Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 102690 |
| Name | oriT_pBamB1895 |
| Organism | Bacillus amyloliquefaciens strain B1895 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_OK210095 (30152..30513 [+], 362 nt) |
| oriT length | 362 nt |
| IRs (inverted repeats) | 290..295, 304..309 (GAAAAA..TTTTTC) 291..296, 302..307 (AAAAAG..CTTTTT) 188..193, 198..203 (CACCAT..ATGGTG) 171..178, 180..187 (AAACCGCA..TGCGGTTT) 116..124, 128..136 (ACATAAACA..TGTTTATGT) 105..110, 117..122 (TTTATG..CATAAA) 62..69, 73..80 (CATAAATT..AATTTATG) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 362 nt
>oriT_pBamB1895
AAAGAGCAATCTCGTCATCGGGGATTAGATTTTAGGGTGAAATGTAGTTGATTTGCGTGTGCATAAATTGAGAATTTATGTGCTTTTCGATTTAAAACACCTAATTTATGGTTGTACATAAACATTTTGTTTATGTGCATTGAGCAATAAATCTGGTACCACGACAAAACAAACCGCACTGCGGTTTCACCATGGGAATGGTGCCAGTTTCCCCTTATGCTCTTTCCTTTTCCACCCCCTCCTTCCTTCGGAATCGGGGGCCGGTTTTTTGCTGCCGCAAAAAACCTTGGAAAAAGAAACACTTTTTTCGAACAGATCATTGCAAAATAAAGTGCCAAAACTGATTAAAAGGAGCGTTCACA
AAAGAGCAATCTCGTCATCGGGGATTAGATTTTAGGGTGAAATGTAGTTGATTTGCGTGTGCATAAATTGAGAATTTATGTGCTTTTCGATTTAAAACACCTAATTTATGGTTGTACATAAACATTTTGTTTATGTGCATTGAGCAATAAATCTGGTACCACGACAAAACAAACCGCACTGCGGTTTCACCATGGGAATGGTGCCAGTTTCCCCTTATGCTCTTTCCTTTTCCACCCCCTCCTTCCTTCGGAATCGGGGGCCGGTTTTTTGCTGCCGCAAAAAACCTTGGAAAAAGAAACACTTTTTTCGAACAGATCATTGCAAAATAAAGTGCCAAAACTGATTAAAAGGAGCGTTCACA
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file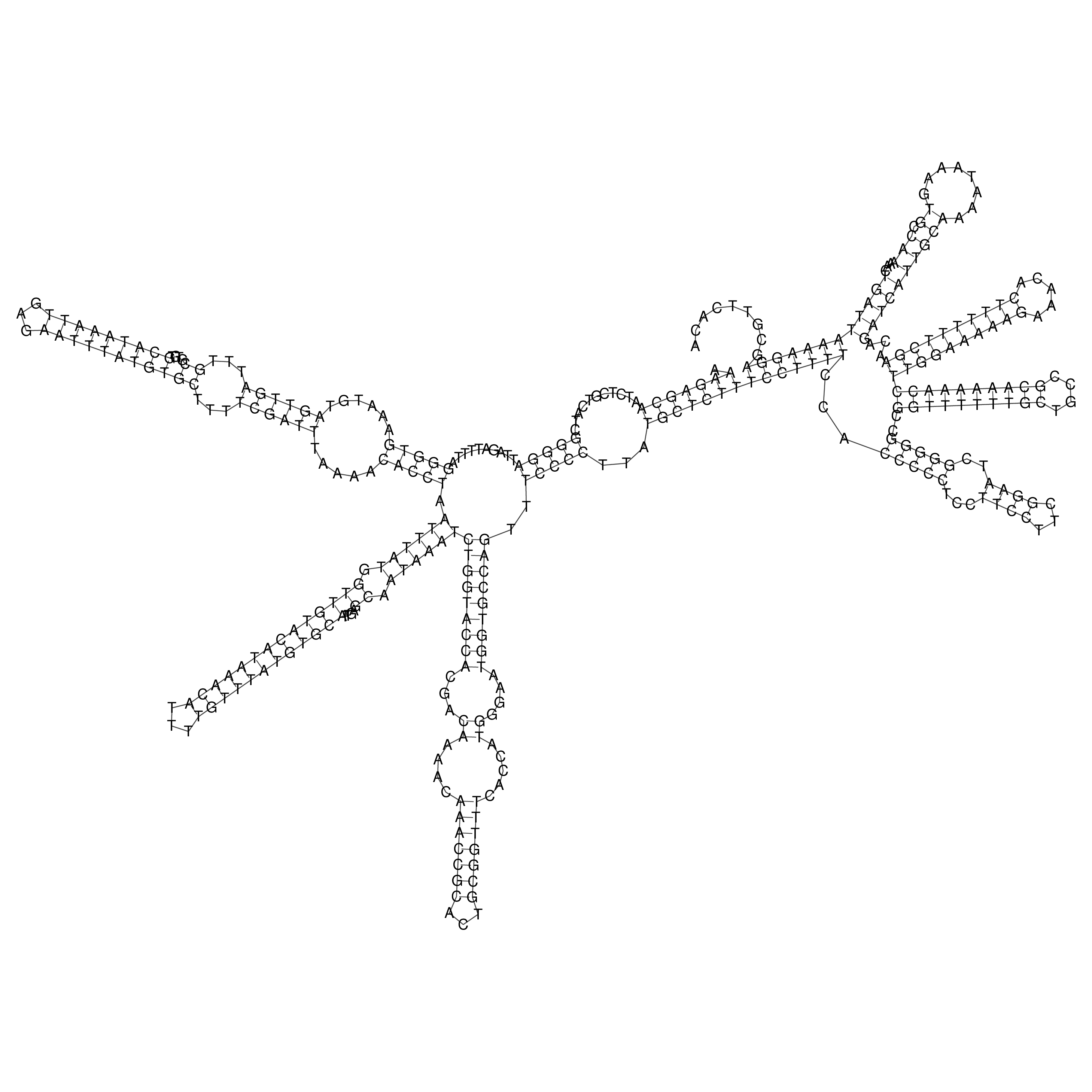
Relaxase
| ID | 1983 | GenBank | WP_208557299 |
| Name | mobL_LUX07_RS00170_pBamB1895 |
UniProt ID | _ |
| Length | 410 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 410 a.a. Molecular weight: 48348.01 Da Isoelectric Point: 9.8707
>WP_208557299.1 MULTISPECIES: MobP2 family relaxase [Bacillus]
MDAPGVVLVSKYVSGKSTKFSKYVNYINRDEAVRTEKFQTYNVNKLDGYNQYMGNPEKSSGIFTQFKDTL
SPQEKNQLKEIFRQAQENDSVMWQDVISFDNKWLEEHGIYKPDTGWINEGAIQNSIRKGMKVLLKEEQLE
QSGVWSAAIHYNTDNIHVHIALVEPNPTKEYGVFTNKKTGEVYQARRGNRKLKTLDKMKSKVANTLLDRD
KELAKISQLIHDRIAPKGVKFQPRLDTYMSKMYNHIYANLPEDMRLWRYNNNALNFIRPEIDSVITMYIQ
KYHPEDFKELDQSLKEEMEFRKSVYGDGPIQVERYKEYRENKHKELYAKLGNSMLKEMVEIRKRESQTKQ
FVRTGTYAPGSASQKQWDAKGKLIKRSDINKIKHALDKDYQSMKNMRKYQQMQYEMEQSR
MDAPGVVLVSKYVSGKSTKFSKYVNYINRDEAVRTEKFQTYNVNKLDGYNQYMGNPEKSSGIFTQFKDTL
SPQEKNQLKEIFRQAQENDSVMWQDVISFDNKWLEEHGIYKPDTGWINEGAIQNSIRKGMKVLLKEEQLE
QSGVWSAAIHYNTDNIHVHIALVEPNPTKEYGVFTNKKTGEVYQARRGNRKLKTLDKMKSKVANTLLDRD
KELAKISQLIHDRIAPKGVKFQPRLDTYMSKMYNHIYANLPEDMRLWRYNNNALNFIRPEIDSVITMYIQ
KYHPEDFKELDQSLKEEMEFRKSVYGDGPIQVERYKEYRENKHKELYAKLGNSMLKEMVEIRKRESQTKQ
FVRTGTYAPGSASQKQWDAKGKLIKRSDINKIKHALDKDYQSMKNMRKYQQMQYEMEQSR
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4CP
| ID | 1624 | GenBank | WP_151276507 |
| Name | t4cp2_LUX07_RS00120_pBamB1895 |
UniProt ID | _ |
| Length | 786 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 786 a.a. Molecular weight: 90705.99 Da Isoelectric Point: 4.8260
>WP_151276507.1 MULTISPECIES: VirD4-like conjugal transfer protein, CD1115 family [Bacillus amyloliquefaciens group]
MHKEFKQWKKIFADKYFLIAFTLFCYAAVTCAANFLFHLFKQIPLMLSSFKTFDQSKLSSPFDSYSWKWF
FEFEWDMGIIYGVLYIIASIFIVRQVYRFRIAFRDINKQTKGTARWTTIKEIQETYKAIKDDDVEYEGSA
GMPIVHFNDHLYVDTNSTHTLVVASTQSGKTETYSYPYLDVIMRAKKKDSVVITDIKGDMLKNTHSEFEK
HGYEVMCFNLLNPYWGIAYNPLELVKQAYMKGDFSKAQMLCNTLSYSLFHNDKSHADPMWENASIALVNA
LILAVCDLCIKNEQPENITMYTLTVMLNELGSNPDEDGYTRLDHFFGNLPPSHPAKLQYGTIQFSQGVTR
SGIYTGTMAKLKNYTYDTIARMTAQNDLNIEDLAYGEKPVALFIVYPDWDDSNYTIISTFLSQVNAVLSE
KATLSKESTLPRKVRFLFEEVANIPPIEGLNRSLAVGLSRGMLYTLVIQNVSQLRDVYGDDMATAIMGNL
GNQIYIMSDEWEDAEKFSEKLGVTTIISADRQGGLMDVDKSYSEREEERPLMLPDELRRLKKGEWVVLRT
KKRENLKRKRVVPYPIFASLDNETHMLHRYEYLMHRFNNKIALGDLGFHGNHETLDLESLLIEFEFNEPV
EQKNKVKGKKRSDKKDADPEVPDYNEIPPPEEPIGIADESSEATDSLFELVPSDNTNEEEFYEGPCLIDS
SKNEVLEVSEEDAIMNSLEEVYETSTPISDAIKADQYVYIKFFAQKALTEDEYNYFESLTTIEQLRAFFR
APEKHELYEKIKKHIE
MHKEFKQWKKIFADKYFLIAFTLFCYAAVTCAANFLFHLFKQIPLMLSSFKTFDQSKLSSPFDSYSWKWF
FEFEWDMGIIYGVLYIIASIFIVRQVYRFRIAFRDINKQTKGTARWTTIKEIQETYKAIKDDDVEYEGSA
GMPIVHFNDHLYVDTNSTHTLVVASTQSGKTETYSYPYLDVIMRAKKKDSVVITDIKGDMLKNTHSEFEK
HGYEVMCFNLLNPYWGIAYNPLELVKQAYMKGDFSKAQMLCNTLSYSLFHNDKSHADPMWENASIALVNA
LILAVCDLCIKNEQPENITMYTLTVMLNELGSNPDEDGYTRLDHFFGNLPPSHPAKLQYGTIQFSQGVTR
SGIYTGTMAKLKNYTYDTIARMTAQNDLNIEDLAYGEKPVALFIVYPDWDDSNYTIISTFLSQVNAVLSE
KATLSKESTLPRKVRFLFEEVANIPPIEGLNRSLAVGLSRGMLYTLVIQNVSQLRDVYGDDMATAIMGNL
GNQIYIMSDEWEDAEKFSEKLGVTTIISADRQGGLMDVDKSYSEREEERPLMLPDELRRLKKGEWVVLRT
KKRENLKRKRVVPYPIFASLDNETHMLHRYEYLMHRFNNKIALGDLGFHGNHETLDLESLLIEFEFNEPV
EQKNKVKGKKRSDKKDADPEVPDYNEIPPPEEPIGIADESSEATDSLFELVPSDNTNEEEFYEGPCLIDS
SKNEVLEVSEEDAIMNSLEEVYETSTPISDAIKADQYVYIKFFAQKALTEDEYNYFESLTTIEQLRAFFR
APEKHELYEKIKKHIE
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Host bacterium
| ID | 3134 | GenBank | NZ_OK210095 |
| Plasmid name | pBamB1895 | Incompatibility group | - |
| Plasmid size | 63409 bp | Coordinate of oriT [Strand] | 30152..30513 [+] |
| Host baterium | Bacillus amyloliquefaciens strain B1895 |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |