Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 103485 |
| Name | oriT_AGR_220|unnamed1 |
| Organism | Serratia entomophila strain AGR_220 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_MT039147 (83415..83497 [+], 83 nt) |
| oriT length | 83 nt |
| IRs (inverted repeats) | _ |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 83 nt
>oriT_AGR_220|unnamed1
ACGGGACCAGATGTGTTTTGTAGCACCGCCTGCACGCAGTCGGTTCAACAAAACGCTGGGTCAGGGCAGAGCCCCGACACCCC
ACGGGACCAGATGTGTTTTGTAGCACCGCCTGCACGCAGTCGGTTCAACAAAACGCTGGGTCAGGGCAGAGCCCCGACACCCC
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file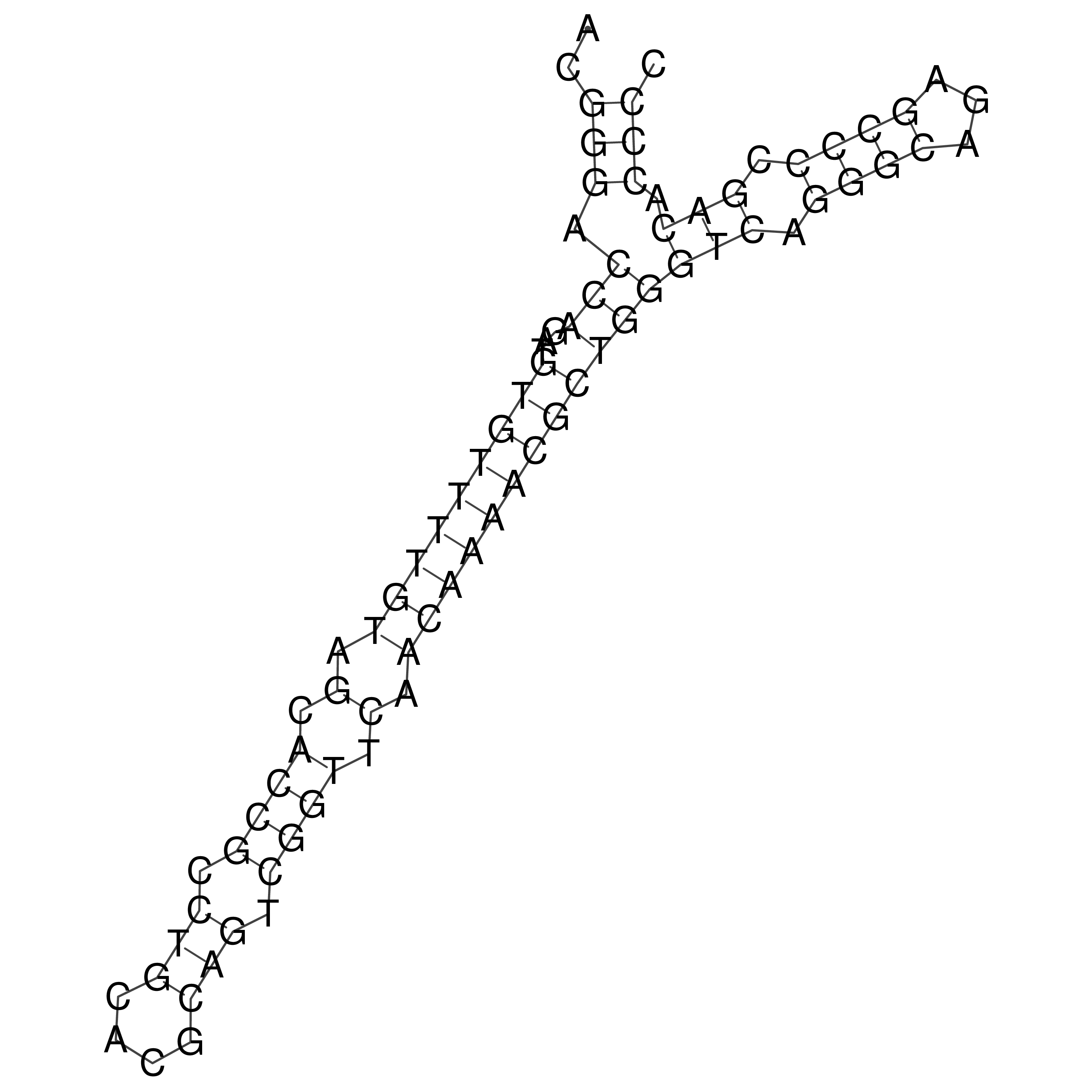
Relaxase
| ID | 2607 | GenBank | WP_241392172 |
| Name | Relaxase_MOO63_RS00450_AGR_220|unnamed1 |
UniProt ID | _ |
| Length | 651 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 651 a.a. Molecular weight: 73366.68 Da Isoelectric Point: 7.6690
>WP_241392172.1 MULTISPECIES: TraI/MobA(P) family conjugative relaxase [Serratia]
MIIRIPEKRRDGKSSFLQLVAYTVVRDDDKPDTPLEPEHPGWRRPRSKDAIFNHLVDYISRNGDAGLQQI
MHADPHGHTQVMFDGVMCETNCFNITTASVEMNAVALQNTRCKDPVLHYFLSWPETDNPSTEHIFDSVRH
SLSALGMSEHQYVAAIHTDTNNIHCHVAANRIHPETYKAADDSFTRIRLQQAARELELKYNWTPTNGFFA
VNDQGEIVRSKREKSLTPSGAKALEYYADVESLHTYAVTECGFKIDEAMADPNLTWKDVHRIMVNAGLAL
KPKGKGLAIYSIDNPELPPVKASSVHPDLTLGCLEKDLGLFQPLENVGTYTKDHTTTATDAIVHEFRYEP
TLHARDLTARLERRLARADARADLKARYQAYKHTWKRPRLDASVVKRRYQNESKRFAWQKARARVAVGDP
LLRKLTYHIIEVERMKAMAALRLAVKEERAAFKADPANRRLSYREWVEQQALSHDQAAIAQLRAFAYRMK
RTQRTASISTNSIVCAVADDTPAFALEGYGTLVTRDGTVQYVRDGRIELQDKGERIEVGDHRVNEGEHIA
GGMALAENKSGEHLKFEGEPVFVQQACSMVRWFNAGGEAPLPLSDPQQRIMAGYDASSQAISTGEGQHKP
SADEHPDVGQNKPRSGYRPQQ
MIIRIPEKRRDGKSSFLQLVAYTVVRDDDKPDTPLEPEHPGWRRPRSKDAIFNHLVDYISRNGDAGLQQI
MHADPHGHTQVMFDGVMCETNCFNITTASVEMNAVALQNTRCKDPVLHYFLSWPETDNPSTEHIFDSVRH
SLSALGMSEHQYVAAIHTDTNNIHCHVAANRIHPETYKAADDSFTRIRLQQAARELELKYNWTPTNGFFA
VNDQGEIVRSKREKSLTPSGAKALEYYADVESLHTYAVTECGFKIDEAMADPNLTWKDVHRIMVNAGLAL
KPKGKGLAIYSIDNPELPPVKASSVHPDLTLGCLEKDLGLFQPLENVGTYTKDHTTTATDAIVHEFRYEP
TLHARDLTARLERRLARADARADLKARYQAYKHTWKRPRLDASVVKRRYQNESKRFAWQKARARVAVGDP
LLRKLTYHIIEVERMKAMAALRLAVKEERAAFKADPANRRLSYREWVEQQALSHDQAAIAQLRAFAYRMK
RTQRTASISTNSIVCAVADDTPAFALEGYGTLVTRDGTVQYVRDGRIELQDKGERIEVGDHRVNEGEHIA
GGMALAENKSGEHLKFEGEPVFVQQACSMVRWFNAGGEAPLPLSDPQQRIMAGYDASSQAISTGEGQHKP
SADEHPDVGQNKPRSGYRPQQ
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4CP
| ID | 2407 | GenBank | WP_241392175 |
| Name | trbC_MOO63_RS00465_AGR_220|unnamed1 |
UniProt ID | _ |
| Length | 725 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 725 a.a. Molecular weight: 81653.39 Da Isoelectric Point: 8.3675
>WP_241392175.1 MULTISPECIES: F-type conjugative transfer protein TrbC [Serratia]
MSRPVEVDPRRVRRPLGYTFINDALLSPIGVQLSLLAGLIVGFVMPATLLLTVPGLLLLVMLFADRPFRM
PLRMPTDIGGMDLTTEREMPKYRKGLGGFFRYVVRTRKYMPAAGVMCLGYARGKGLARELWLTLDDALRH
MLLLATTGSGKTEALLSVFLNSICWGRGICYSDGKGQNTLAFAMWSLARRFGREDDFYVLNFMTGGTDKL
LQLLLNDKKRQPSNTINLFGTANTTFIIQLMESMLPPAGSGDQGWQDKAKSMLSALVYAIYYKCKREKRR
ISQKVIQEYLPLRKLADLYQEAKRDGWHREAYNPLENYFNTLAGFRIELISKPSEWEQGVYDQHGYLIQQ
FNRMLTMFNDLYGHIFSTDAGDIDIEDVLHNDRILCTTIPALELSKGEASNIGKLYISAIRMTMARDLGC
ELEGMMNDVLIVKKYSGKFPYPIAMDELGAYFGPGMDNLASQMRSLGYMLIVSAQDIQRFIAEHKGEYMT
VNANLLTKWFMTLQDEKDTFELARITGGKGYYAELGSVEQAGGFITPNYEDAANNYIREKDRLDLGDLKD
MSPGEGMISFKSALVPSNAIYIKDDDKMTSSLPMRINRFIDVETPTEAELFTLNPSLKRKLPPTAQEIDG
ILQRLDQAMNTTTSAGMMDPVLKRVAAVALDLDNRADVSYSPTQRGVLLFEAAREALHQNKRNWRQLPSP
PKPIRVSKEVAQSLANAGTNEITFR
MSRPVEVDPRRVRRPLGYTFINDALLSPIGVQLSLLAGLIVGFVMPATLLLTVPGLLLLVMLFADRPFRM
PLRMPTDIGGMDLTTEREMPKYRKGLGGFFRYVVRTRKYMPAAGVMCLGYARGKGLARELWLTLDDALRH
MLLLATTGSGKTEALLSVFLNSICWGRGICYSDGKGQNTLAFAMWSLARRFGREDDFYVLNFMTGGTDKL
LQLLLNDKKRQPSNTINLFGTANTTFIIQLMESMLPPAGSGDQGWQDKAKSMLSALVYAIYYKCKREKRR
ISQKVIQEYLPLRKLADLYQEAKRDGWHREAYNPLENYFNTLAGFRIELISKPSEWEQGVYDQHGYLIQQ
FNRMLTMFNDLYGHIFSTDAGDIDIEDVLHNDRILCTTIPALELSKGEASNIGKLYISAIRMTMARDLGC
ELEGMMNDVLIVKKYSGKFPYPIAMDELGAYFGPGMDNLASQMRSLGYMLIVSAQDIQRFIAEHKGEYMT
VNANLLTKWFMTLQDEKDTFELARITGGKGYYAELGSVEQAGGFITPNYEDAANNYIREKDRLDLGDLKD
MSPGEGMISFKSALVPSNAIYIKDDDKMTSSLPMRINRFIDVETPTEAELFTLNPSLKRKLPPTAQEIDG
ILQRLDQAMNTTTSAGMMDPVLKRVAAVALDLDNRADVSYSPTQRGVLLFEAAREALHQNKRNWRQLPSP
PKPIRVSKEVAQSLANAGTNEITFR
Protein domains
No domain identified.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 88965..118242
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| MOO63_RS00455 (220p1_00097) | 85870..86193 | - | 324 | WP_241392173 | hypothetical protein | - |
| MOO63_RS00460 (220p1_00098) | 86203..86739 | - | 537 | WP_241392174 | phospholipase D family protein | - |
| MOO63_RS00465 (220p1_00099) | 86750..88927 | - | 2178 | WP_241392175 | F-type conjugative transfer protein TrbC | - |
| MOO63_RS00470 (220p1_00100) | 88965..89978 | - | 1014 | WP_241392176 | hypothetical protein | trbB |
| MOO63_RS00475 (220p1_00101) | 89992..91251 | - | 1260 | WP_241392177 | conjugal transfer protein TrbA | trbA |
| MOO63_RS00480 (220p1_00102) | 91444..92022 | - | 579 | WP_241392178 | hypothetical protein | - |
| MOO63_RS00485 (220p1_00103) | 92019..94163 | - | 2145 | WP_241392179 | DotA/TraY family protein | traY |
| MOO63_RS00490 (220p1_00104) | 94227..94790 | - | 564 | WP_241392180 | conjugal transfer protein TraX | - |
| MOO63_RS00495 (220p1_00105) | 94783..96009 | - | 1227 | WP_241392181 | conjugal transfer protein TraW | traW |
| MOO63_RS00500 (220p1_00107) | 96150..99224 | - | 3075 | WP_241392182 | type IV secretory system conjugative DNA transfer family protein | traU |
| MOO63_RS00505 (220p1_00108) | 99227..99832 | - | 606 | WP_241392183 | hypothetical protein | traT |
| MOO63_RS00510 (220p1_00109) | 99915..100316 | - | 402 | WP_241392184 | DUF6750 family protein | traR |
| MOO63_RS00515 (220p1_00110) | 100332..100862 | - | 531 | WP_241392185 | conjugal transfer protein TraQ | traQ |
| MOO63_RS00520 (220p1_00111) | 100871..101614 | - | 744 | WP_241392186 | hypothetical protein | traP |
| MOO63_RS00525 (220p1_00112) | 101614..102846 | - | 1233 | WP_241392187 | conjugal transfer protein TraO | traO |
| MOO63_RS00530 (220p1_00113) | 102851..103858 | - | 1008 | WP_241392188 | DotH/IcmK family type IV secretion protein | traN |
| MOO63_RS00535 (220p1_00114) | 103861..104535 | - | 675 | WP_241392189 | DotI/IcmL/TraM family protein | traM |
| MOO63_RS00540 (220p1_00115) | 104628..105119 | - | 492 | WP_241392190 | hypothetical protein | traL |
| MOO63_RS00545 (220p1_00116) | 105129..108362 | - | 3234 | WP_241392191 | LPD7 domain-containing protein | - |
| MOO63_RS00550 (220p1_00118) | 108505..108771 | - | 267 | WP_241392192 | IcmT/TraK family protein | traK |
| MOO63_RS00555 (220p1_00119) | 108771..109928 | - | 1158 | WP_241392193 | plasmid transfer ATPase TraJ | virB11 |
| MOO63_RS00560 (220p1_00120) | 109959..110747 | - | 789 | WP_241392194 | type IV secretory system conjugative DNA transfer family protein | traI |
| MOO63_RS00565 (220p1_00121) | 110744..111208 | - | 465 | WP_241392195 | DotD/TraH family lipoprotein | - |
| MOO63_RS00570 (220p1_00122) | 111302..112378 | - | 1077 | WP_241392196 | acyltransferase | - |
| MOO63_RS00575 (220p1_00123) | 112458..113810 | - | 1353 | WP_241392197 | shufflon system plasmid conjugative transfer pilus tip adhesin PilV | - |
| MOO63_RS00580 (220p1_00124) | 113807..114475 | - | 669 | WP_241392198 | prepilin peptidase | - |
| MOO63_RS00585 (220p1_00125) | 114475..114960 | - | 486 | WP_241392199 | lytic transglycosylase domain-containing protein | virB1 |
| MOO63_RS00590 (220p1_00126) | 115015..115605 | - | 591 | WP_241392200 | type 4 pilus major pilin | - |
| MOO63_RS00595 (220p1_00127) | 115640..116755 | - | 1116 | WP_241392201 | type II secretion system F family protein | - |
| MOO63_RS00600 (220p1_00128) | 116752..118242 | - | 1491 | WP_241392202 | ATPase, T2SS/T4P/T4SS family | virB11 |
| MOO63_RS00605 (220p1_00129) | 118248..118832 | - | 585 | WP_241392203 | type IV pilus biogenesis protein PilP | - |
| MOO63_RS00610 (220p1_00130) | 118819..120123 | - | 1305 | WP_241392204 | type 4b pilus protein PilO2 | - |
| MOO63_RS00615 (220p1_00131) | 120127..121761 | - | 1635 | WP_241392205 | PilN family type IVB pilus formation outer membrane protein | - |
| MOO63_RS00620 (220p1_00132) | 121784..122209 | - | 426 | WP_241392206 | type IV pilus biogenesis protein PilM | - |
| MOO63_RS00625 (220p1_00133) | 122209..123147 | - | 939 | WP_241392207 | toxin co-regulated pilus biosynthesis Q family protein | - |
Host bacterium
| ID | 3928 | GenBank | NZ_MT039147 |
| Plasmid name | AGR_220|unnamed1 | Incompatibility group | - |
| Plasmid size | 145178 bp | Coordinate of oriT [Strand] | 83415..83497 [+] |
| Host baterium | Serratia entomophila strain AGR_220 |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |