Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 103544 |
| Name | oriT_112|P2 |
| Organism | Citrobacter freundii isolate 112 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_OW848790 (13737..13874 [-], 138 nt) |
| oriT length | 138 nt |
| IRs (inverted repeats) | _ |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 138 nt
>oriT_112|P2
CGAATGGGCAGGCGTCCGTTGGAACCACGTGGCGAAGCCATGTGTGTTACAACGGTCATCCTGTATTGCTCATCCGCTCTACTATCGTATCAGCAGTTTAGGTGCACGGGACGCACTTTGCAACAAGCACAAATGGCG
CGAATGGGCAGGCGTCCGTTGGAACCACGTGGCGAAGCCATGTGTGTTACAACGGTCATCCTGTATTGCTCATCCGCTCTACTATCGTATCAGCAGTTTAGGTGCACGGGACGCACTTTGCAACAAGCACAAATGGCG
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file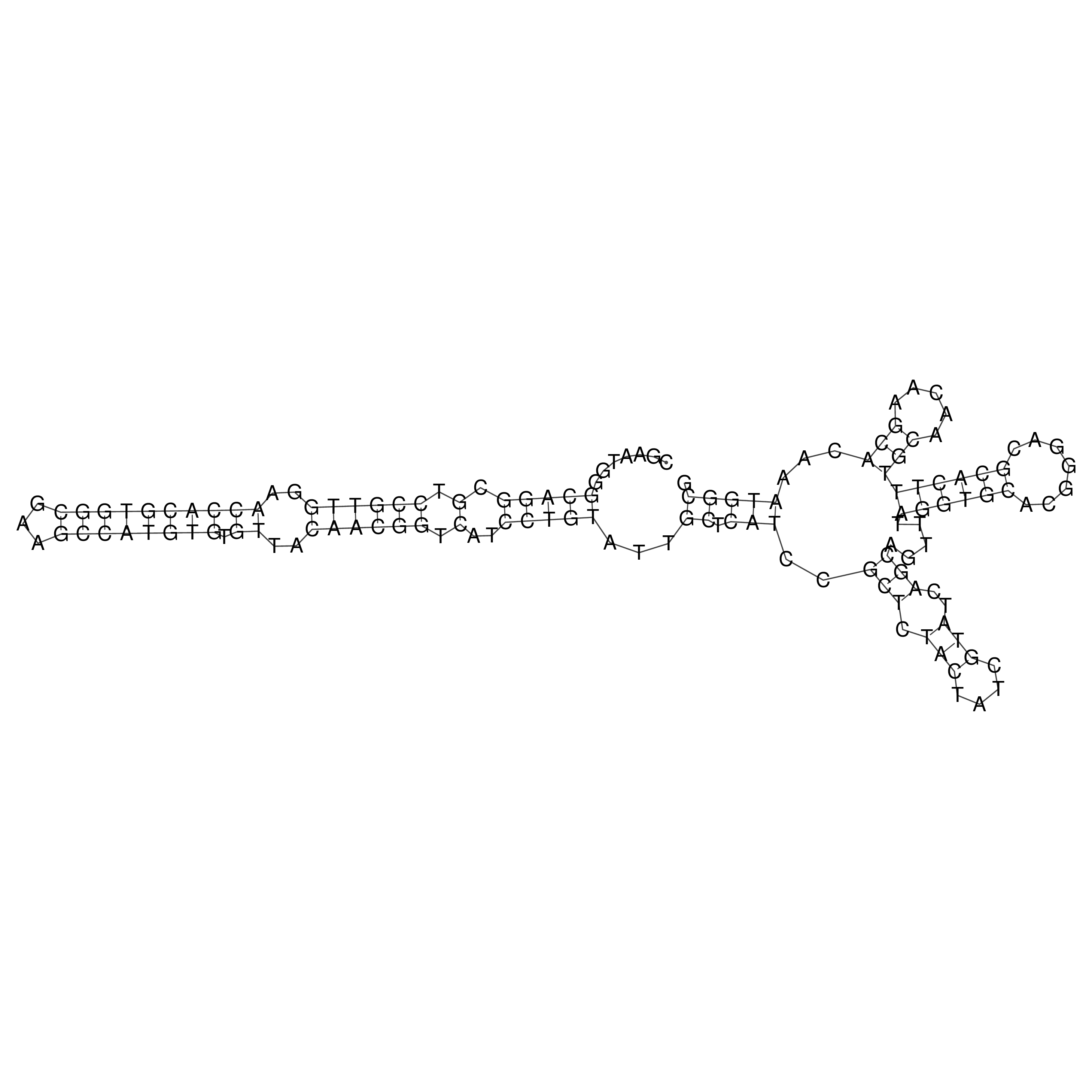
Relaxase
| ID | 2652 | GenBank | WP_033996285 |
| Name | Relaxase_LQ087_RS25230_112|P2 |
UniProt ID | A0A0U2DHM7 |
| Length | 890 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 890 a.a. Molecular weight: 101454.99 Da Isoelectric Point: 9.8485
>WP_033996285.1 MULTISPECIES: TraI/MobA(P) family conjugative relaxase [Pseudomonadota]
MIVKKVKNPQKAASKAVRVSRLTDYIREPERENSQEKCIHAGARGFITDEPQSQTAEMIALSQEAVRSKD
TINHYVLSWREGEQPSPEQVEEAVSIFMDELGVKDHQAIYGLHADTDNLHLHLAINRVHPETLKVVKINN
GFDIEAAHKAIARIENAQGWQREQNGRYQVLENGELGREHIDKDKPRQPAQPKRDMENRTGEKSAERIAI
EDGAPIIKKAQTWEQLHRELAAKGMRYEKTGSGATLFVGDVGVKASSADRDASLSKLQKRLGAYQPPPQR
QQVAQREPEPIKPDVPGWKDYITGRKAHYAEKNAAKLALDKRQEQERKQIAEQQKARRDELMRGNWKGKG
EVLNAMRSVIAAEQAAEKAALKEKHQKQREQHRQQFRPYPDLEQWQRMQKSPELAEQWRHRASEPQRIEG
ASGEPPTPRDIRAYQPEIVGQQVHYSRKEEAGAGGGVSFVDKGKSIDIHDWRNRDSTLAALQLSAQKWGS
FTVTGNDEYKAMCAKLAAEHGFKITNPELQERIQQERQRIQQERAQAMKSEQLKQFELYAEAVGAERYRV
TSIKMQADGRKQTFILDKKDGITRGFTPQEIEQRTPEMLRLQRRGENLYYTPLSDKKHHILIDDMNREKL
ERLIRDGYRPAVVLESSPGNYQAIITVPKLGTAHDKDVGNRLSDALNREYGDPKLSGAIHPHRAPGYENR
KPKHQREDGSYPEVRLLKAERRECVKALALSSQIDAEYQRQAALKAQQPERSKAKPALELAAASGSAIDA
YQRHYRDVIKRQRGGEVDLSRVDSMIAVRMRVTGHDQAAIEGAIRQCAPATRQKDEGRDWNDYAQRTARY
AYSAAGDRQAAELGKYRQQWEKLEGREPVRQQEQAKAQKIERDNSPGMSR
MIVKKVKNPQKAASKAVRVSRLTDYIREPERENSQEKCIHAGARGFITDEPQSQTAEMIALSQEAVRSKD
TINHYVLSWREGEQPSPEQVEEAVSIFMDELGVKDHQAIYGLHADTDNLHLHLAINRVHPETLKVVKINN
GFDIEAAHKAIARIENAQGWQREQNGRYQVLENGELGREHIDKDKPRQPAQPKRDMENRTGEKSAERIAI
EDGAPIIKKAQTWEQLHRELAAKGMRYEKTGSGATLFVGDVGVKASSADRDASLSKLQKRLGAYQPPPQR
QQVAQREPEPIKPDVPGWKDYITGRKAHYAEKNAAKLALDKRQEQERKQIAEQQKARRDELMRGNWKGKG
EVLNAMRSVIAAEQAAEKAALKEKHQKQREQHRQQFRPYPDLEQWQRMQKSPELAEQWRHRASEPQRIEG
ASGEPPTPRDIRAYQPEIVGQQVHYSRKEEAGAGGGVSFVDKGKSIDIHDWRNRDSTLAALQLSAQKWGS
FTVTGNDEYKAMCAKLAAEHGFKITNPELQERIQQERQRIQQERAQAMKSEQLKQFELYAEAVGAERYRV
TSIKMQADGRKQTFILDKKDGITRGFTPQEIEQRTPEMLRLQRRGENLYYTPLSDKKHHILIDDMNREKL
ERLIRDGYRPAVVLESSPGNYQAIITVPKLGTAHDKDVGNRLSDALNREYGDPKLSGAIHPHRAPGYENR
KPKHQREDGSYPEVRLLKAERRECVKALALSSQIDAEYQRQAALKAQQPERSKAKPALELAAASGSAIDA
YQRHYRDVIKRQRGGEVDLSRVDSMIAVRMRVTGHDQAAIEGAIRQCAPATRQKDEGRDWNDYAQRTARY
AYSAAGDRQAAELGKYRQQWEKLEGREPVRQQEQAKAQKIERDNSPGMSR
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A0U2DHM7 |
Host bacterium
| ID | 3987 | GenBank | NZ_OW848790 |
| Plasmid name | 112|P2 | Incompatibility group | IncP6 |
| Plasmid size | 38483 bp | Coordinate of oriT [Strand] | 13737..13874 [-] |
| Host baterium | Citrobacter freundii isolate 112 |
Cargo genes
| Drug resistance gene | blaKPC-2 |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |