Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 105287 |
| Name | oriT_pRHBSTW-00355_4 |
| Organism | Citrobacter freundii strain RHBSTW-00355 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP056589 (12123..12212 [-], 90 nt) |
| oriT length | 90 nt |
| IRs (inverted repeats) | 34..41, 44..51 (AGTGCGAT..ATCGCACT) 20..25, 39..44 (TAAATC..GATTTA) |
| Location of nic site | 62..63 |
| Conserved sequence flanking the nic site |
GGTGTATAGC |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 90 nt
>oriT_pRHBSTW-00355_4
TATTTCATTTTTTGTTTTTTAAATCAGTTAGTTAGTGCGATTTATCGCACTGCGTTAGGTGTATAGCAGGTTAAGGGCTAAAAAATAATC
TATTTCATTTTTTGTTTTTTAAATCAGTTAGTTAGTGCGATTTATCGCACTGCGTTAGGTGTATAGCAGGTTAAGGGCTAAAAAATAATC
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file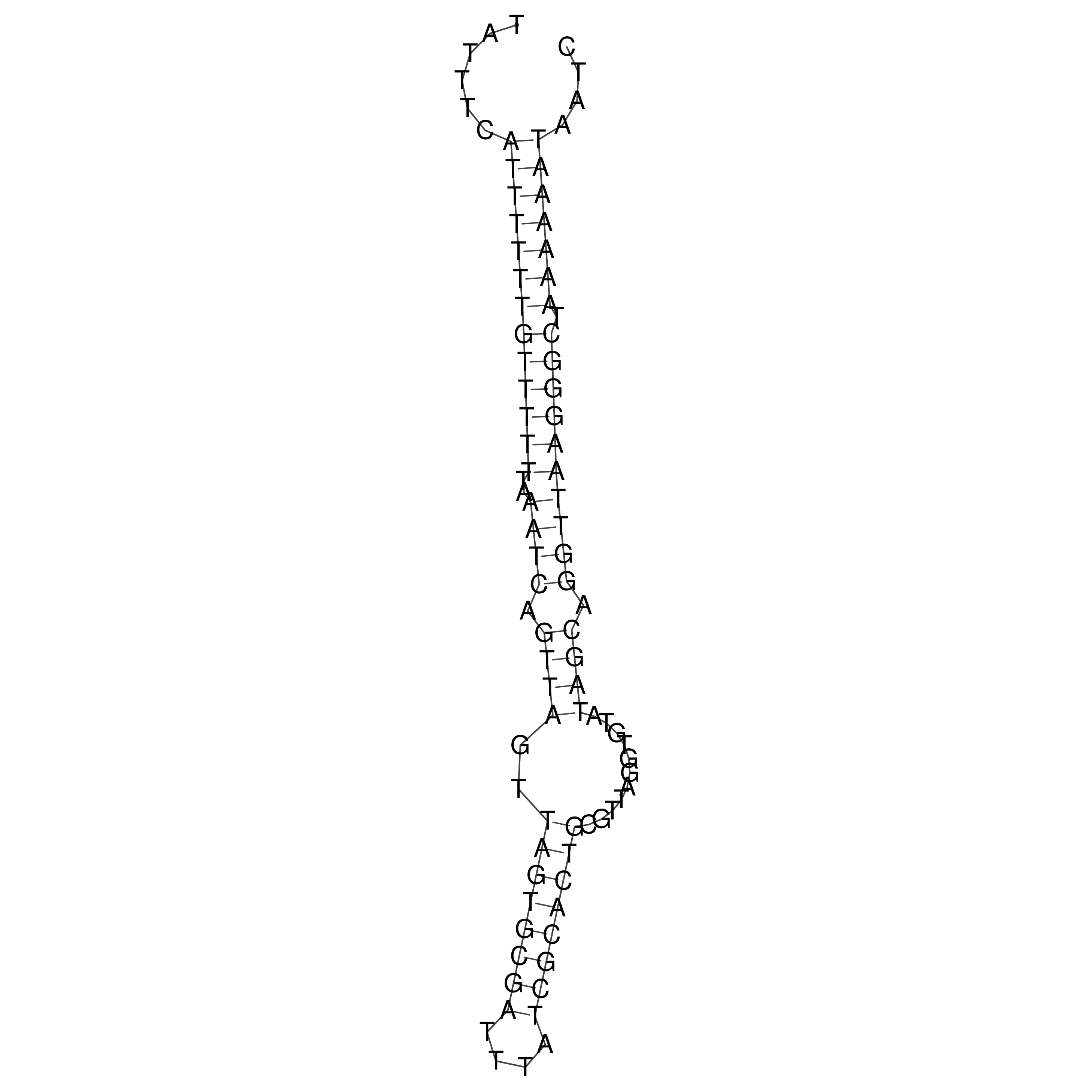
Relaxase
| ID | 3618 | GenBank | WP_050136475 |
| Name | mobF_HV173_RS26870_pRHBSTW-00355_4 |
UniProt ID | _ |
| Length | 1072 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1072 a.a. Molecular weight: 118873.51 Da Isoelectric Point: 6.4635
>WP_050136475.1 MULTISPECIES: MobF family relaxase [Enterobacterales]
MLDITTITRIDIASVVGYYADAKDDYYSKDSSFTAWHGQGAEALGLSGEVSSQRFKELLVGDIDPFTQMK
RSSGDATKERLGYDLTFSAPKGVSMQALIHGDKAIIAAHEKAVAAAVQEAEKLAQARTTHAGKTITQNTG
NLVVASFRHETSRALDPELHTHAFVMNMTQRGDGQWRALKNDELMRAKMHLGDVYKQELALELTKAGYEL
RYNNKNNTFDMAHFTDEQIRGFSRRSAQIEEGLAAMGLTRETADAQTKSRVSMATRDKKTDDYSREDIHK
EWVNRANELGIDFNDLSWAGKGKDGPQTSMNTVPNFTSPEVKADRAIQFALKSLSERDASFDQSRLLAVA
NKKAIGQASKQDVEAAYYRAVAKGAIIEGEARYTTTTDQRSKDAPPLPAMTKAEWQAQLTDKKGMNDSQA
KALVAEGIKSGRLKKTSHRVTTVEGIRVEKAVLNIEARGRGKVIAALTSEQVTAQLVGKTLKAEQQKSVV
DICTTTDRFVAAHGFAGTGKSYMTMSAKAVLESQGYNVTALAPYGTQKKSLEDEGMPARTVAAFVKAKDK
KIDEKSVVFIDEAGVIPARQMKVLMETIEKAGARAVFLGDTSQTKAVEAGKPFEQLISAGMQTSYMKDIQ
RQKNDVLLEAVKLAAEGHISGSLARLSNISAEKDQDKRLEAVANRWLSFSPEQREGTLIISGTNESRVIL
NSQIREALQLQGKGVEVNFLERVDSTQAERRDSKYYQVGQIIIPEKDYKNGLQRGESYRVLDTGPGNLLT
VAGSDGQKISFSPRTHKNLSVYTSVAAELAVGDKVMVTRNDKTLDVANGDRFTVTGTTEKGTFTLTNSKG
REIELDQKQAAYLSYAYATTVHKAQGLTCDRVLFNIDTRSLTTSKDVFYVGISRARHEVEIFTDDKKRLL
SSASRNSPKTTASEIDRFLGMEERYKDVNRSYGHSDGQQEKSYGHDAGHQTEEKENTMKNATEAAHDGPD
TSGTENTSQDLRQREQEEVNAAIRYEQRQEATPEMEANPYHDYERYADEGPDYGDMQDYSTYEEYERARD
VELPQHEHENHQHDHDKEGYGL
MLDITTITRIDIASVVGYYADAKDDYYSKDSSFTAWHGQGAEALGLSGEVSSQRFKELLVGDIDPFTQMK
RSSGDATKERLGYDLTFSAPKGVSMQALIHGDKAIIAAHEKAVAAAVQEAEKLAQARTTHAGKTITQNTG
NLVVASFRHETSRALDPELHTHAFVMNMTQRGDGQWRALKNDELMRAKMHLGDVYKQELALELTKAGYEL
RYNNKNNTFDMAHFTDEQIRGFSRRSAQIEEGLAAMGLTRETADAQTKSRVSMATRDKKTDDYSREDIHK
EWVNRANELGIDFNDLSWAGKGKDGPQTSMNTVPNFTSPEVKADRAIQFALKSLSERDASFDQSRLLAVA
NKKAIGQASKQDVEAAYYRAVAKGAIIEGEARYTTTTDQRSKDAPPLPAMTKAEWQAQLTDKKGMNDSQA
KALVAEGIKSGRLKKTSHRVTTVEGIRVEKAVLNIEARGRGKVIAALTSEQVTAQLVGKTLKAEQQKSVV
DICTTTDRFVAAHGFAGTGKSYMTMSAKAVLESQGYNVTALAPYGTQKKSLEDEGMPARTVAAFVKAKDK
KIDEKSVVFIDEAGVIPARQMKVLMETIEKAGARAVFLGDTSQTKAVEAGKPFEQLISAGMQTSYMKDIQ
RQKNDVLLEAVKLAAEGHISGSLARLSNISAEKDQDKRLEAVANRWLSFSPEQREGTLIISGTNESRVIL
NSQIREALQLQGKGVEVNFLERVDSTQAERRDSKYYQVGQIIIPEKDYKNGLQRGESYRVLDTGPGNLLT
VAGSDGQKISFSPRTHKNLSVYTSVAAELAVGDKVMVTRNDKTLDVANGDRFTVTGTTEKGTFTLTNSKG
REIELDQKQAAYLSYAYATTVHKAQGLTCDRVLFNIDTRSLTTSKDVFYVGISRARHEVEIFTDDKKRLL
SSASRNSPKTTASEIDRFLGMEERYKDVNRSYGHSDGQQEKSYGHDAGHQTEEKENTMKNATEAAHDGPD
TSGTENTSQDLRQREQEEVNAAIRYEQRQEATPEMEANPYHDYERYADEGPDYGDMQDYSTYEEYERARD
VELPQHEHENHQHDHDKEGYGL
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4CP
| ID | 3584 | GenBank | WP_040223430 |
| Name | t4cp2_HV173_RS26865_pRHBSTW-00355_4 |
UniProt ID | _ |
| Length | 510 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 510 a.a. Molecular weight: 57917.09 Da Isoelectric Point: 9.6871
>WP_040223430.1 MULTISPECIES: type IV secretion system DNA-binding domain-containing protein [Enterobacterales]
MTEQERGLIFLCSLLGPIPFVWFITAKFTYGINSETAHYLVPYLLKNTFNLWPLWAAIIAGVVIGLCGFI
GFIIYDKGRVYKGPRFKRILRGTKLARASTLAEKTKERGKKQLMIAGVPVPTEYETLHFSIGGATGTGKS
TVIKEAIYSSLKRGTDKRIVLDPNGEYVETFYRPGDVILSPYDKRSPGWSMFNEIRDDFDYDLYAVSIIQ
ESKERSAEEWYRYGRLIYREVSKKLLTLKRNPTMEEVMHWCTIVENDKLKEFVKGTPAEALFIKGAERAT
GSARFVLGDKLAPHLKMPAGDFSLRDWLEDDKPGTLFITWQEQMKKSLNPLISCWLDIMFSSLLGMGKRD
QRTLFFIDELESLEYLPNLQDLLTKGRKSGASVWAGYQTYSQLIERYGINIAETLLGSMRSGIVMGSSRL
GKETLEQMSRALGEIEGEVEHKQGNPFQAGIGNRPRIETRTVRAVTPTEISELKNLTGYVYFPGDLPVGK
FKTKYVKYTRKNPVPGIIMR
MTEQERGLIFLCSLLGPIPFVWFITAKFTYGINSETAHYLVPYLLKNTFNLWPLWAAIIAGVVIGLCGFI
GFIIYDKGRVYKGPRFKRILRGTKLARASTLAEKTKERGKKQLMIAGVPVPTEYETLHFSIGGATGTGKS
TVIKEAIYSSLKRGTDKRIVLDPNGEYVETFYRPGDVILSPYDKRSPGWSMFNEIRDDFDYDLYAVSIIQ
ESKERSAEEWYRYGRLIYREVSKKLLTLKRNPTMEEVMHWCTIVENDKLKEFVKGTPAEALFIKGAERAT
GSARFVLGDKLAPHLKMPAGDFSLRDWLEDDKPGTLFITWQEQMKKSLNPLISCWLDIMFSSLLGMGKRD
QRTLFFIDELESLEYLPNLQDLLTKGRKSGASVWAGYQTYSQLIERYGINIAETLLGSMRSGIVMGSSRL
GKETLEQMSRALGEIEGEVEHKQGNPFQAGIGNRPRIETRTVRAVTPTEISELKNLTGYVYFPGDLPVGK
FKTKYVKYTRKNPVPGIIMR
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Host bacterium
| ID | 5725 | GenBank | NZ_CP056589 |
| Plasmid name | pRHBSTW-00355_4 | Incompatibility group | IncN3 |
| Plasmid size | 63589 bp | Coordinate of oriT [Strand] | 12123..12212 [-] |
| Host baterium | Citrobacter freundii strain RHBSTW-00355 |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |