Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 102055 |
| Name | oriT_ISU 995|unnamed4 |
| Organism | Staphylococcus aureus strain ISU 995 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_LKYO01000056 (1680..1720 [-], 41 nt) |
| oriT length | 41 nt |
| IRs (inverted repeats) | 2..8, 13..19 (ACTTTAT..ATAAAGT) |
| Location of nic site | _ |
| Conserved sequence flanking the nic site |
_ |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 41 nt
>oriT_ISU 995|unnamed4
CACTTTATGAATATAAAGTATAATGTGTTATACTTTACATG
CACTTTATGAATATAAAGTATAATGTGTTATACTTTACATG
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file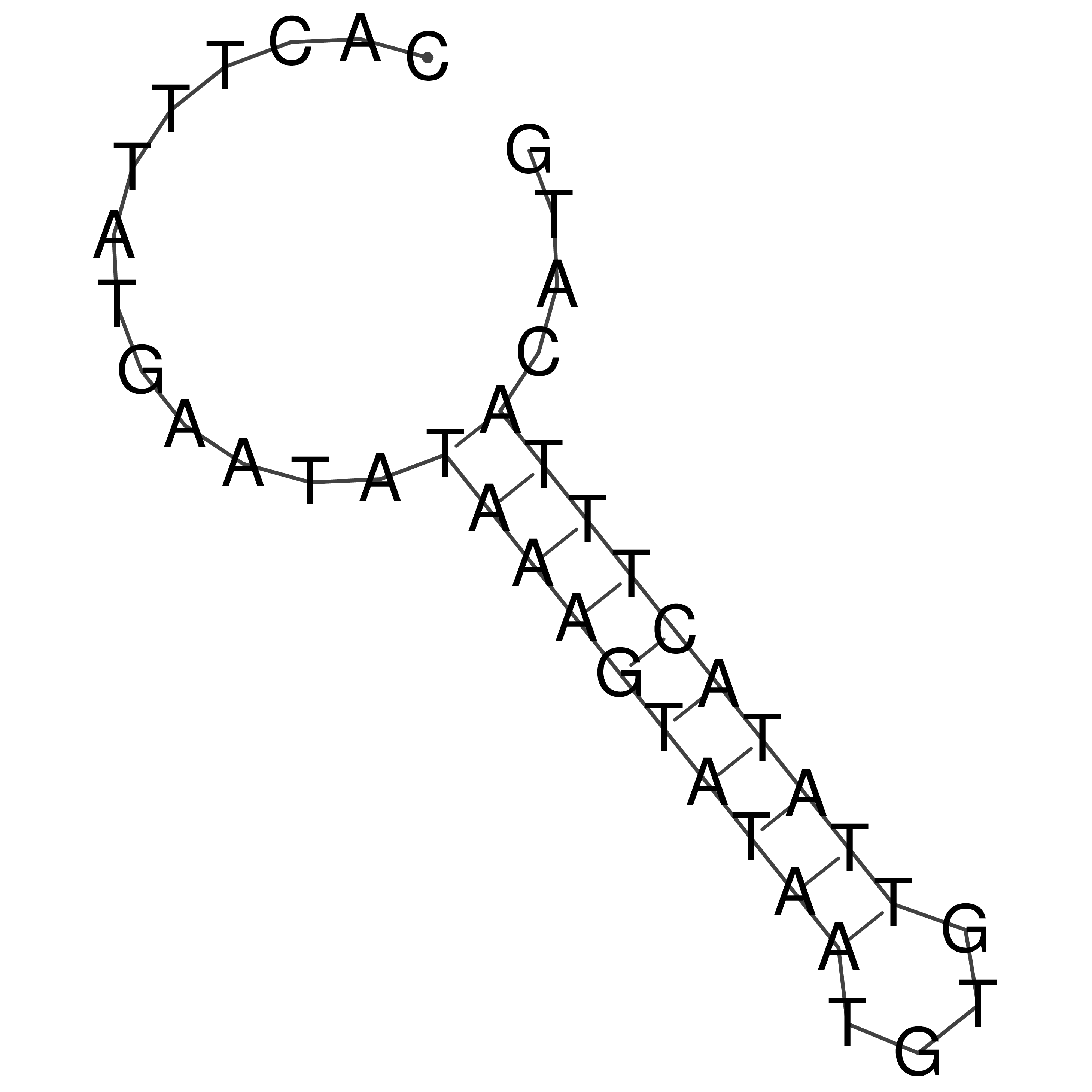
Relaxase
| ID | 1642 | GenBank | WP_015382137 |
| Name | mobV_APW15_RS03635_ISU 995|unnamed4 |
UniProt ID | L8BAH8 |
| Length | 409 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 409 a.a. Molecular weight: 48447.75 Da Isoelectric Point: 7.0106
>WP_015382137.1 MULTISPECIES: MobV family relaxase [Bacilli]
MSMIVARMQKMKAENLVGIGNHNQRKTKNHSNPDIDTSLSKLNYDLVDRTQNYKTDIENFINENKTTTRA
VRKDAVLVNEWIISSDKDFFDNLTESEIENFFERSKDYFAEKFGEKNIRYATVHLDESTPHMHMGIVPFD
KDNKLSAKRVFNRQALRDVQEELPKYLQDFQFEIERGQKGSERKNLTVPEFKKLKEEEREIKKELEIKKD
ELMAYTKENKIDKEIDITPIKEMEDVEIETDEKSLFGITKTKTVRQWTGNIVLSEKDYLKLRQKVNKGKQ
AEGKLEAILETDVYQENKELKNELKDQIDKNDKDIDDYNDLVKRYNNLYEENTSLKNQIGDLKEEIKLIY
QSTKRFFKDRISDFKAFKEVFKELADSISNISREKGLDSSFKKEFDRENKKKRTKGMSR
MSMIVARMQKMKAENLVGIGNHNQRKTKNHSNPDIDTSLSKLNYDLVDRTQNYKTDIENFINENKTTTRA
VRKDAVLVNEWIISSDKDFFDNLTESEIENFFERSKDYFAEKFGEKNIRYATVHLDESTPHMHMGIVPFD
KDNKLSAKRVFNRQALRDVQEELPKYLQDFQFEIERGQKGSERKNLTVPEFKKLKEEEREIKKELEIKKD
ELMAYTKENKIDKEIDITPIKEMEDVEIETDEKSLFGITKTKTVRQWTGNIVLSEKDYLKLRQKVNKGKQ
AEGKLEAILETDVYQENKELKNELKDQIDKNDKDIDDYNDLVKRYNNLYEENTSLKNQIGDLKEEIKLIY
QSTKRFFKDRISDFKAFKEVFKELADSISNISREKGLDSSFKKEFDRENKKKRTKGMSR
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | L8BAH8 |
Host bacterium
| ID | 2499 | GenBank | NZ_LKYO01000056 |
| Plasmid name | ISU 995|unnamed4 | Incompatibility group | - |
| Plasmid size | 2206 bp | Coordinate of oriT [Strand] | 1680..1720 [-] |
| Host baterium | Staphylococcus aureus strain ISU 995 |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |