Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 101597 |
| Name | oriT_pCrieF1144_1 |
| Organism | Escherichia coli strain CrieF1144 |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_JBAGRT010000003 (180213..180466 [-], 254 nt) |
| oriT length | 254 nt |
| IRs (inverted repeats) | 152..157, 159..164 (AAAAGT..ACTTTT) 121..129, 134..142 (TTAAGGCTT..AAGCCTTAA) 50..55, 59..64 (AATTTT..AAAATT) |
| Location of nic site | 77..78 |
| Conserved sequence flanking the nic site |
TTTGGTTAAA |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 254 nt
>oriT_pCrieF1144_1
GGATTTAGGTTTTTTTTTAATCGCTTCACATTTCGTTAGCATGCGAAGAAATTTTGATAAAATTCTGGTCAGTTTGGTTAAAAAGTGTTACAAGTAAGCGGTGTGGTTGAAGGGATAGATTTAAGGCTTATTCAAGCCTTAAGAAAATACTAAAAGTTACTTTTCACCCTACCGAACACCTAACAAAAAATCCATGTTGAAGATTTGAACAATTGTAATGGCGCAAGGACAATCAGCACATGTCAGAATCTGAT
GGATTTAGGTTTTTTTTTAATCGCTTCACATTTCGTTAGCATGCGAAGAAATTTTGATAAAATTCTGGTCAGTTTGGTTAAAAAGTGTTACAAGTAAGCGGTGTGGTTGAAGGGATAGATTTAAGGCTTATTCAAGCCTTAAGAAAATACTAAAAGTTACTTTTCACCCTACCGAACACCTAACAAAAAATCCATGTTGAAGATTTGAACAATTGTAATGGCGCAAGGACAATCAGCACATGTCAGAATCTGAT
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file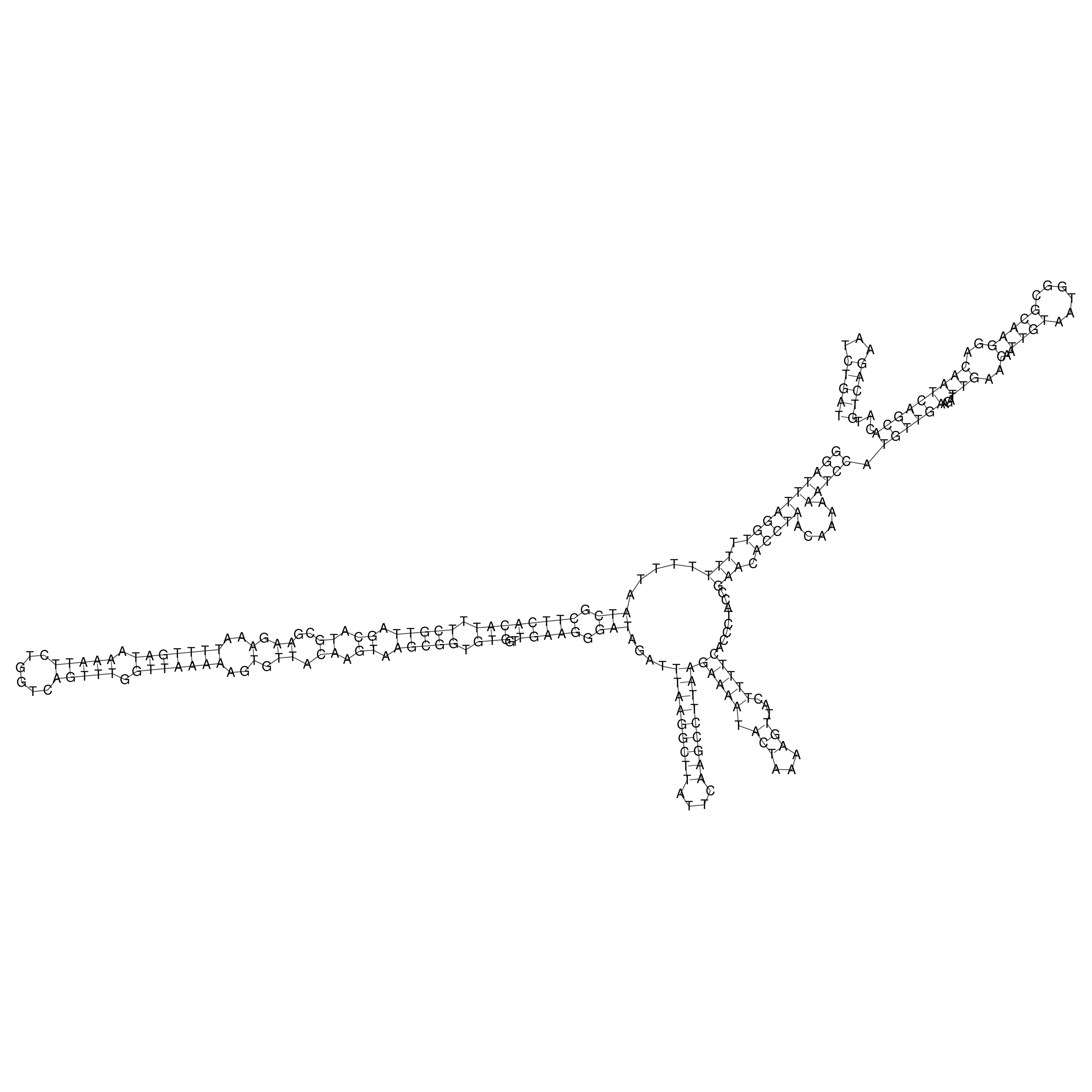
Relaxase
| ID | 1416 | GenBank | WP_049589872 |
| Name | mobH_V5G27_RS25095_pCrieF1144_1 |
UniProt ID | A0A5X7KNX7 |
| Length | 1048 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1048 a.a. Molecular weight: 117787.06 Da Isoelectric Point: 4.6054
>WP_049589872.1 MULTISPECIES: MobH family relaxase [Enterobacteriaceae]
MNFRALYLCIKRILGIFSSQENDATSVMIEDISSLSPFAQILGDQKYTVPDHPNPEVLKFIEYPTRPTGI
QTFNEQSILSLYREKLHSISMMLAISDSDIRDDAYTFTNLVLKPLVEYVRWIHLLPASENHHHNGIGGLL
SHSLEVAILSLKNAHHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLASPIIWTPS
SQSLLDWARENDVVEYEIHWRKRIHNQHNIWSSVFLERILNPVCLAFLDRVNKERVYSKMITALNVYTDG
NDFLSKCVRTADFYSTGTDLNVLRDPIMGLRSNDAAARAISTIKHNFTSININNYNAKPMHIIIVNGEVY
LNENAFLDFVLNDFELHKYNFPQGEAGKTVLVESLVQRGYVEPYDDERVVHYFIPGIYSENEISNIFRNG
IGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRDAITKITDTVE
TAVLKVNDLGRSTASIDVDIHSKKNEGSSDDFEKKAESDNEIDNDTQIVKSEGEEAADPVIPDIEESEDE
SAKDTESHVLVNQLHELLLSAPLSNDYIVCVDAVPYLNIDATMALLPGLDEKAFSEEPYFQLTFREGSLD
GMWIVRDIDDLRLVQLGDNCAGFQLTYHEPRRPTTLKSLFNTSMYQALVINDESSVENSAPRPKQTLELP
PPRVNALEEHSGDVEYHGTDSASATGPLKTEAVEYEHYQHLFEEEDEEHEIIDYTDFTQLSVSRPEVGSC
ATSSSVHNEKLLPEPSELPELNREQNADPQGTIERSMDVSVGQENSEPDTEGNCPPPAEVVYSQTEAAAT
SVMASEEPALPPVLEESNGEHAPTDAKGHHLSPALARLFAPTAPVEKQNPKRNRNKSSDKAEVQKPASPV
SGHNLNSKVFATTESDQNGEFSLISEGDVTELEFVEIASVLHQILSKMEVAFKRKRKNRFMVSTPNTLYL
TQSCVEKFGSQLEAQDLFNKLPQYLVNSGAVINTKCHAFNMPTLLAASDRAKVDIERIINNLKEAGNL
MNFRALYLCIKRILGIFSSQENDATSVMIEDISSLSPFAQILGDQKYTVPDHPNPEVLKFIEYPTRPTGI
QTFNEQSILSLYREKLHSISMMLAISDSDIRDDAYTFTNLVLKPLVEYVRWIHLLPASENHHHNGIGGLL
SHSLEVAILSLKNAHHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLASPIIWTPS
SQSLLDWARENDVVEYEIHWRKRIHNQHNIWSSVFLERILNPVCLAFLDRVNKERVYSKMITALNVYTDG
NDFLSKCVRTADFYSTGTDLNVLRDPIMGLRSNDAAARAISTIKHNFTSININNYNAKPMHIIIVNGEVY
LNENAFLDFVLNDFELHKYNFPQGEAGKTVLVESLVQRGYVEPYDDERVVHYFIPGIYSENEISNIFRNG
IGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRDAITKITDTVE
TAVLKVNDLGRSTASIDVDIHSKKNEGSSDDFEKKAESDNEIDNDTQIVKSEGEEAADPVIPDIEESEDE
SAKDTESHVLVNQLHELLLSAPLSNDYIVCVDAVPYLNIDATMALLPGLDEKAFSEEPYFQLTFREGSLD
GMWIVRDIDDLRLVQLGDNCAGFQLTYHEPRRPTTLKSLFNTSMYQALVINDESSVENSAPRPKQTLELP
PPRVNALEEHSGDVEYHGTDSASATGPLKTEAVEYEHYQHLFEEEDEEHEIIDYTDFTQLSVSRPEVGSC
ATSSSVHNEKLLPEPSELPELNREQNADPQGTIERSMDVSVGQENSEPDTEGNCPPPAEVVYSQTEAAAT
SVMASEEPALPPVLEESNGEHAPTDAKGHHLSPALARLFAPTAPVEKQNPKRNRNKSSDKAEVQKPASPV
SGHNLNSKVFATTESDQNGEFSLISEGDVTELEFVEIASVLHQILSKMEVAFKRKRKNRFMVSTPNTLYL
TQSCVEKFGSQLEAQDLFNKLPQYLVNSGAVINTKCHAFNMPTLLAASDRAKVDIERIINNLKEAGNL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A5X7KNX7 |
T4CP
| ID | 1072 | GenBank | WP_001284076 |
| Name | traD_V5G27_RS25100_pCrieF1144_1 |
UniProt ID | _ |
| Length | 694 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 694 a.a. Molecular weight: 78137.93 Da Isoelectric Point: 8.0177
>WP_001284076.1 MULTISPECIES: conjugative transfer system coupling protein TraD [Enterobacterales]
MSDSKRTNLHAQENFYRPILEYRSASILLICSVSMLYMGLSSDGLDIAPIVLFTSILLFLLCLYRCKTAA
PFLMAHWRVFKRHFMFVSLDSLRVINKSNFFSNERKYRQLVQDYQNKNKDIPERKSYFCDGFEWGPEHAD
RAYQIANLSSDKREIELPFVFNPIKRHFDAMARKMGGSNAIFAVERREPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVVVVIDPKNDAEWRESLMEEAKTLGLPFYKFHPGQPASSVCIDVCNTYTNVSD
LTSRLLSLVTVPGEVNPFVQYAKALVSNVISGLSYIEKKPSIYLIHKNMKSHMSIVNLTVKVMESCYARY
YGYDVWTEKVKYVANETLPVRFKRLAEWFTAHFMNYEGSEQIDWLDTVSQLIDYSMSDPEHMAKMTAGIM
PVFDMLIEKPLNELLSPNPNSVSSREIVTSEGMFSTGGVLYISLDGLSNPDTAAAISQLIMSDLTSCAGS
RYNAQDGDMSANSRISIFVDEAHSAINNPMINLLAQGRAAKIALFICTQTISDFIAAASVETANRITGLC
NNYISLRVNDTPTQTLVVENFGKSAISTNMVTYTTGSETSLPHNNFSGSISERKQTTLEESIPKDLLGQV
PMFHIVARLQDGRKVVGQIPIAVAEKQMKPNTTLSEMLFKKAGKVTLRQNLDIKNLNKFLRKLH
MSDSKRTNLHAQENFYRPILEYRSASILLICSVSMLYMGLSSDGLDIAPIVLFTSILLFLLCLYRCKTAA
PFLMAHWRVFKRHFMFVSLDSLRVINKSNFFSNERKYRQLVQDYQNKNKDIPERKSYFCDGFEWGPEHAD
RAYQIANLSSDKREIELPFVFNPIKRHFDAMARKMGGSNAIFAVERREPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVVVVIDPKNDAEWRESLMEEAKTLGLPFYKFHPGQPASSVCIDVCNTYTNVSD
LTSRLLSLVTVPGEVNPFVQYAKALVSNVISGLSYIEKKPSIYLIHKNMKSHMSIVNLTVKVMESCYARY
YGYDVWTEKVKYVANETLPVRFKRLAEWFTAHFMNYEGSEQIDWLDTVSQLIDYSMSDPEHMAKMTAGIM
PVFDMLIEKPLNELLSPNPNSVSSREIVTSEGMFSTGGVLYISLDGLSNPDTAAAISQLIMSDLTSCAGS
RYNAQDGDMSANSRISIFVDEAHSAINNPMINLLAQGRAAKIALFICTQTISDFIAAASVETANRITGLC
NNYISLRVNDTPTQTLVVENFGKSAISTNMVTYTTGSETSLPHNNFSGSISERKQTTLEESIPKDLLGQV
PMFHIVARLQDGRKVVGQIPIAVAEKQMKPNTTLSEMLFKKAGKVTLRQNLDIKNLNKFLRKLH
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 3779..12986
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| V5G27_RS24100 (V5G27_24100) | 1..876 | + | 876 | WP_000594612 | RepB family plasmid replication initiator protein | - |
| V5G27_RS24105 (V5G27_24105) | 2025..2378 | + | 354 | WP_000423602 | hypothetical protein | - |
| V5G27_RS24110 (V5G27_24110) | 2468..3580 | + | 1113 | WP_001300563 | IS4-like element IS421 family transposase | - |
| V5G27_RS24115 (V5G27_24115) | 3779..4096 | + | 318 | WP_000043357 | type IV conjugative transfer system protein TraL | traL |
| V5G27_RS24120 (V5G27_24120) | 4108..4896 | + | 789 | WP_000783153 | TraE/TraK family type IV conjugative transfer system protein | traE |
| V5G27_RS24125 (V5G27_24125) | 4896..6167 | + | 1272 | WP_000592090 | type-F conjugative transfer system secretin TraK | traK |
| V5G27_RS24130 (V5G27_24130) | 6169..6630 | + | 462 | WP_000521240 | plasmid transfer protein HtdO | - |
| V5G27_RS24135 (V5G27_24135) | 6620..7975 | + | 1356 | WP_000351841 | IncHI-type conjugal transfer protein TrhB | traB |
| V5G27_RS24140 (V5G27_24140) | 7983..8465 | + | 483 | WP_000377633 | hypothetical protein | - |
| V5G27_RS24145 (V5G27_24145) | 8584..9336 | + | 753 | WP_183079335 | protein-disulfide isomerase HtdT | - |
| V5G27_RS24150 (V5G27_24150) | 9346..10296 | + | 951 | WP_001022587 | IncHI-type conjugal transfer lipoprotein TrhV | traV |
| V5G27_RS24155 (V5G27_24155) | 10305..12986 | + | 2682 | WP_000387412 | TraC family protein | virb4 |
Host bacterium
| ID | 2041 | GenBank | NZ_JBAGRT010000003 |
| Plasmid name | pCrieF1144_1 | Incompatibility group | IncHI2A |
| Plasmid size | 251464 bp | Coordinate of oriT [Strand] | 180213..180466 [-] |
| Host baterium | Escherichia coli strain CrieF1144 |
Cargo genes
| Drug resistance gene | blaTEM-1B, aadA2, sul3, ant(3'')-Ia, floR, sul1, qacE, lnu(F), tet(A), mcr-1.1, mph(A), dfrA14 |
| Virulence gene | - |
| Metal resistance gene | terE, terD, terC, terB, terA, terZ, terW |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |