Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 105134 |
| Name | oriT_pRHB03-C23_2 |
| Organism | Escherichia fergusonii strain RHB03-C23 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP055807 (41032..41380 [-], 349 nt) |
| oriT length | 349 nt |
| IRs (inverted repeats) | 134..140, 154..160 (AAAATAT..ATATTTT) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 349 nt
>oriT_pRHB03-C23_2
ACTGCGATGCAGGATAGGGCAAACGCCGCAAAATGACGTCTCTGACGCCATTCCGCAACGTTTGCCCGGTGACCTACAGTCGGTTCTTGTTGGAGAGTTTTATGAAATATTTGGTCAAAGAGTTTATTAACGAAAAATATACTAAGGCTGTTAATATTTTAAAGGATAACCTTAAAGAACACTATCATGTTTTTTATTGTGTGAGATTAAGTGAGATTCTTTTTCCTGCCAGTGAATATGGAACTGAACAGTTTTTTAGTGATTTTGAAAAGATAAACTCCATTTCATTACCTGTCTTAGTGTTTGATCTCAAAGAGGGTGTTCCTGTGATTGTCATTAGTCTTGATGA
ACTGCGATGCAGGATAGGGCAAACGCCGCAAAATGACGTCTCTGACGCCATTCCGCAACGTTTGCCCGGTGACCTACAGTCGGTTCTTGTTGGAGAGTTTTATGAAATATTTGGTCAAAGAGTTTATTAACGAAAAATATACTAAGGCTGTTAATATTTTAAAGGATAACCTTAAAGAACACTATCATGTTTTTTATTGTGTGAGATTAAGTGAGATTCTTTTTCCTGCCAGTGAATATGGAACTGAACAGTTTTTTAGTGATTTTGAAAAGATAAACTCCATTTCATTACCTGTCTTAGTGTTTGATCTCAAAGAGGGTGTTCCTGTGATTGTCATTAGTCTTGATGA
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file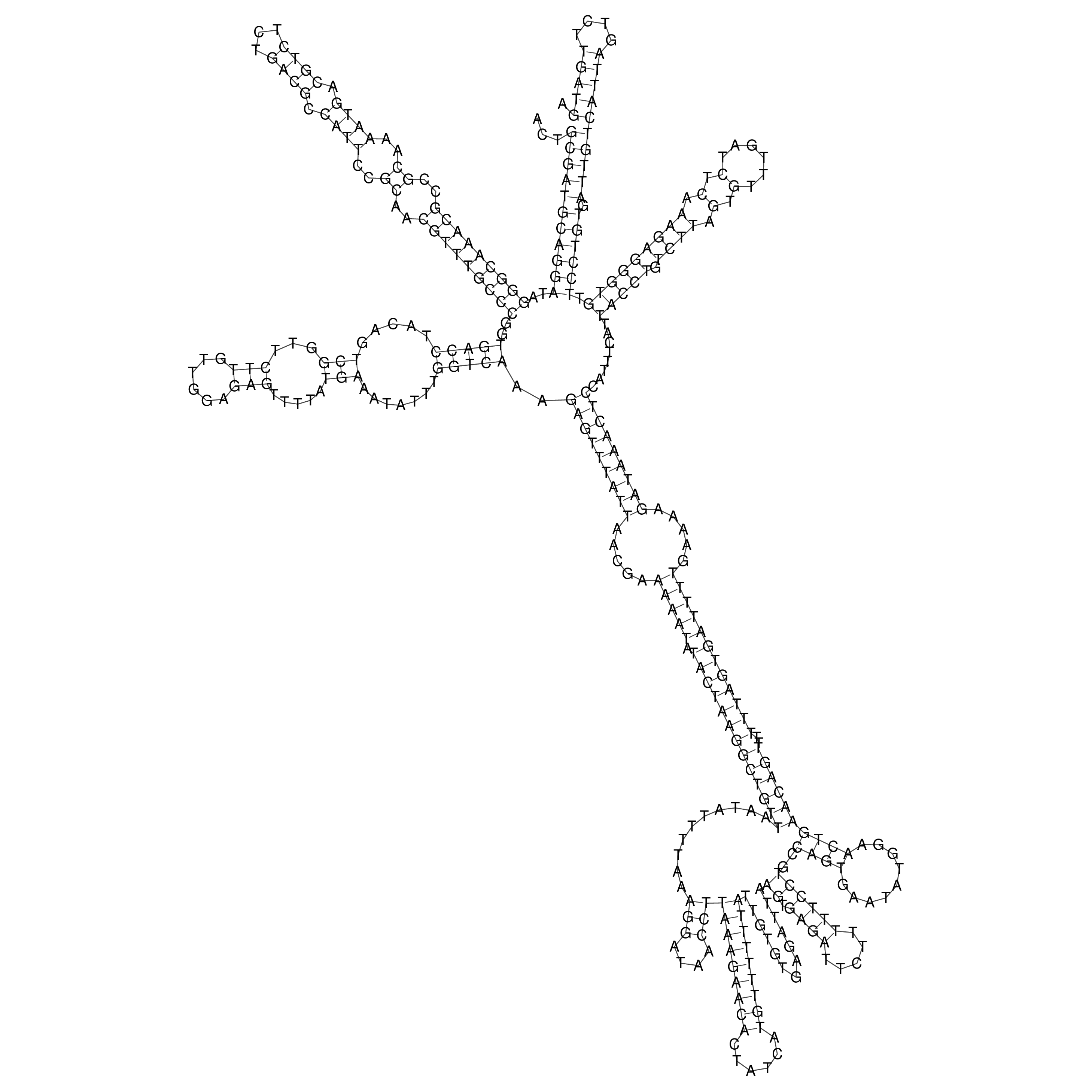
Relaxase
| ID | 3557 | GenBank | WP_015056831 |
| Name | mobP1_HVV65_RS23155_pRHB03-C23_2 |
UniProt ID | H9ZMK2 |
| Length | 388 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 388 a.a. Molecular weight: 44539.89 Da Isoelectric Point: 10.5067
>WP_015056831.1 MULTISPECIES: MobP1 family relaxase [Escherichia]
MGVYVDKEYRVKRKSSENGRKSAFAHKVKNGGKNYSRNVQERINRKGASKEVVVKISGGAVTRQGIRNSI
DYMSRESELPVMSESGRVWTGDEILEAKDHMIDRANDPQHMMNDKGKENKKITQNIVFSPPVSAKVKPED
LLESVRKTMQKKYPNHRFVLGYHCDKKEHPHVHVVFRIRDNDGKRADIRKKDLREIRTGFCEELKLKGYD
VKATHKQQHGLNQSVKDAHNTAPKRQKGVYEVVDVGYDHYQNDKTKSKQHFIKLKTLNKGVEKTYWGADF
GDLCSRESVKAGDLVRLKKLGQKEVKIPALDKNGVQHGWKTVHRNEWQLENLGVKGVDRTPSASKELVLN
SPDMLLKQQQRMAQFTQQKASTLQSEQKLKTGIKFWSL
MGVYVDKEYRVKRKSSENGRKSAFAHKVKNGGKNYSRNVQERINRKGASKEVVVKISGGAVTRQGIRNSI
DYMSRESELPVMSESGRVWTGDEILEAKDHMIDRANDPQHMMNDKGKENKKITQNIVFSPPVSAKVKPED
LLESVRKTMQKKYPNHRFVLGYHCDKKEHPHVHVVFRIRDNDGKRADIRKKDLREIRTGFCEELKLKGYD
VKATHKQQHGLNQSVKDAHNTAPKRQKGVYEVVDVGYDHYQNDKTKSKQHFIKLKTLNKGVEKTYWGADF
GDLCSRESVKAGDLVRLKKLGQKEVKIPALDKNGVQHGWKTVHRNEWQLENLGVKGVDRTPSASKELVLN
SPDMLLKQQQRMAQFTQQKASTLQSEQKLKTGIKFWSL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | H9ZMK2 |
T4CP
| ID | 3523 | GenBank | WP_000053820 |
| Name | t4cp2_HVV65_RS22865_pRHB03-C23_2 |
UniProt ID | _ |
| Length | 611 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 611 a.a. Molecular weight: 69389.23 Da Isoelectric Point: 7.1491
>WP_000053820.1 MULTISPECIES: type IV secretory system conjugative DNA transfer family protein [Enterobacteriaceae]
MSLKLPDKGQWAFIGLVMCLVTYYTGSVAVYFLNGKTPLYIWKNFDSMLLWRIITESNIRTDIRLTAIPS
LLSGMVSSLIVPVFIIWQLNKTDVALYGDAKFASDNDLRKSKLLKWEKENDTDILVGAYKGKYLWYTAPD
FVSLGAGTRAGKGAAIGIPNLLVRKHSLIALDPKQELWKITSKVREILLGNKVYLLDPFNSKTHQFNPLF
YIDLKAESGAKDLLKLIEILFPSYGMTGAEAHFNNLAGQYWTGLAKLLHFFINYEPSWLNEFGLKPVFSI
GSVVDLYSNIDRELILSKREELEGTNGLDENALYHLRDALTKIREYHETEDEQRSSIDGSFRKKMSLFYL
PTVRKCTDGNDFDLRQLRREDITVYVGVNAEDISLAYDFLNLFFNFVVEVTLRENPDFDPTLKHDCLMFL
DEFPSIGYMPIIKKGSGYIAGFKLKLLTIYQNISQLNEIYGIEGAKTLMSAHPCRIIYAVSEEDDAAKIS
EKLGYITTTSKSTSKSRGRSTSQGESESEARRALVLPQELGTLDFKEEFIILKGENPVKAEKALYYLDPY
FMDRLMKVSPKLASLTMELNKTKKIFGVKGLKYPSKEKMLSVGELESEVLL
MSLKLPDKGQWAFIGLVMCLVTYYTGSVAVYFLNGKTPLYIWKNFDSMLLWRIITESNIRTDIRLTAIPS
LLSGMVSSLIVPVFIIWQLNKTDVALYGDAKFASDNDLRKSKLLKWEKENDTDILVGAYKGKYLWYTAPD
FVSLGAGTRAGKGAAIGIPNLLVRKHSLIALDPKQELWKITSKVREILLGNKVYLLDPFNSKTHQFNPLF
YIDLKAESGAKDLLKLIEILFPSYGMTGAEAHFNNLAGQYWTGLAKLLHFFINYEPSWLNEFGLKPVFSI
GSVVDLYSNIDRELILSKREELEGTNGLDENALYHLRDALTKIREYHETEDEQRSSIDGSFRKKMSLFYL
PTVRKCTDGNDFDLRQLRREDITVYVGVNAEDISLAYDFLNLFFNFVVEVTLRENPDFDPTLKHDCLMFL
DEFPSIGYMPIIKKGSGYIAGFKLKLLTIYQNISQLNEIYGIEGAKTLMSAHPCRIIYAVSEEDDAAKIS
EKLGYITTTSKSTSKSRGRSTSQGESESEARRALVLPQELGTLDFKEEFIILKGENPVKAEKALYYLDPY
FMDRLMKVSPKLASLTMELNKTKKIFGVKGLKYPSKEKMLSVGELESEVLL
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Host bacterium
| ID | 5572 | GenBank | NZ_CP055807 |
| Plasmid name | pRHB03-C23_2 | Incompatibility group | IncX1 |
| Plasmid size | 52614 bp | Coordinate of oriT [Strand] | 41032..41380 [-] |
| Host baterium | Escherichia fergusonii strain RHB03-C23 |
Cargo genes
| Drug resistance gene | ant(3'')-Ia, aph(6)-Id, dfrA14, sul2 |
| Virulence gene | - |
| Metal resistance gene | arsH |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |