Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 102978 |
| Name | oriT_FDAARGOS_95|unnamed |
| Organism | Escherichia coli strain FDAARGOS_95 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP014090 (65535..65819 [-], 285 nt) |
| oriT length | 285 nt |
| IRs (inverted repeats) | 184..189, 191..196 (AAAAGT..ACTTTT) |
| Location of nic site | 109..110 |
| Conserved sequence flanking the nic site |
TTTGGTTAAA |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 285 nt
>oriT_FDAARGOS_95|unnamed
GGTTATTGCTACTTAATGCCGATAACGACTCAGGCTTTGAGGTTTTTTTATACGGTTCACATTTCGTTAGCAAGGTCAGGGTTTTTTGATAAAATTCTGGTTAGTTTGGTTAAAAAGTGTTACAAGTAAGGGTAATGGCTGAAAGGTTAGTTTTAAGGTTCAAAGCGGCAGTATTAAAATTCCAAAAGTTACTTTTCATCCTTCAGAATCCAGACCTTAATTTCATGTAGAAGATTCGTACAATTGTATTGGCGCAAGGACAATCCGCACATGTCAGAATCAGAT
GGTTATTGCTACTTAATGCCGATAACGACTCAGGCTTTGAGGTTTTTTTATACGGTTCACATTTCGTTAGCAAGGTCAGGGTTTTTTGATAAAATTCTGGTTAGTTTGGTTAAAAAGTGTTACAAGTAAGGGTAATGGCTGAAAGGTTAGTTTTAAGGTTCAAAGCGGCAGTATTAAAATTCCAAAAGTTACTTTTCATCCTTCAGAATCCAGACCTTAATTTCATGTAGAAGATTCGTACAATTGTATTGGCGCAAGGACAATCCGCACATGTCAGAATCAGAT
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file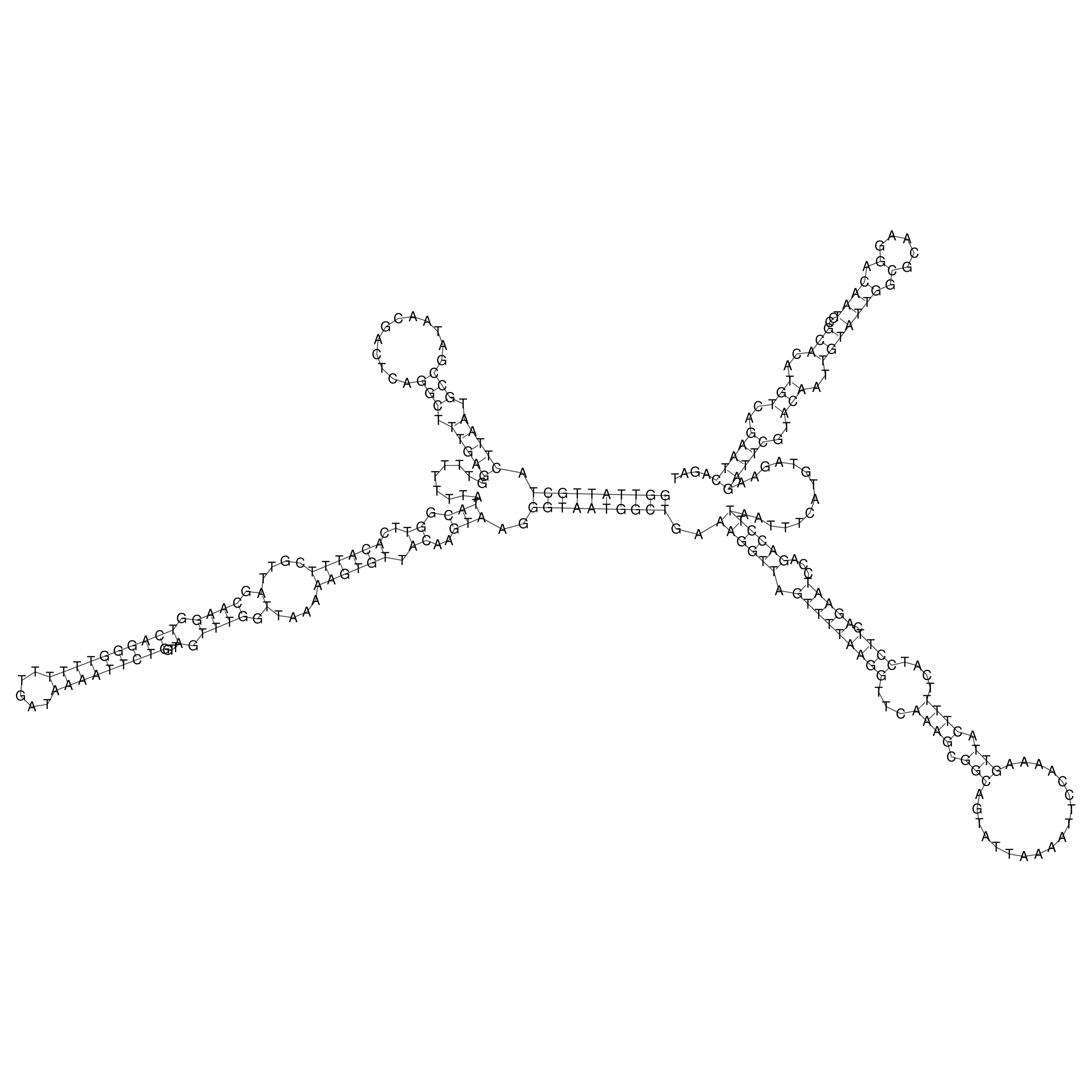
Relaxase
| ID | 2233 | GenBank | WP_016695058 |
| Name | TraI_2_AL551_RS01645_FDAARGOS_95|unnamed |
UniProt ID | A0A140CLT5 |
| Length | 1011 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1011 a.a. Molecular weight: 113053.03 Da Isoelectric Point: 4.5108
>WP_016695058.1 MULTISPECIES: MobH family relaxase [Enterobacteriaceae]
MNFRALFLSMQRVFGIFSRRENDVSELMMKDAANFSPFAQIIGEQKYTVPDHPNPEVLKFIEYPTRPAGI
QTFNEQSILSLYRDKLHSISMMLAISDGDIREDAYTFTNLVLKPLIEYIRWIHLLPASENHHHNGIGGLL
SHSLEVAMISLKNANHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLSETLTWAPS
SQSLLDWARENNVVEYEIHWRKRIHNQHNIWSSVFLERILDPVCMSFLDRVKKERVYAKMVTALNVYNDG
NDFLSKCVRTSDYYSTGTDLNVLRDPIMGLRSNDAAARAIGTIKHNFTSININNYKTKPMHIIIVNGEVY
LNENAFLDFVLSDFAAHKFNFPQGDAGKTVLVESLVQRGYVEPYDDERVVHYFIPGTYSENEIASIFRNG
IGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRESVTRVTDTVN
DAIENTPQYGLQLVNSPDADSNNIISSENTESITDSLEESGADISNEIFETQVVTAIDTAETVNADEPEQ
VEEHDDRSQIHLVEQLHEMLLSAPLPHHAVINIDSVPYLDLDAAIALIPGIDEAAFCNGPFFQLTYRDGS
LDGMWIVRDVNNLRLIQLGDNCAGMQVSTSEPRNTSSLKSLFDTSMYQPLDIPEAPSVNEAASPPQTPLE
LPQPRLNAPVAEEASSVAEQTNAHSEPDSVIATEYEQYGHLLEETLDSDGEAYSDLIASDSTEAEYPATD
PQSSDFAQLPRETALSVAPGDLDYSEGAIKPPAPDATGKETILTSPEPAEDVRETVAAVEKASHLSPALA
RLFAVSTHAEKKHEKTQEPSPVKEVKNPTSSTTVKAPISIEPPGAEEKEAVEEFTLLNDGEVTELEYVEI
ATMLHQILTKLSGSFKRKRKNRFMVLTQNTFYLTQSCIEKYGTQLNAPELFNQLPQYQVTSGAVVNTKCI
AFNIPTLVAASDRAKVDIELIINKLKEVGNL
MNFRALFLSMQRVFGIFSRRENDVSELMMKDAANFSPFAQIIGEQKYTVPDHPNPEVLKFIEYPTRPAGI
QTFNEQSILSLYRDKLHSISMMLAISDGDIREDAYTFTNLVLKPLIEYIRWIHLLPASENHHHNGIGGLL
SHSLEVAMISLKNANHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLSETLTWAPS
SQSLLDWARENNVVEYEIHWRKRIHNQHNIWSSVFLERILDPVCMSFLDRVKKERVYAKMVTALNVYNDG
NDFLSKCVRTSDYYSTGTDLNVLRDPIMGLRSNDAAARAIGTIKHNFTSININNYKTKPMHIIIVNGEVY
LNENAFLDFVLSDFAAHKFNFPQGDAGKTVLVESLVQRGYVEPYDDERVVHYFIPGTYSENEIASIFRNG
IGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRESVTRVTDTVN
DAIENTPQYGLQLVNSPDADSNNIISSENTESITDSLEESGADISNEIFETQVVTAIDTAETVNADEPEQ
VEEHDDRSQIHLVEQLHEMLLSAPLPHHAVINIDSVPYLDLDAAIALIPGIDEAAFCNGPFFQLTYRDGS
LDGMWIVRDVNNLRLIQLGDNCAGMQVSTSEPRNTSSLKSLFDTSMYQPLDIPEAPSVNEAASPPQTPLE
LPQPRLNAPVAEEASSVAEQTNAHSEPDSVIATEYEQYGHLLEETLDSDGEAYSDLIASDSTEAEYPATD
PQSSDFAQLPRETALSVAPGDLDYSEGAIKPPAPDATGKETILTSPEPAEDVRETVAAVEKASHLSPALA
RLFAVSTHAEKKHEKTQEPSPVKEVKNPTSSTTVKAPISIEPPGAEEKEAVEEFTLLNDGEVTELEYVEI
ATMLHQILTKLSGSFKRKRKNRFMVLTQNTFYLTQSCIEKYGTQLNAPELFNQLPQYQVTSGAVVNTKCI
AFNIPTLVAASDRAKVDIELIINKLKEVGNL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A140CLT5 |
T4CP
| ID | 1921 | GenBank | WP_236923368 |
| Name | t4cp1_AL551_RS01650_FDAARGOS_95|unnamed |
UniProt ID | _ |
| Length | 402 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 402 a.a. Molecular weight: 46033.27 Da Isoelectric Point: 9.3172
>WP_236923368.1 ATP-binding protein [Escherichia coli]
MTKSKRTNLHAQENFYRPILEYRSASVLLICAVIMLVMGFRSDGVNIAPIILYTAVFLLLVCLYRCKTAH
PYLMAHWRVFQRQIMFISLKSLRTINKSNFFSNERKYRQLVQEYKKNNRPVPDRKTYFCNGFEWGPEHAD
RAYQIANLSSDKREIALPFVLSPIARHFETMARSMGGNNAIFAVDRRAPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVIVVIDPKNDAEWRQSLMDEASELGLPFYKFHPAQPSSSVCIDVCNAYTNVSD
LTSRLLSLVSVPGEVNPFVQYAEALVSTVITGLSYTDKKPSIYLIHKNMKSHMSVVNLTIKVMECCFARH
YGPDVWMEKVKYASNDTLQVRFKRLTEWFNAHFLNYEGAEPIELNRPGNPGD
MTKSKRTNLHAQENFYRPILEYRSASVLLICAVIMLVMGFRSDGVNIAPIILYTAVFLLLVCLYRCKTAH
PYLMAHWRVFQRQIMFISLKSLRTINKSNFFSNERKYRQLVQEYKKNNRPVPDRKTYFCNGFEWGPEHAD
RAYQIANLSSDKREIALPFVLSPIARHFETMARSMGGNNAIFAVDRRAPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVIVVIDPKNDAEWRQSLMDEASELGLPFYKFHPAQPSSSVCIDVCNAYTNVSD
LTSRLLSLVSVPGEVNPFVQYAEALVSTVITGLSYTDKKPSIYLIHKNMKSHMSVVNLTIKVMECCFARH
YGPDVWMEKVKYASNDTLQVRFKRLTEWFNAHFLNYEGAEPIELNRPGNPGD
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Host bacterium
| ID | 3421 | GenBank | NZ_CP014090 |
| Plasmid name | FDAARGOS_95|unnamed | Incompatibility group | IncFIA |
| Plasmid size | 86763 bp | Coordinate of oriT [Strand] | 65535..65819 [-] |
| Host baterium | Escherichia coli strain FDAARGOS_95 |
Cargo genes
| Drug resistance gene | aph(6)-Id, aph(3'')-Ib, tet(B) |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |