Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 103174 |
| Name | oriT_p1220-CTXM |
| Organism | Klebsiella pneumoniae strain 1220 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_KY174332 (78298..78347 [-], 50 nt) |
| oriT length | 50 nt |
| IRs (inverted repeats) | 7..14, 17..24 (GCAAAATT..AATTTTGC) |
| Location of nic site | 33..34 |
| Conserved sequence flanking the nic site |
TGTGTGGTGA |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 50 nt
>oriT_p1220-CTXM
AAATCTGCAAAATTTTAATTTTGCGTAGTGTGTGGTGATTTTGTGGTGAG
AAATCTGCAAAATTTTAATTTTGCGTAGTGTGTGGTGATTTTGTGGTGAG
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file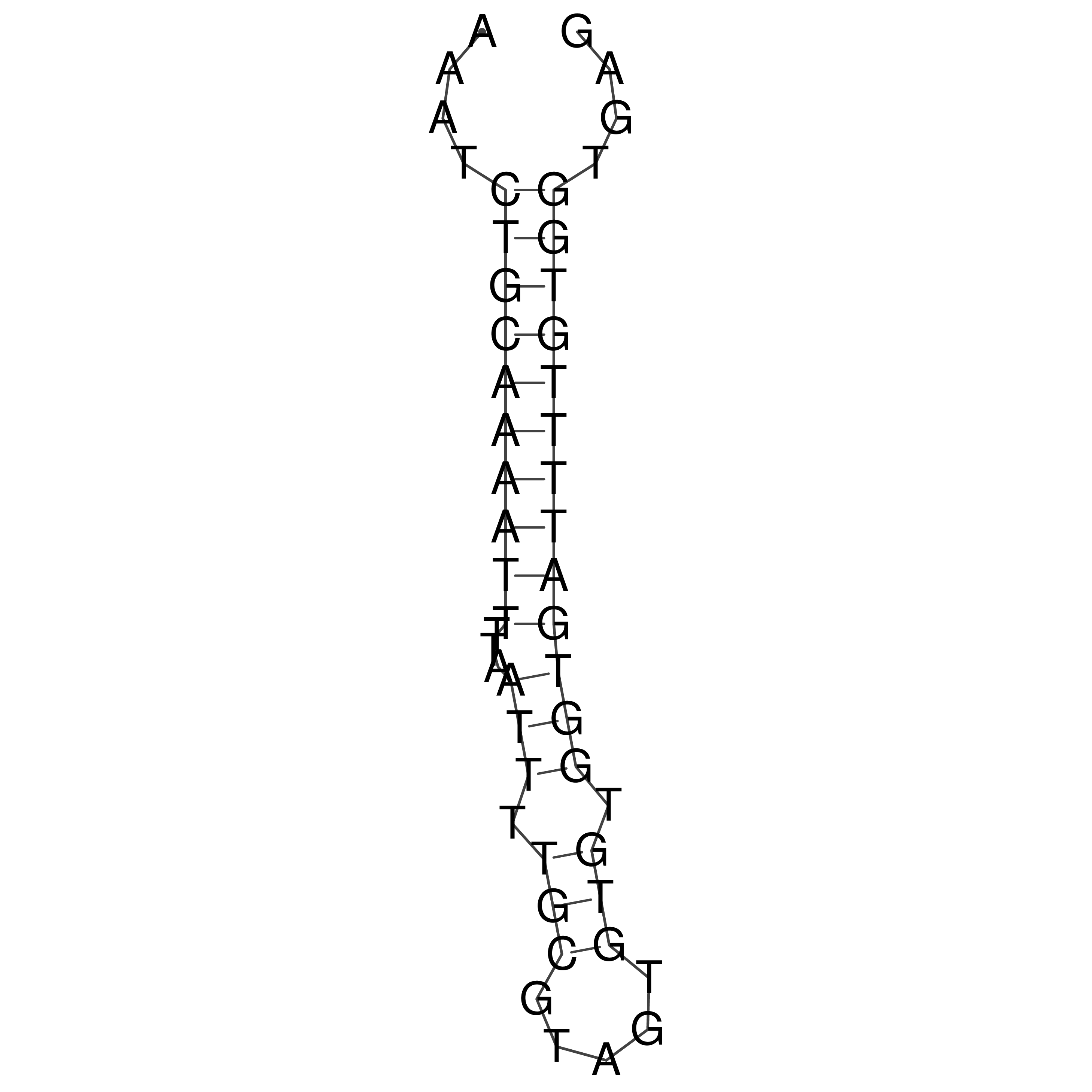
Relaxase
| ID | 2375 | GenBank | WP_043907038 |
| Name | traI_HTL29_RS00665_p1220-CTXM |
UniProt ID | A0A0S1TKY9 |
| Length | 1752 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1752 a.a. Molecular weight: 190458.19 Da Isoelectric Point: 6.0263
>WP_043907038.1 MULTISPECIES: conjugative transfer relaxase/helicase TraI [Enterobacteriaceae]
MLSLSQVKSAGSAGNYYTEKDNYYVIGSMEERWQGKGAELLGLEGKVDKQVFTELLQGKLPDGSDLTRIQ
DGVNKHRPGYDLTFSAPKSVSMLAMLGGDKRLIDAHNRAVTVALNQVESLASTRVKKDGVSETVLTGNLI
IARFNHDTSRAQDPQIHTHSVVINATQNGDKWLTLASDTVGKTGFSETILANRIAFGKIYQNSLRADVES
MGYKTVDAGRNGMWEMEGVPVESFSTRSQELREAAGPDASLKSRDVAALDTRKSKEAIDPAEKMVEWMNT
LKETGFDIRGYREAADARAAELARAPAAPVNTDGPDITDVVTKAIAGLSDRKVQFTYADLLARTVGQLEA
KDGVFELARKGIDAAIEREQLIPLDREKGLFTSNIHVLDELAVKALSQEVQRQNHVSVTPDASVVRQVPF
SDAVSVLAQDRPVMGIVSGQGGATGQRERVAELTLMAREQGRDVHILAADNRSRDFLAGDVRLAGETVTG
KSALQDGTAFIPGGTLIVDQAEKLSLKETISLLDGAMRHNVQVLLSDSGKRSGTGSALTVLKDSGVNTYR
WQGGHQTTADIISEPDKGARYSRLAQEFAVSVREGQESVAQISGTREQSVLNGLIRDSLRQEGVLGEKDT
TITALTPVWLDSKSRGVRDYYREGMVMERWDPETRTHDRFVIDRVTASSNMLTLKDREGDRLDLKVSAVD
SQWTLFRADTLPVAEGERLAVLGKIPDTRLKGGESVTVMKVEDGQLTVQRPGQKTTQTLAVGAGVFDGIK
VGHGWVESPGRSVSETATVFASVTQRELDNATLNQLAQSGSHLRLYSAQDAARTTEKLSRHTAFSVVSEQ
LKSRSGETDLDAAIAQQKAGLHTPAEQAIHLAIPLLESQDLTFSRPQLLATAMETGGGKVSMADIDTTIQ
AQIRSGQLLNVPVAPGRGNDLLISRQAWDAEKSILTRVLEGKDAVAPLMDRVPDSLMTELTAGQRAATRM
ILESTDRFTVVQGYAGVGKTTQFRAVMSAISLLPEETRPRVIGLAPTHRAVGEMQSAGVDARTTASFLHD
TQLLQRNGQTPDFSNTLFLLDESSMVGLADMAKAHSLIAAGGGRAVPSGDTDQLPPIASGQPFRLTQQRS
AADVAIMKEIVRQVPELRPAVYSLIERDVHRALTTIEQVTPEQVPRKEGAWAPGSSVVEFTPKQEKAIEK
ALSEGKTLPEGQPATLYEALVKDYTGRTPEAQSQTLVITHLNKDRRALNSLIHDARRENGETGKEEITLP
VLVTSNIRDGELRKLSTWTAHKEAVALVDNVYHRISKVDKDNQLITLTDSEGKERFISPREASAEGVTLY
RQEKITVSQGDRMRFSKSDPERGYVANSIWEVQSVSGDSVTLSDGKITRTLTPKAEQAQQHIDLAYAITA
HGAQGASEPYAIALEGVAGGREQMASFESAYVALSRMKQHVQVYTDNREGWIKAIKNSPEKATAHDILEP
RNDRAVKTADLLFGRARPLDETAAGRAALQQSGLAQGSSPGKFISPGKKYPQPHVALPAFDKNGKAAGIW
LSPLTDRDGRLEAIGGEGRIMGNEDARFVALQNSRNGESLLAGNMGEGVRMARDNPDTGVVVRLAGDDRP
WNPGAMTGGRVWAEPAPVAPVPQAGADIILPPEVLAQRAAEEQQRREMEKQAEQTAREVAGEARKAGEPA
DRVKEVIGDVIRGLERDRPGTEKTTLPDDPQFRRQEEAIQQVASERLQRERLQAVERDMVRDLNREKTLG
GD
MLSLSQVKSAGSAGNYYTEKDNYYVIGSMEERWQGKGAELLGLEGKVDKQVFTELLQGKLPDGSDLTRIQ
DGVNKHRPGYDLTFSAPKSVSMLAMLGGDKRLIDAHNRAVTVALNQVESLASTRVKKDGVSETVLTGNLI
IARFNHDTSRAQDPQIHTHSVVINATQNGDKWLTLASDTVGKTGFSETILANRIAFGKIYQNSLRADVES
MGYKTVDAGRNGMWEMEGVPVESFSTRSQELREAAGPDASLKSRDVAALDTRKSKEAIDPAEKMVEWMNT
LKETGFDIRGYREAADARAAELARAPAAPVNTDGPDITDVVTKAIAGLSDRKVQFTYADLLARTVGQLEA
KDGVFELARKGIDAAIEREQLIPLDREKGLFTSNIHVLDELAVKALSQEVQRQNHVSVTPDASVVRQVPF
SDAVSVLAQDRPVMGIVSGQGGATGQRERVAELTLMAREQGRDVHILAADNRSRDFLAGDVRLAGETVTG
KSALQDGTAFIPGGTLIVDQAEKLSLKETISLLDGAMRHNVQVLLSDSGKRSGTGSALTVLKDSGVNTYR
WQGGHQTTADIISEPDKGARYSRLAQEFAVSVREGQESVAQISGTREQSVLNGLIRDSLRQEGVLGEKDT
TITALTPVWLDSKSRGVRDYYREGMVMERWDPETRTHDRFVIDRVTASSNMLTLKDREGDRLDLKVSAVD
SQWTLFRADTLPVAEGERLAVLGKIPDTRLKGGESVTVMKVEDGQLTVQRPGQKTTQTLAVGAGVFDGIK
VGHGWVESPGRSVSETATVFASVTQRELDNATLNQLAQSGSHLRLYSAQDAARTTEKLSRHTAFSVVSEQ
LKSRSGETDLDAAIAQQKAGLHTPAEQAIHLAIPLLESQDLTFSRPQLLATAMETGGGKVSMADIDTTIQ
AQIRSGQLLNVPVAPGRGNDLLISRQAWDAEKSILTRVLEGKDAVAPLMDRVPDSLMTELTAGQRAATRM
ILESTDRFTVVQGYAGVGKTTQFRAVMSAISLLPEETRPRVIGLAPTHRAVGEMQSAGVDARTTASFLHD
TQLLQRNGQTPDFSNTLFLLDESSMVGLADMAKAHSLIAAGGGRAVPSGDTDQLPPIASGQPFRLTQQRS
AADVAIMKEIVRQVPELRPAVYSLIERDVHRALTTIEQVTPEQVPRKEGAWAPGSSVVEFTPKQEKAIEK
ALSEGKTLPEGQPATLYEALVKDYTGRTPEAQSQTLVITHLNKDRRALNSLIHDARRENGETGKEEITLP
VLVTSNIRDGELRKLSTWTAHKEAVALVDNVYHRISKVDKDNQLITLTDSEGKERFISPREASAEGVTLY
RQEKITVSQGDRMRFSKSDPERGYVANSIWEVQSVSGDSVTLSDGKITRTLTPKAEQAQQHIDLAYAITA
HGAQGASEPYAIALEGVAGGREQMASFESAYVALSRMKQHVQVYTDNREGWIKAIKNSPEKATAHDILEP
RNDRAVKTADLLFGRARPLDETAAGRAALQQSGLAQGSSPGKFISPGKKYPQPHVALPAFDKNGKAAGIW
LSPLTDRDGRLEAIGGEGRIMGNEDARFVALQNSRNGESLLAGNMGEGVRMARDNPDTGVVVRLAGDDRP
WNPGAMTGGRVWAEPAPVAPVPQAGADIILPPEVLAQRAAEEQQRREMEKQAEQTAREVAGEARKAGEPA
DRVKEVIGDVIRGLERDRPGTEKTTLPDDPQFRRQEEAIQQVASERLQRERLQAVERDMVRDLNREKTLG
GD
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4CP
| ID | 2123 | GenBank | WP_013214038 |
| Name | traD_HTL29_RS00660_p1220-CTXM |
UniProt ID | _ |
| Length | 770 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 770 a.a. Molecular weight: 86010.07 Da Isoelectric Point: 5.1787
>WP_013214038.1 MULTISPECIES: type IV conjugative transfer system coupling protein TraD [Enterobacterales]
MSFNAKDMTQGGQIANMRFRMFGQIANIIFYVLFILFWVLCGLMLMYRLSWQTFVNGCVYWWCTTLGPMR
DIIRSQPVYTIQYYGQSLEYTSEQILADKYTIWCGEQLWTSFVFAAVVSLVICIVTFFIASWVLGRQGKQ
QSEDENTGGRQLSDKPKEVARQMKRDGMASDIKIGDLPILKNSEIQNFCLHGTVGSGKSEVIRRLLNYVR
ARGDMAIIYDRSCEFVKSYYDPSLDKILNPLDSRCAAWDLWKECLTLPDFDNISNTLIPMGTKEDPFWQG
SGRTIFAEGAYLMREDKDRSYEKLVDTMLSIKIDKLRAYLQNTPAANLVEEKIEKTAISIRAVLTNYVKA
IRYLQGIEKNGEPFTIRDWMRGVREDRPNGWLFISSNADTHASLKPVISMWLSIAIRGLLAMGENRNRRV
WIFADELPTLHKLPDLVEILPEARKFGGCYVFGIQSYAQLEDIYGVKPAATLFDVMNTRAFFRSPSKEIA
EFAAGEIGEKEILKASEQYSYGADPVRDGVSTGKEKERETLVSYSDIQTLPDLSCYVTLPGPYPAVKLAL
KYKPRPKIAEGFIPRTLDARVDARLSALLEAREAEGSLARALFTPDAPASGPADTNSHAGEQPEPVSQPA
PADMTVSPEPVKAPPTIKRPAAEPSVRTTEPSVLRVTTVPLIKPKAAAAAAAASTASSSGAPATAAGGTQ
QELAQQSAEQGQDMLPAGMNEDGVIEDMQAYDAWLADEQTQRDMQRREEVNINHSHRHDEQDDVEIGGNF
MSFNAKDMTQGGQIANMRFRMFGQIANIIFYVLFILFWVLCGLMLMYRLSWQTFVNGCVYWWCTTLGPMR
DIIRSQPVYTIQYYGQSLEYTSEQILADKYTIWCGEQLWTSFVFAAVVSLVICIVTFFIASWVLGRQGKQ
QSEDENTGGRQLSDKPKEVARQMKRDGMASDIKIGDLPILKNSEIQNFCLHGTVGSGKSEVIRRLLNYVR
ARGDMAIIYDRSCEFVKSYYDPSLDKILNPLDSRCAAWDLWKECLTLPDFDNISNTLIPMGTKEDPFWQG
SGRTIFAEGAYLMREDKDRSYEKLVDTMLSIKIDKLRAYLQNTPAANLVEEKIEKTAISIRAVLTNYVKA
IRYLQGIEKNGEPFTIRDWMRGVREDRPNGWLFISSNADTHASLKPVISMWLSIAIRGLLAMGENRNRRV
WIFADELPTLHKLPDLVEILPEARKFGGCYVFGIQSYAQLEDIYGVKPAATLFDVMNTRAFFRSPSKEIA
EFAAGEIGEKEILKASEQYSYGADPVRDGVSTGKEKERETLVSYSDIQTLPDLSCYVTLPGPYPAVKLAL
KYKPRPKIAEGFIPRTLDARVDARLSALLEAREAEGSLARALFTPDAPASGPADTNSHAGEQPEPVSQPA
PADMTVSPEPVKAPPTIKRPAAEPSVRTTEPSVLRVTTVPLIKPKAAAAAAAASTASSSGAPATAAGGTQ
QELAQQSAEQGQDMLPAGMNEDGVIEDMQAYDAWLADEQTQRDMQRREEVNINHSHRHDEQDDVEIGGNF
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 77740..106352
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| HTL29_RS00465 | 73132..73359 | + | 228 | WP_001568050 | hypothetical protein | - |
| HTL29_RS00470 | 73451..73681 | + | 231 | WP_001568051 | hypothetical protein | - |
| HTL29_RS00475 | 73733..75088 | + | 1356 | WP_014343515 | DUF3560 domain-containing protein | - |
| HTL29_RS00480 | 75136..75699 | + | 564 | WP_014343514 | methyltransferase | - |
| HTL29_RS00485 | 76524..77345 | + | 822 | WP_004152492 | DUF932 domain-containing protein | - |
| HTL29_RS00490 | 77378..77707 | + | 330 | WP_013214015 | DUF5983 family protein | - |
| HTL29_RS00495 | 77740..78225 | - | 486 | WP_014343495 | transglycosylase SLT domain-containing protein | virB1 |
| HTL29_RS00500 | 78616..79032 | + | 417 | WP_014343494 | conjugal transfer relaxosome DNA-binding protein TraM | - |
| HTL29_RS00505 | 79232..79969 | + | 738 | WP_013214018 | conjugal transfer protein TrbJ | - |
| HTL29_RS00510 | 80084..80290 | + | 207 | WP_171773970 | TraY domain-containing protein | - |
| HTL29_RS00515 | 80352..80720 | + | 369 | WP_004194426 | type IV conjugative transfer system pilin TraA | - |
| HTL29_RS00520 | 80734..81039 | + | 306 | WP_004144424 | type IV conjugative transfer system protein TraL | traL |
| HTL29_RS00525 | 81059..81625 | + | 567 | WP_004144423 | type IV conjugative transfer system protein TraE | traE |
| HTL29_RS00530 | 81612..82352 | + | 741 | WP_013214019 | type-F conjugative transfer system secretin TraK | traK |
| HTL29_RS00535 | 82352..83776 | + | 1425 | WP_013214020 | F-type conjugal transfer pilus assembly protein TraB | traB |
| HTL29_RS00540 | 83848..84432 | + | 585 | WP_014343493 | type IV conjugative transfer system lipoprotein TraV | traV |
| HTL29_RS00545 | 84755..84973 | + | 219 | WP_004195468 | hypothetical protein | - |
| HTL29_RS00550 | 84974..85285 | + | 312 | WP_013214022 | hypothetical protein | - |
| HTL29_RS00555 | 85352..85756 | + | 405 | WP_004197817 | hypothetical protein | - |
| HTL29_RS00560 | 86133..86531 | + | 399 | WP_014343490 | hypothetical protein | - |
| HTL29_RS00565 | 86603..89242 | + | 2640 | WP_013214024 | type IV secretion system protein TraC | virb4 |
| HTL29_RS00570 | 89242..89631 | + | 390 | WP_013214025 | type-F conjugative transfer system protein TrbI | - |
| HTL29_RS00575 | 89631..90266 | + | 636 | WP_013214026 | type-F conjugative transfer system protein TraW | traW |
| HTL29_RS00580 | 90301..90702 | + | 402 | WP_004194979 | hypothetical protein | - |
| HTL29_RS00585 | 90699..91688 | + | 990 | WP_013214027 | conjugal transfer pilus assembly protein TraU | traU |
| HTL29_RS00590 | 91701..92339 | + | 639 | WP_004193871 | type-F conjugative transfer system pilin assembly protein TrbC | trbC |
| HTL29_RS00595 | 92398..94353 | + | 1956 | WP_014343489 | type-F conjugative transfer system mating-pair stabilization protein TraN | traN |
| HTL29_RS00600 | 94385..94639 | + | 255 | WP_004152674 | conjugal transfer protein TrbE | - |
| HTL29_RS00605 | 94617..94865 | + | 249 | WP_013214029 | hypothetical protein | - |
| HTL29_RS00610 | 94878..95204 | + | 327 | WP_013214030 | hypothetical protein | - |
| HTL29_RS00615 | 95225..95977 | + | 753 | WP_004152677 | type-F conjugative transfer system pilin assembly protein TraF | traF |
| HTL29_RS00620 | 95988..96227 | + | 240 | WP_004144400 | type-F conjugative transfer system pilin chaperone TraQ | - |
| HTL29_RS00625 | 96199..96756 | + | 558 | WP_013214031 | type-F conjugative transfer system pilin assembly thiol-disulfide isomerase TrbB | traF |
| HTL29_RS00630 | 96802..97245 | + | 444 | WP_013214032 | F-type conjugal transfer protein TrbF | - |
| HTL29_RS00635 | 97223..98602 | + | 1380 | WP_074160414 | conjugal transfer pilus assembly protein TraH | traH |
| HTL29_RS00640 | 98602..101424 | + | 2823 | WP_013214034 | conjugal transfer mating-pair stabilization protein TraG | traG |
| HTL29_RS00645 | 101447..102049 | + | 603 | WP_013214035 | conjugal transfer protein TraS | - |
| HTL29_RS00650 | 102300..103031 | + | 732 | WP_013214036 | conjugal transfer complement resistance protein TraT | - |
| HTL29_RS00655 | 103224..103913 | + | 690 | WP_014343485 | hypothetical protein | - |
| HTL29_RS00660 | 104040..106352 | + | 2313 | WP_013214038 | type IV conjugative transfer system coupling protein TraD | virb4 |
Host bacterium
| ID | 3617 | GenBank | NZ_KY174332 |
| Plasmid name | p1220-CTXM | Incompatibility group | IncFII |
| Plasmid size | 137060 bp | Coordinate of oriT [Strand] | 78298..78347 [-] |
| Host baterium | Klebsiella pneumoniae strain 1220 |
Cargo genes
| Drug resistance gene | blaCTX-M-14, sul1, blaDHA-1, qnrB4 |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | AcrIE9 |