Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 101944 |
| Name | oriT_158900|unnamed2 |
| Organism | Salmonella enterica subsp. enterica serovar Java strain 158900 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_VUIY01000028 (2697..3303 [-], 607 nt) |
| oriT length | 607 nt |
| IRs (inverted repeats) | 590..595, 600..605 (ATGCCC..GGGCAT) 517..523, 529..535 (CCTCCCG..CGGGAGG) 453..458, 462..467 (GTTCGC..GCGAAC) 196..202, 204..210 (TCAATTC..GAATTGA) 175..180, 187..192 (AAATCA..TGATTT) 79..84, 93..98 (GCTTTA..TAAAGC) 29..37, 46..54 (TTTTGATAA..TTATCAAAA) |
| Location of nic site | 313..314 |
| Conserved sequence flanking the nic site |
TCCTGCATCG |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 607 nt
>oriT_158900|unnamed2
CTTTGTTTACCTGTTAAGTACATCGTTGTTTTGATAAAATCATTATTATCAAAAAACGTATTTATATCCTTACTGCCAGCTTTACTTTTTAATAAAGCATTTGCCAAAAGTTTCATGCCTTTAGCTACTGTTAAAGCTGGTTCATCTGGATAAAGAGTTTTAAGGCAATCCAGCAAATCATCATGCTGATTTTCATCAATTCTGAATTGAACTTTTTTCATAAAATTCTCCAAAAAGAACCGACTGTAGGTCACCGGGCAAACGTTGCGGAATGGCGTCAGAGACGTCATTTTGCGGCGTTTGCCCTATCCTGCATCGCAGTGAATCTGGCGCCGCTCATAAATTGTGCTTGAGCATCAACAAAAATGCAAAAAGGCTGATGTTGTCATGGCTGAACATACGTACGTCTGCGACCGGAAACCCGATCAATGCGGTAAGGACAGTGCCGTCAGGTTCGCTCCGCGAACGGTCTGGCAGAAAGCGCAAGCGCAGCCCCGCGCAATATATCAGAAAGCACCTCCCGAAATACGGGAGGGCTTCGGCTATTCAGCTAAGGTTGCTCTGTTCAGGTTGCTGAGTGTAGCCTTCCATGCCCTGACGGGCATCA
CTTTGTTTACCTGTTAAGTACATCGTTGTTTTGATAAAATCATTATTATCAAAAAACGTATTTATATCCTTACTGCCAGCTTTACTTTTTAATAAAGCATTTGCCAAAAGTTTCATGCCTTTAGCTACTGTTAAAGCTGGTTCATCTGGATAAAGAGTTTTAAGGCAATCCAGCAAATCATCATGCTGATTTTCATCAATTCTGAATTGAACTTTTTTCATAAAATTCTCCAAAAAGAACCGACTGTAGGTCACCGGGCAAACGTTGCGGAATGGCGTCAGAGACGTCATTTTGCGGCGTTTGCCCTATCCTGCATCGCAGTGAATCTGGCGCCGCTCATAAATTGTGCTTGAGCATCAACAAAAATGCAAAAAGGCTGATGTTGTCATGGCTGAACATACGTACGTCTGCGACCGGAAACCCGATCAATGCGGTAAGGACAGTGCCGTCAGGTTCGCTCCGCGAACGGTCTGGCAGAAAGCGCAAGCGCAGCCCCGCGCAATATATCAGAAAGCACCTCCCGAAATACGGGAGGGCTTCGGCTATTCAGCTAAGGTTGCTCTGTTCAGGTTGCTGAGTGTAGCCTTCCATGCCCTGACGGGCATCA
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file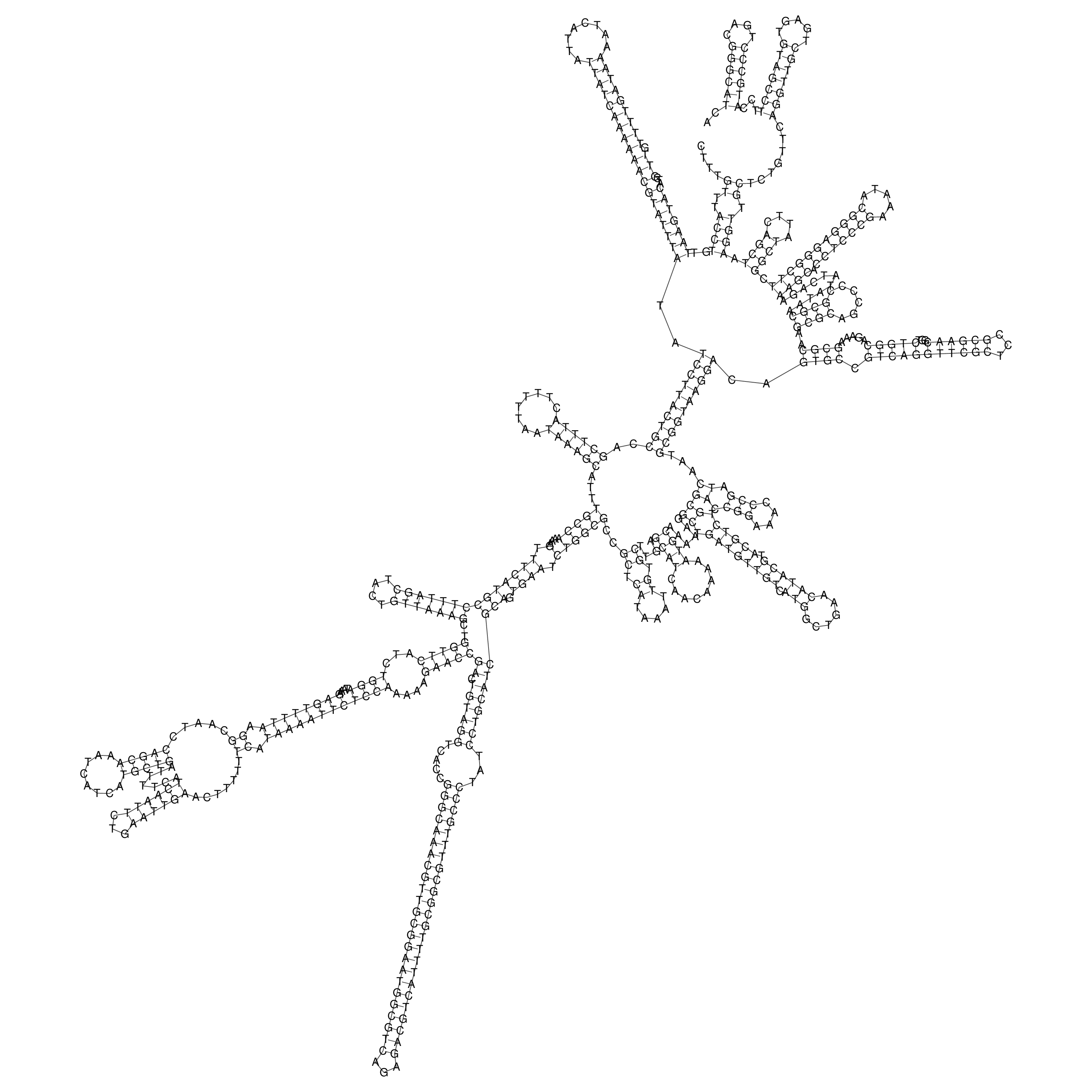
Relaxase
| ID | 1570 | GenBank | WP_000539538 |
| Name | mobP1_F1E34_RS22350_158900|unnamed2 |
UniProt ID | A0A5J2A6K9 |
| Length | 388 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 388 a.a. Molecular weight: 44507.83 Da Isoelectric Point: 10.5067
>WP_000539538.1 MULTISPECIES: MobP1 family relaxase [Enterobacterales]
MGVYVDKEYRVKRKSSENGRKSAFAHKVKNGGKNYSRNVQERINRKGASKEVVVKISGGAVTRQGIRNSI
DYMSRESELPVMSESGRVWTGDEILEAKDHMIDRANDPQHVMNDKGKENKKITQNIVFSPPVSAKVKPED
LLESVRKTMQKKYPNHRFVLGYHCDKKEHPHVHVVFRIRDNDGKRADIRKKDLREIRTGFCEELKLKGYD
VKATHKQQHGLNQSVKDAHNTAPKRQKGVYEVVDVGYDHYQNDKTKSKQHFIKLKTLNKGVEKTYWGADF
GDLCSRESVKAGDLVRLKKLGQKEVKIPALDKNGVQHGWKTVHRNEWQLENLGVKGVDRTPSASKELVLN
SPDMLLKQQQRMAQFTQQKASTLQSEQKLKTGIKFWSL
MGVYVDKEYRVKRKSSENGRKSAFAHKVKNGGKNYSRNVQERINRKGASKEVVVKISGGAVTRQGIRNSI
DYMSRESELPVMSESGRVWTGDEILEAKDHMIDRANDPQHVMNDKGKENKKITQNIVFSPPVSAKVKPED
LLESVRKTMQKKYPNHRFVLGYHCDKKEHPHVHVVFRIRDNDGKRADIRKKDLREIRTGFCEELKLKGYD
VKATHKQQHGLNQSVKDAHNTAPKRQKGVYEVVDVGYDHYQNDKTKSKQHFIKLKTLNKGVEKTYWGADF
GDLCSRESVKAGDLVRLKKLGQKEVKIPALDKNGVQHGWKTVHRNEWQLENLGVKGVDRTPSASKELVLN
SPDMLLKQQQRMAQFTQQKASTLQSEQKLKTGIKFWSL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A5J2A6K9 |
Host bacterium
| ID | 2388 | GenBank | NZ_VUIY01000028 |
| Plasmid name | 158900|unnamed2 | Incompatibility group | - |
| Plasmid size | 6652 bp | Coordinate of oriT [Strand] | 2697..3303 [-] |
| Host baterium | Salmonella enterica subsp. enterica serovar Java strain 158900 |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |