEQFSL
LAVDRIIPAETREFLSKSGPSIMEAMLEAISGTSVNQPVTKLMLRADQESVSRDLRQSRLDVDEFSYGVP
VERYVFDAIRRLVKTGKWKVNEPGAKVWHLNQGVFIAWKQLGDLYDLISHDKIPGIPRDPDTLADILIER
GFAVPNTVQEKGERAYYRYWEVLPEMLQEAAGSVKILMLRLESNDLVFTTEPPAAVAAEVVGDVEDAEIE
FVDPEEADDGDDQEEGEAALNDDMLAAEQEAEKALAGLGFGDAMEMLKSTSDAVEEKPEQKDAGPTESSK
PDAGKKGKPQSKPGKAKPKSDTEKQPHKPEAKEDLSPQDIAKNAPPLANDNPLQALKDVGGGLGDIDFPF
DAFNASAETTSTDATNSEIPDVAMPGKQEEQPKQDFVPQEQNSLQGDDFPMFGGSDEPPSWAIEPLPMLT
DAPEQPTHTPEMPHTDNVNQHEKDAKTLLVEMLSGYGEASALLEQAIMPVLEGKTTLGEVLCLMKGQAVI
LYPDGARSLGAPSEVLSKLSHANAIVPDPIMPGRKVRDFSGVKAIVLAEQLSDAVVAAIKDAEASMGGYQ
DAFELVSPPGLDASKNKSAPKQQSRKKAQQQKPEVNAGKASPEQKAKGKDSQPQPKEKKVDVTSPVEEQQ
RKPVQEKQNVARLPKREAQPVAPEPKVEREKELGHVEVREREDPEVREFEPPKAKTNPKDINAEDFLPSG
VTPQKALQMLKDMIQKRSGRWLVTPVLEEDGCLVTSDKAFDMIAGENIGISKHILCGMLSRAQRRPLLKK
RQGKLYLEVNET
MLKALNKLFGGRSGVIETAPSARVLPLKDVEDEEIPRYPPFAKGLPVAPLDKILATQAELIEKVRNSLGF
TVDDFNRLVLPVIQRYAAFVHLLPASESHHHRGAGGLFRHGLEVAFWAAQASESVIFSIEGTPRERRDNE
PRWRLASCFSGLLHDVGKPLSDVSITDKDGSITWNPYSESLHDWAHRHEIDRYFIRWRDKRHKRHEQFSL
LAVDRIIPAETREFLSKSGPSIMEAMLEAISGTSVNQPVTKLMLRADQESVSRDLRQSRLDVDEFSYGVP
VERYVFDAIRRLVKTGKWKVNEPGAKVWHLNQGVFIAWKQLGDLYDLISHDKIPGIPRDPDTLADILIER
GFAVPNTVQEKGERAYYRYWEVLPEMLQEAAGSVKILMLRLESNDLVFTTEPPAAVAAEVVGDVEDAEIE
FVDPEEADDGDDQEEGEAALNDDMLAAEQEAEKALAGLGFGDAMEMLKSTSDAVEEKPEQKDAGPTESSK
PDAGKKGKPQSKPGKAKPKSDTEKQPHKPEAKEDLSPQDIAKNAPPLANDNPLQALKDVGGGLGDIDFPF
DAFNASAETTSTDATNSEIPDVAMPGKQEEQPKQDFVPQEQNSLQGDDFPMFGGSDEPPSWAIEPLPMLT
DAPEQPTHTPEMPHTDNVNQHEKDAKTLLVEMLSGYGEASALLEQAIMPVLEGKTTLGEVLCLMKGQAVI
LYPDGARSLGAPSEVLSKLSHANAIVPDPIMPGRKVRDFSGVKAIVLAEQLSDAVVAAIKDAEASMGGYQ
DAFELVSPPGLDASKNKSAPKQQSRKKAQQQKPEVNAGKASPEQKAKGKDSQPQPKEKKVDVTSPVEEQQ
RKPVQEKQNVARLPKREAQPVAPEPKVEREKELGHVEVREREDPEVREFEPPKAKTNPKDINAEDFLPSG
VTPQKALQMLKDMIQKRSGRWLVTPVLEEDGCLVTSDKAFDMIAGENIGISKHILCGMLSRAQRRPLLKK
RQGKLYLEVNET
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A709FRI2 |
T4CP
| ID | 3759 | GenBank | WP_000637386 |
| Name | traC_CF168_RS21035_pSHE-CTX-M |
UniProt ID | _ |
| Length | 815 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 815 a.a. Molecular weight: 92590.87 Da Isoelectric Point: 5.9222
>WP_000637386.1 MULTISPECIES: type IV secretion system protein TraC [Gammaproteobacteria]
MIVTIKKKLEETLIPEHLRAAGIIPVLAYDEDDHVFLMDDHSAGFGFMCEPLCGADEKVQERMNGFLNQE
FPSKTTLQFVLFRSPDINQEMYRMMGLRDGFRHELLTSVIKERINFLQHHTTERIFAKTNKGIYDNGLIQ
DLKLFVTCKVPIKNNNPTESELQQLAQLRTKVESSLQTVGLRPRTMTAVNYIRIMSTILNWGPDASWRHD
SVDWEMDKPICEQIFDYGTDVEVSKNGIRLGDYHAKVMSAKKLPDVFYFGDALTYAGDLSGGNSSIKENY
MVVTNVFFPEAESTKNTLERKRQFTVNQAYGPMLKFVPVLADKKESFDTLYESMKEGAKPVKITYSVVLF
APTKERVEAAAMAARNIWRESRFELMEDKFVALPMFLNCLPFCTDRDAVRDLFRYKTMTTEQAAVVLPVF
GEWKGTGTYHAALISRNGQLMSLSLHDSNTNKNLVIAAESGSGKSFLTNELIFSYLSEGAQVWVIDAGKS
YQKLSEMLNGDFVHFEEGTHVCLNPFELIQNYEDEEDAIVSLVCAMASAKGLLDEWQISALKQVLSRLWE
EKGKEMKVDDIAERCLEEENDQRLKDIGQQLYAFTSKGSYGKYFSRKNNVSFQNQFTVLELDELQGRKHL
RQVVLLQLIYQIQQEVFLGERNRKKVVIVDEAWDLLKEGEVSVFMEHAYRKFRKYGGSVVIATQSINDLY
ENAVGRAIAENSASMYLLGQTEETVESVKRSGRLTLSEGGFHTLKTVHTIQGVYSEIFIKSKSGMGVGRL
IVGDFQKLLYSTDPVDVNAIDQFVKQGMSIPEAIKAVMRSRQQAA
MIVTIKKKLEETLIPEHLRAAGIIPVLAYDEDDHVFLMDDHSAGFGFMCEPLCGADEKVQERMNGFLNQE
FPSKTTLQFVLFRSPDINQEMYRMMGLRDGFRHELLTSVIKERINFLQHHTTERIFAKTNKGIYDNGLIQ
DLKLFVTCKVPIKNNNPTESELQQLAQLRTKVESSLQTVGLRPRTMTAVNYIRIMSTILNWGPDASWRHD
SVDWEMDKPICEQIFDYGTDVEVSKNGIRLGDYHAKVMSAKKLPDVFYFGDALTYAGDLSGGNSSIKENY
MVVTNVFFPEAESTKNTLERKRQFTVNQAYGPMLKFVPVLADKKESFDTLYESMKEGAKPVKITYSVVLF
APTKERVEAAAMAARNIWRESRFELMEDKFVALPMFLNCLPFCTDRDAVRDLFRYKTMTTEQAAVVLPVF
GEWKGTGTYHAALISRNGQLMSLSLHDSNTNKNLVIAAESGSGKSFLTNELIFSYLSEGAQVWVIDAGKS
YQKLSEMLNGDFVHFEEGTHVCLNPFELIQNYEDEEDAIVSLVCAMASAKGLLDEWQISALKQVLSRLWE
EKGKEMKVDDIAERCLEEENDQRLKDIGQQLYAFTSKGSYGKYFSRKNNVSFQNQFTVLELDELQGRKHL
RQVVLLQLIYQIQQEVFLGERNRKKVVIVDEAWDLLKEGEVSVFMEHAYRKFRKYGGSVVIATQSINDLY
ENAVGRAIAENSASMYLLGQTEETVESVKRSGRLTLSEGGFHTLKTVHTIQGVYSEIFIKSKSGMGVGRL
IVGDFQKLLYSTDPVDVNAIDQFVKQGMSIPEAIKAVMRSRQQAA
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
| ID | 3760 | GenBank | WP_000178856 |
| Name | traD_CF168_RS22165_pSHE-CTX-M |
UniProt ID | _ |
| Length | 621 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 621 a.a. Molecular weight: 69197.66 Da Isoelectric Point: 6.9403
>WP_000178856.1 MULTISPECIES: conjugative transfer system coupling protein TraD [Gammaproteobacteria]
MTMSYDPLAYEMPWRPNYEKNAVAGWLAASGAALAVEQVSTMPPEPFYWMTGICGVMAMARLPKAIKLHL
LQKHLKGRDLEFISIAELQKYIKDTPDDMWLGSGFLWENRHAQRVFEILKRDWTSIVGRESTVKKVVRKI
QGKKKELPIGQPWIHGVEPKEEKLMQPLKHTEGHSLIVGTTGSGKTRMFDILISQAILRGEAVIIIDPKG
DKEMRDNARRACEAMGQPERFVSFHPAFPEESVRIDPLRNFTRVTEIASRLAALIPSEAGADPFKSFGWQ
ALNNIAQGLVITHDRPNLTKLRRFLEGGAAGLVIKAVQAYSERVMPDWEAEAAAYLEKAKNGSREKIAFA
LMKFYYDIIQPEHPNSDLEGLLSMFQHDQTHFSKMVANLLPIMNMLTSGELGPLLSPDSSDLSDERQITD
SAKIINNAQVAYLGLDSLTDNMVGSAMGSIFLSDLTAVAGDRYNYGVNNRPVNIFVDEAAEVINDPFIQL
LNKGRGAKLRLFVATQTFADFAARLGSKDKALQVLGNINNTFALRIVDGETQEYIADNLPKTRLKYVMRT
QGQNSDGKEPIMHGGNQGERLMEEEADLFPAQLLGMLPNLEYIAKISGGTIVKGRLPILTQ
MTMSYDPLAYEMPWRPNYEKNAVAGWLAASGAALAVEQVSTMPPEPFYWMTGICGVMAMARLPKAIKLHL
LQKHLKGRDLEFISIAELQKYIKDTPDDMWLGSGFLWENRHAQRVFEILKRDWTSIVGRESTVKKVVRKI
QGKKKELPIGQPWIHGVEPKEEKLMQPLKHTEGHSLIVGTTGSGKTRMFDILISQAILRGEAVIIIDPKG
DKEMRDNARRACEAMGQPERFVSFHPAFPEESVRIDPLRNFTRVTEIASRLAALIPSEAGADPFKSFGWQ
ALNNIAQGLVITHDRPNLTKLRRFLEGGAAGLVIKAVQAYSERVMPDWEAEAAAYLEKAKNGSREKIAFA
LMKFYYDIIQPEHPNSDLEGLLSMFQHDQTHFSKMVANLLPIMNMLTSGELGPLLSPDSSDLSDERQITD
SAKIINNAQVAYLGLDSLTDNMVGSAMGSIFLSDLTAVAGDRYNYGVNNRPVNIFVDEAAEVINDPFIQL
LNKGRGAKLRLFVATQTFADFAARLGSKDKALQVLGNINNTFALRIVDGETQEYIADNLPKTRLKYVMRT
QGQNSDGKEPIMHGGNQGERLMEEEADLFPAQLLGMLPNLEYIAKISGGTIVKGRLPILTQ
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 906..20700
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| CF168_RS22410 | 241..384 | + | 144 | WP_001275801 | hypothetical protein | - |
| CF168_RS20990 (CF168_20990) | 396..761 | + | 366 | WP_001052531 | hypothetical protein | - |
| CF168_RS20995 (CF168_20995) | 906..1187 | + | 282 | WP_000805625 | type IV conjugative transfer system protein TraL | traL |
| CF168_RS21000 (CF168_21000) | 1184..1810 | + | 627 | WP_001049716 | TraE/TraK family type IV conjugative transfer system protein | traE |
| CF168_RS21005 (CF168_21005) | 1794..2711 | + | 918 | WP_000794249 | type-F conjugative transfer system secretin TraK | traK |
| CF168_RS21010 (CF168_21010) | 2711..4024 | + | 1314 | WP_024131605 | TraB/VirB10 family protein | traB |
| CF168_RS21015 (CF168_21015) | 4021..4599 | + | 579 | WP_000793435 | type IV conjugative transfer system lipoprotein TraV | traV |
| CF168_RS21020 (CF168_21020) | 4603..4995 | + | 393 | WP_000479535 | TraA family conjugative transfer protein | - |
| CF168_RS21025 (CF168_21025) | 5196..10727 | + | 5532 | WP_000606833 | Ig-like domain-containing protein | - |
| CF168_RS21030 (CF168_21030) | 10876..11583 | + | 708 | WP_001259347 | DsbC family protein | - |
| CF168_RS21035 (CF168_21035) | 11580..14027 | + | 2448 | WP_000637386 | type IV secretion system protein TraC | virb4 |
| CF168_RS21040 (CF168_21040) | 14042..14359 | + | 318 | WP_000351984 | hypothetical protein | - |
| CF168_RS21045 (CF168_21045) | 14356..14886 | + | 531 | WP_001010738 | S26 family signal peptidase | - |
| CF168_RS21050 (CF168_21050) | 14849..16114 | + | 1266 | WP_000621288 | TrbC family F-type conjugative pilus assembly protein | traW |
| CF168_RS21055 (CF168_21055) | 16111..16782 | + | 672 | WP_001337754 | EAL domain-containing protein | - |
| CF168_RS21060 (CF168_21060) | 16782..17786 | + | 1005 | WP_043940352 | TraU family protein | traU |
| CF168_RS21065 (CF168_21065) | 17902..20700 | + | 2799 | WP_001256487 | conjugal transfer mating pair stabilization protein TraN | traN |
| CF168_RS21070 (CF168_21070) | 20739..21599 | - | 861 | WP_000709517 | hypothetical protein | - |
| CF168_RS21075 (CF168_21075) | 21722..22363 | - | 642 | WP_000796664 | hypothetical protein | - |
| CF168_RS21080 (CF168_21080) | 22657..22977 | + | 321 | WP_000547566 | hypothetical protein | - |
| CF168_RS21085 (CF168_21085) | 23281..23466 | + | 186 | WP_001186917 | hypothetical protein | - |
| CF168_RS21090 (CF168_21090) | 23686..24654 | + | 969 | WP_000085162 | AAA family ATPase | - |
| CF168_RS21095 (CF168_21095) | 24665..25573 | + | 909 | WP_000739139 | hypothetical protein | - |
Host bacterium
| ID | 5966 | GenBank | NZ_CP022359 |
| Plasmid name | pSHE-CTX-M | Incompatibility group | IncA/C2 |
| Plasmid size | 193338 bp | Coordinate of oriT [Strand] | 190763..190913 [+] |
| Host baterium | Shewanella bicestrii strain JAB-1 |
Cargo genes
| Drug resistance gene | blaCTX-M-15, aac(3)-IIa, blaOXA-1, aac(6')-Ib-cr, tet(A), blaSHV-12, aph(6)-Id, aph(3'')-Ib, sul2 |
| Virulence gene | - |
| Metal resistance gene | merR, merT, merP, merC, merA, merD, merE |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |