Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 103278 |
| Name | oriT_pCERC6 |
| Organism | Escherichia coli strain 5.1-R1 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_MH287044 (34085..34157 [-], 73 nt) |
| oriT length | 73 nt |
| IRs (inverted repeats) | _ |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 73 nt
>oriT_pCERC6
GTCGGGGCAAAGCCCTGACCAGGTGGGGAATGTCTGAGTGCGCGTGCGCGGTCCGACATTCCCACATCCTGTC
GTCGGGGCAAAGCCCTGACCAGGTGGGGAATGTCTGAGTGCGCGTGCGCGGTCCGACATTCCCACATCCTGTC
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file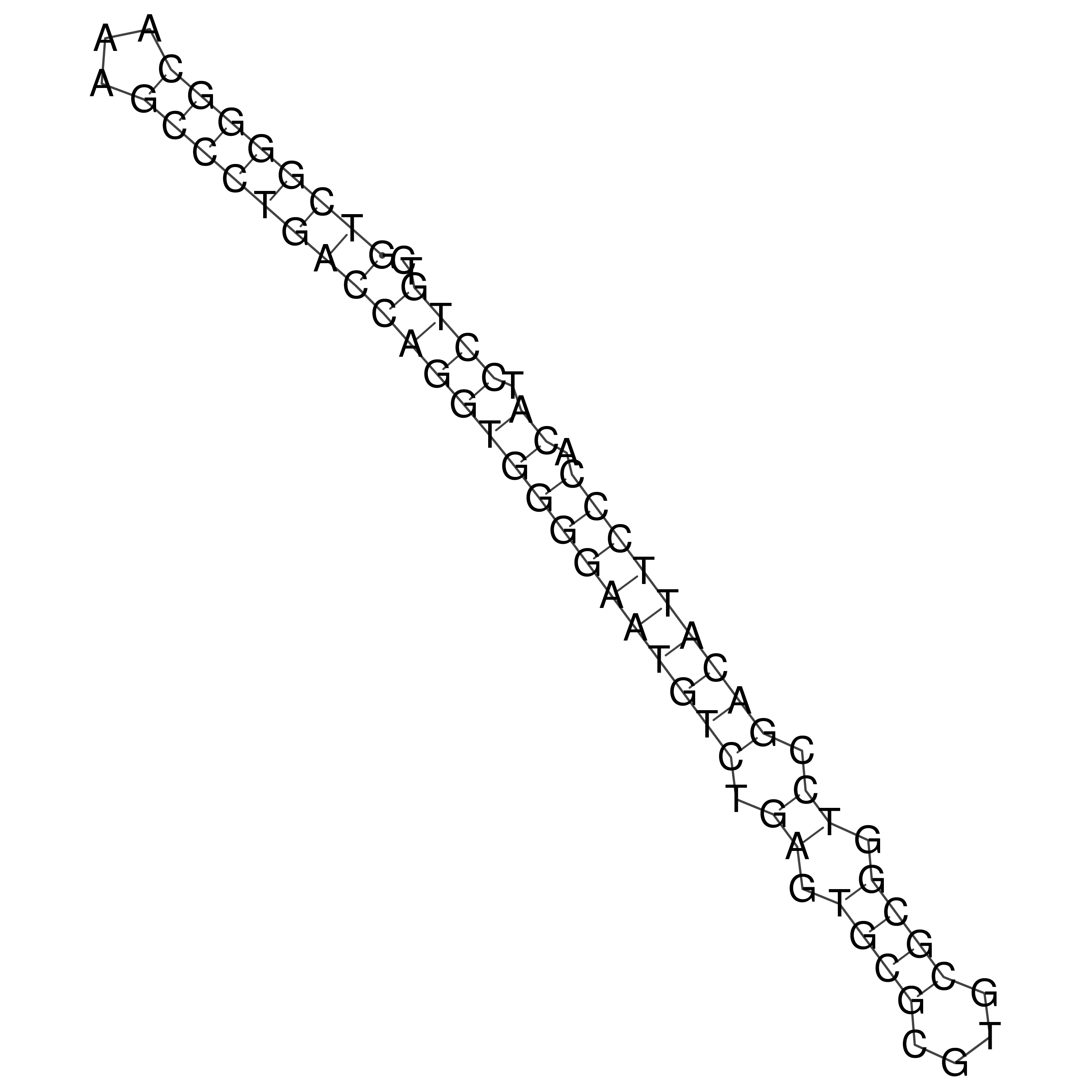
Relaxase
| ID | 2447 | GenBank | WP_000991387 |
| Name | Relaxase_HTT41_RS00250_pCERC6 |
UniProt ID | A0A7I0L2J7 |
| Length | 903 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 903 a.a. Molecular weight: 104567.95 Da Isoelectric Point: 7.5259
>WP_000991387.1 MULTISPECIES: relaxase/mobilization nuclease domain-containing protein [Enterobacteriaceae]
MNAIIPHKRRDGRSSFEDLIAYTSVRDDVQENELTEDSEIKSDVPHRNRFRRLVDYATRLRNEKFVSLID
VMKDGSQWVNFYGVTCFHNCNSLETAAEEMQYKADKAVFSRNDTDPVFHYILSWPAHESPRPEQLFDCVR
HTLKSLELSKHQYVAAVHTDTDNLHVHVAVNRVHPETGYINWLSWSQEKLSRACRELELKHGFAPDNGCF
VHAPGNRIVRKTALVRERRNAWRRGKKQTFREYIAQMSIAGLREEPAQDWLSLHKRLASDGLYITMQEGE
LVVKDGWDRAREGVALSSFGPSWTAEKLGRKLGEYQPVPTDIFSQVGTPGRYDPEAINVDIRPEKVAETE
SLKQYACRYFAERLPEMARNGELESCLDVHRTLAKAGLWMGIQHGHLVLHDGFDKQQTPVRADSVWPLMT
LDYMQDLDGGWQPVPKDIFTQVIPGERFRGRNLGTQAVSDYEWYRMRMGTGPQGAIKRELFSDKESLWGY
TTVQCESLIEDMIAGGNFSWQACHEMFARKGLMLQKQHHGLVIVDAFNHELTPVKASSIHPDLTLSRAEL
QAGPFEIAGADIFERVKPECRYNPELAASDEVEPGFRRDPVLRRERREARAAAREDLRARYLAWKEHWRK
PDLRYGERLREIHAACRRRKAYIRVQFRDPQLRKLHYHIAEVQRMQALIRLKESVKEERLSLIAEGKWYP
LSYRQWVEQQAVQGDRAALSQLRGWDYRDRRKDKRRTTNADRCVILCEPGGTPLYEDTGVLEARLQKDGS
VRFRDRRNGELVCVDYGDRVVFYHHQDRNELVDKLNLIAPVLFDREPGMGFEPEGSYQQFNDVFAEMVAW
HNAAGITGNGHFVISRPDVDLHRQRSEQYYHEYIRQQKSISGGHGASYAPVQDNEWTPPSPGM
MNAIIPHKRRDGRSSFEDLIAYTSVRDDVQENELTEDSEIKSDVPHRNRFRRLVDYATRLRNEKFVSLID
VMKDGSQWVNFYGVTCFHNCNSLETAAEEMQYKADKAVFSRNDTDPVFHYILSWPAHESPRPEQLFDCVR
HTLKSLELSKHQYVAAVHTDTDNLHVHVAVNRVHPETGYINWLSWSQEKLSRACRELELKHGFAPDNGCF
VHAPGNRIVRKTALVRERRNAWRRGKKQTFREYIAQMSIAGLREEPAQDWLSLHKRLASDGLYITMQEGE
LVVKDGWDRAREGVALSSFGPSWTAEKLGRKLGEYQPVPTDIFSQVGTPGRYDPEAINVDIRPEKVAETE
SLKQYACRYFAERLPEMARNGELESCLDVHRTLAKAGLWMGIQHGHLVLHDGFDKQQTPVRADSVWPLMT
LDYMQDLDGGWQPVPKDIFTQVIPGERFRGRNLGTQAVSDYEWYRMRMGTGPQGAIKRELFSDKESLWGY
TTVQCESLIEDMIAGGNFSWQACHEMFARKGLMLQKQHHGLVIVDAFNHELTPVKASSIHPDLTLSRAEL
QAGPFEIAGADIFERVKPECRYNPELAASDEVEPGFRRDPVLRRERREARAAAREDLRARYLAWKEHWRK
PDLRYGERLREIHAACRRRKAYIRVQFRDPQLRKLHYHIAEVQRMQALIRLKESVKEERLSLIAEGKWYP
LSYRQWVEQQAVQGDRAALSQLRGWDYRDRRKDKRRTTNADRCVILCEPGGTPLYEDTGVLEARLQKDGS
VRFRDRRNGELVCVDYGDRVVFYHHQDRNELVDKLNLIAPVLFDREPGMGFEPEGSYQQFNDVFAEMVAW
HNAAGITGNGHFVISRPDVDLHRQRSEQYYHEYIRQQKSISGGHGASYAPVQDNEWTPPSPGM
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A7I0L2J7 |
Auxiliary protein
| ID | 1045 | GenBank | WP_001291058 |
| Name | WP_001291058_pCERC6 |
UniProt ID | A0A7I0L241 |
| Length | 110 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
Auxiliary protein sequence
Download Length: 110 a.a. Molecular weight: 12730.52 Da Isoelectric Point: 9.9656
>WP_001291058.1 MULTISPECIES: plasmid mobilization protein MobA [Enterobacteriaceae]
MSEKKTRTGSENRKRIVKFTARFTEDEAEIVREKAESSGQTVSTFIRSSSLDKVVNCRTDDRMIDEIMRL
GRLQKHLFVEGKRTGDKEYAEVLVAITEYVNALRRDLMGR
MSEKKTRTGSENRKRIVKFTARFTEDEAEIVREKAESSGQTVSTFIRSSSLDKVVNCRTDDRMIDEIMRL
GRLQKHLFVEGKRTGDKEYAEVLVAITEYVNALRRDLMGR
Protein domains
No domain identified.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A7I0L241 |
T4CP
| ID | 2221 | GenBank | WP_000077663 |
| Name | trbC_HTT41_RS00305_pCERC6 |
UniProt ID | _ |
| Length | 767 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 767 a.a. Molecular weight: 87012.13 Da Isoelectric Point: 7.3008
>WP_000077663.1 MULTISPECIES: F-type conjugative transfer protein TrbC [Enterobacteriaceae]
MSQHHVNPDLIHRTAWGNPLWNALHNLNITGLCLAGSIITALIWPLALPVCLLFTLVTSVIFMLQRWRCP
LRMPMTLSLDDPSQDRKVRRSLFSFWPTLFQYEADETFPARGIFYVGYRRINDIGRELWLSMDDLTRHVM
FFATTGGGKTETTFAWLLNPLCWGRGFTFVDGKAQNDTTRTIWYLSRRFGREDDIEVINFMNGGKSRSEI
IQSGEKSRPQSNTWNPFAFSTEAFTAETMQSMLPQNVQGGEWQSRAIAMNKALVFGTKFWCVREGKTMSL
QMLREHMTLEGMAKLYCRGLDDQWPEEAIAPLRNYLQDVPGFDMSLVRTPSAWTEEPRKQHAYLSGQFSE
TFTTFAETFGDVFAADAGDIDIRDSIHSDRILIVMIPALDTSAHTTSALGRMFVTQQSMLLARDLGYRLE
GTDAQTLEVKKYKGSFPYICILDEVGAYYTDRIAVEATQVRSLEFSLIMTAQDQERIEGQTSATNTATLM
QNAGTKFAGRIVSDDKTARTVKNAAGEEARARMGSLQRHDGVMGESWVDGNQITIQMESKIDVQDLIRLN
AGEFFTVFQGDVVPSASVYIPDSEKSCDSDPVVINRYISVEAPRLERLRRLVPRTVQRRLPTPEHVSSII
GVLTAKPSRKRRKNRTEPYRIIDTFQRQLATAQTSLDLLPQYDTDIESRANELWKKAVHTINNTTREERR
VCYITLNRPEEEHSGPEDIPSVKAILNTLLPLEMLLPVPDLTASPPHKKNVAQTPSGGKQESRKKRF
MSQHHVNPDLIHRTAWGNPLWNALHNLNITGLCLAGSIITALIWPLALPVCLLFTLVTSVIFMLQRWRCP
LRMPMTLSLDDPSQDRKVRRSLFSFWPTLFQYEADETFPARGIFYVGYRRINDIGRELWLSMDDLTRHVM
FFATTGGGKTETTFAWLLNPLCWGRGFTFVDGKAQNDTTRTIWYLSRRFGREDDIEVINFMNGGKSRSEI
IQSGEKSRPQSNTWNPFAFSTEAFTAETMQSMLPQNVQGGEWQSRAIAMNKALVFGTKFWCVREGKTMSL
QMLREHMTLEGMAKLYCRGLDDQWPEEAIAPLRNYLQDVPGFDMSLVRTPSAWTEEPRKQHAYLSGQFSE
TFTTFAETFGDVFAADAGDIDIRDSIHSDRILIVMIPALDTSAHTTSALGRMFVTQQSMLLARDLGYRLE
GTDAQTLEVKKYKGSFPYICILDEVGAYYTDRIAVEATQVRSLEFSLIMTAQDQERIEGQTSATNTATLM
QNAGTKFAGRIVSDDKTARTVKNAAGEEARARMGSLQRHDGVMGESWVDGNQITIQMESKIDVQDLIRLN
AGEFFTVFQGDVVPSASVYIPDSEKSCDSDPVVINRYISVEAPRLERLRRLVPRTVQRRLPTPEHVSSII
GVLTAKPSRKRRKNRTEPYRIIDTFQRQLATAQTSLDLLPQYDTDIESRANELWKKAVHTINNTTREERR
VCYITLNRPEEEHSGPEDIPSVKAILNTLLPLEMLLPVPDLTASPPHKKNVAQTPSGGKQESRKKRF
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 49211..88476
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| HTT41_RS00285 | 44277..44384 | - | 108 | Protein_55 | aminoglycoside phosphotransferase family protein | - |
| HTT41_RS00290 | 44384..45187 | - | 804 | WP_001082319 | aminoglycoside O-phosphotransferase APH(3'')-Ib | - |
| HTT41_RS00295 | 45248..46063 | - | 816 | WP_001043260 | sulfonamide-resistant dihydropteroate synthase Sul2 | - |
| HTT41_RS00300 | 46520..46813 | - | 294 | WP_000334996 | hypothetical protein | - |
| HTT41_RS00305 | 46927..49230 | - | 2304 | WP_000077663 | F-type conjugative transfer protein TrbC | - |
| HTT41_RS00310 | 49211..50335 | - | 1125 | WP_000152378 | DsbC family protein | trbB |
| HTT41_RS00315 | 50332..51639 | - | 1308 | WP_000121918 | IncI1-type conjugal transfer protein TrbA | trbA |
| HTT41_RS00320 | 51830..52123 | + | 294 | WP_023636565 | hypothetical protein | - |
| HTT41_RS00600 | 52176..52271 | + | 96 | WP_001388983 | DinQ-like type I toxin DqlB | - |
| HTT41_RS00605 | 52860..52955 | + | 96 | WP_001388983 | DinQ-like type I toxin DqlB | - |
| HTT41_RS00330 | 53018..53194 | - | 177 | WP_001054907 | hypothetical protein | - |
| HTT41_RS00335 | 53344..53811 | - | 468 | WP_000598962 | thermonuclease family protein | - |
| HTT41_RS00340 | 53971..54594 | + | 624 | WP_001155250 | helix-turn-helix domain-containing protein | - |
| HTT41_RS00345 | 54792..55007 | - | 216 | WP_000266543 | hypothetical protein | - |
| HTT41_RS00350 | 55011..55379 | - | 369 | WP_001345852 | hypothetical protein | - |
| HTT41_RS00355 | 55670..55912 | - | 243 | WP_000650923 | hypothetical protein | - |
| HTT41_RS00360 | 56242..56394 | + | 153 | WP_001335079 | Hok/Gef family protein | - |
| HTT41_RS00620 | 56532..58122 | - | 1591 | Protein_72 | TnsA endonuclease N-terminal domain-containing protein | - |
| HTT41_RS00370 | 58396..59046 | - | 651 | WP_001044247 | plasmid IncI1-type surface exclusion protein ExcA | - |
| HTT41_RS00375 | 59135..61300 | - | 2166 | WP_000691810 | DotA/TraY family protein | traY |
| HTT41_RS00380 | 61375..61944 | - | 570 | WP_000650752 | conjugal transfer protein TraX | - |
| HTT41_RS00385 | 61941..63146 | - | 1206 | WP_001189549 | conjugal transfer protein TraW | traW |
| HTT41_RS00390 | 63104..63724 | - | 621 | WP_000286854 | hypothetical protein | traV |
| HTT41_RS00395 | 63724..66768 | - | 3045 | WP_001024766 | conjugal transfer protein | traU |
| HTT41_RS00400 | 66819..67070 | + | 252 | WP_001136184 | hypothetical protein | - |
| HTT41_RS00405 | 67063..67689 | - | 627 | WP_000785708 | hypothetical protein | traT |
| HTT41_RS00595 | 67796..68047 | - | 252 | WP_001277573 | hypothetical protein | - |
| HTT41_RS00415 | 68104..68502 | - | 399 | WP_000088813 | DUF6750 family protein | traR |
| HTT41_RS00420 | 68549..69079 | - | 531 | WP_000987021 | conjugal transfer protein TraQ | traQ |
| HTT41_RS00425 | 69076..69789 | - | 714 | WP_000787002 | conjugal transfer protein TraP | traP |
| HTT41_RS00430 | 69786..71123 | - | 1338 | WP_021516923 | conjugal transfer protein TraO | traO |
| HTT41_RS00435 | 71127..72101 | - | 975 | WP_000547386 | DotH/IcmK family type IV secretion protein | traN |
| HTT41_RS00440 | 72112..72807 | - | 696 | WP_001287545 | DotI/IcmL family type IV secretion protein | traM |
| HTT41_RS00445 | 72819..73169 | - | 351 | WP_001056367 | hypothetical protein | traL |
| HTT41_RS00450 | 73186..77247 | - | 4062 | WP_077561291 | LPD7 domain-containing protein | - |
| HTT41_RS00455 | 77311..77601 | - | 291 | WP_000817801 | IcmT/TraK family protein | traK |
| HTT41_RS00460 | 77598..78746 | - | 1149 | WP_000775210 | plasmid transfer ATPase TraJ | virB11 |
| HTT41_RS00465 | 78730..79566 | - | 837 | WP_000745049 | type IV secretory system conjugative DNA transfer family protein | traI |
| HTT41_RS00470 | 79563..80021 | - | 459 | WP_001170111 | DotD/TraH family lipoprotein | - |
| HTT41_RS00475 | 80125..81327 | - | 1203 | WP_000979993 | conjugal transfer protein TraF | - |
| HTT41_RS00480 | 81429..82250 | - | 822 | WP_000845373 | hypothetical protein | traE |
| HTT41_RS00590 | 82415..82609 | - | 195 | Protein_96 | integrase | - |
| HTT41_RS00485 | 82723..84084 | - | 1362 | WP_001168171 | shufflon system plasmid conjugative transfer pilus tip adhesin PilV | - |
| HTT41_RS00490 | 84102..84728 | - | 627 | WP_000939938 | prepilin peptidase | - |
| HTT41_RS00495 | 84744..85229 | - | 486 | WP_001080767 | lytic transglycosylase domain-containing protein | virB1 |
| HTT41_RS00500 | 85274..85810 | - | 537 | WP_000111521 | type 4 pilus major pilin | - |
| HTT41_RS00505 | 85872..86966 | - | 1095 | WP_000697154 | type II secretion system F family protein | - |
| HTT41_RS00510 | 86968..88476 | - | 1509 | WP_000880806 | ATPase, T2SS/T4P/T4SS family | virB11 |
| HTT41_RS00515 | 88579..89037 | - | 459 | WP_000095884 | type IV pilus biogenesis protein PilP | - |
| HTT41_RS00520 | 89027..90322 | - | 1296 | WP_000129885 | type 4b pilus protein PilO2 | - |
| HTT41_RS00525 | 90343..91962 | - | 1620 | WP_001035559 | PilN family type IVB pilus formation outer membrane protein | - |
| HTT41_RS00530 | 91994..92407 | - | 414 | WP_001604201 | type IV pilus biogenesis protein PilM | - |
Host bacterium
| ID | 3721 | GenBank | NZ_MH287044 |
| Plasmid name | pCERC6 | Incompatibility group | IncB/O/K/Z |
| Plasmid size | 98766 bp | Coordinate of oriT [Strand] | 34085..34157 [-] |
| Host baterium | Escherichia coli strain 5.1-R1 |
Cargo genes
| Drug resistance gene | aph(6)-Id, blaTEM-1C, aph(3'')-Ib, sul2 |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |