Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 101270 |
| Name | oriT_F2W|unnamed1 |
| Organism | Enterobacter asburiae strain F2W |
| Sequence Completeness | - |
| NCBI accession of oriT (coordinates [strand]) | NZ_JACGGB010000019 (19305..19558 [-], 254 nt) |
| oriT length | 254 nt |
| IRs (inverted repeats) | 152..157, 159..164 (AAAAGT..ACTTTT) 121..129, 134..142 (TTAAGGCTT..AAGCCTTAA) 50..55, 59..64 (AATTTT..AAAATT) |
| Location of nic site | 77..78 |
| Conserved sequence flanking the nic site |
TTTGGTTAAA |
| Note | Predicted by oriTfinder 2.0 |
oriT sequence
Download Length: 254 nt
>oriT_F2W|unnamed1
GGATTTAGGTTTTTTTTTAATCGCTTCACATTTCGTTAGCATGCGAAGAAATTTTGATAAAATTCTGGTCAGTTTGGTTAAAAAGTGTTACAAGTAAGCGGTGTGGTTGAAGGGATAGATTTAAGGCTTATTCAAGCCTTAAGAAAATACTAAAAGTTACTTTTCACCCTACCGAACACCTAACAAAAAATCCATGTTGAAGATTTGAACAATTGTAATGGCGCAAGGACAATCAGCACATGTCAGAATCTGAT
GGATTTAGGTTTTTTTTTAATCGCTTCACATTTCGTTAGCATGCGAAGAAATTTTGATAAAATTCTGGTCAGTTTGGTTAAAAAGTGTTACAAGTAAGCGGTGTGGTTGAAGGGATAGATTTAAGGCTTATTCAAGCCTTAAGAAAATACTAAAAGTTACTTTTCACCCTACCGAACACCTAACAAAAAATCCATGTTGAAGATTTGAACAATTGTAATGGCGCAAGGACAATCAGCACATGTCAGAATCTGAT
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file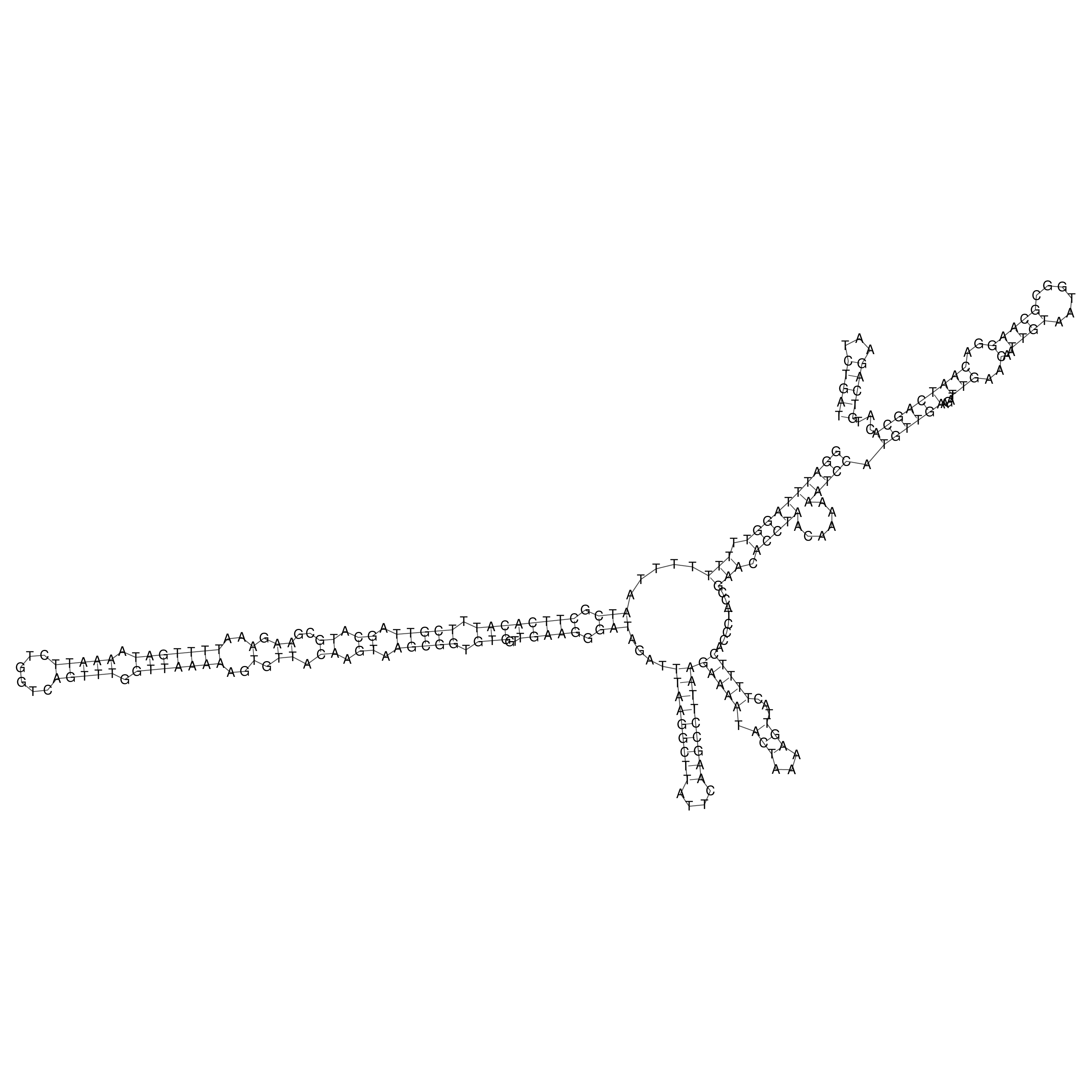
Relaxase
| ID | 1227 | GenBank | WP_001011151 |
| Name | mobH_H4F75_RS20365_F2W|unnamed1 |
UniProt ID | A0A1P8VU63 |
| Length | 1048 a.a. | PDB ID | |
| Note | Predicted by oriTfinder 2.0 | ||
Relaxase protein sequence
Download Length: 1048 a.a. Molecular weight: 117773.04 Da Isoelectric Point: 4.6054
>WP_001011151.1 MULTISPECIES: MobH family relaxase [Enterobacterales]
MNFRALYLCIKRILGIFSSQENDATSVMIEDISSLSPFAQILGDQKYTVPDHPNPEVLKFIEYPTRPTGI
QTFNEQSILSLYREKLHSISMMLAISDSDIRDDAYTFTNLVLKPLVEYVRWIHLLPASENHHHNGIGGLL
SHSLEVAILSLKNAHHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLASPIIWTPS
SQSLLDWARENDVVEYEIHWRKRIHNQHNIWSSVFLERILNPVCLAFLDRVNKERVYSKMITALNVYTDG
NDFLSKCVRTADFYSTGTDLNVLRDPIMGLRSNDAAARAISTIKHNFTSININNYNAKPMHIIIVNGEVY
LNENAFLDFVLNDFELHKYNFPQGEAGKTVLVESLVQRGYVEPYDDERVVHYFIPGIYSENEISNIFRNG
IGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRDAITKITDTVE
TAVLKVNDLGRSTASIDVDIHSKKNEGSSDDFEKKAESDNEIDNDTQIVKSEGEEAADPVIPDIEESEDE
SAKDTESHVLVNQLHELLLSAPLSNDYIVCVDAVPYLNIDATMALLPGLDEKAFSEEPYFQLTFREGSLD
GMWIVRDIDDLRLVQLGDNCAGFQLTYHEPRRPTTLKSLFNTSMYQALVINDESSVENSAPRPKQTLELP
PPRVNAVEEHSGDVEYHGTDSASATGPLKTEAVEYEHYQHLFEEEDEEHEIIDYTDFTQLSVSRPEVGSC
ATSSSVHNEKLLPEPSELPELNREQNADPQGTIERSMDVSVGQENSEPDTEGNCPPPAEVVYSQTEAAAT
SVMASEEPALPPVLEESNGEHAPTDAKGHHLSPALARLFAPTAPVEKQNPKRNRNKSSDKAEVQKPASPV
SGHNLNSKVFATTESDQNGEFSLISEGDVTELEFVEIASVLHQILSKMEVAFKRKRKNRFMVSTPNTLYL
TQSCVEKFGSQLEAQDLFNKLPQYLVNSGAVINTKCHAFNMPTLLAASDRAKVDIERIINNLKEAGNL
MNFRALYLCIKRILGIFSSQENDATSVMIEDISSLSPFAQILGDQKYTVPDHPNPEVLKFIEYPTRPTGI
QTFNEQSILSLYREKLHSISMMLAISDSDIRDDAYTFTNLVLKPLVEYVRWIHLLPASENHHHNGIGGLL
SHSLEVAILSLKNAHHSELRPIGYQDEEVVRRKVYLYAAFICGLVHDAGKVYDLDIVSLNLASPIIWTPS
SQSLLDWARENDVVEYEIHWRKRIHNQHNIWSSVFLERILNPVCLAFLDRVNKERVYSKMITALNVYTDG
NDFLSKCVRTADFYSTGTDLNVLRDPIMGLRSNDAAARAISTIKHNFTSININNYNAKPMHIIIVNGEVY
LNENAFLDFVLNDFELHKYNFPQGEAGKTVLVESLVQRGYVEPYDDERVVHYFIPGIYSENEISNIFRNG
IGKLEFYNLLKLRWIGLIFDSYKIPDSVPGLFSVNANKDFIYIDEQKTVTEYRRPVPGRDAITKITDTVE
TAVLKVNDLGRSTASIDVDIHSKKNEGSSDDFEKKAESDNEIDNDTQIVKSEGEEAADPVIPDIEESEDE
SAKDTESHVLVNQLHELLLSAPLSNDYIVCVDAVPYLNIDATMALLPGLDEKAFSEEPYFQLTFREGSLD
GMWIVRDIDDLRLVQLGDNCAGFQLTYHEPRRPTTLKSLFNTSMYQALVINDESSVENSAPRPKQTLELP
PPRVNAVEEHSGDVEYHGTDSASATGPLKTEAVEYEHYQHLFEEEDEEHEIIDYTDFTQLSVSRPEVGSC
ATSSSVHNEKLLPEPSELPELNREQNADPQGTIERSMDVSVGQENSEPDTEGNCPPPAEVVYSQTEAAAT
SVMASEEPALPPVLEESNGEHAPTDAKGHHLSPALARLFAPTAPVEKQNPKRNRNKSSDKAEVQKPASPV
SGHNLNSKVFATTESDQNGEFSLISEGDVTELEFVEIASVLHQILSKMEVAFKRKRKNRFMVSTPNTLYL
TQSCVEKFGSQLEAQDLFNKLPQYLVNSGAVINTKCHAFNMPTLLAASDRAKVDIERIINNLKEAGNL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | A0A1P8VU63 |
T4CP
| ID | 837 | GenBank | WP_001284076 |
| Name | traD_H4F75_RS20370_F2W|unnamed1 |
UniProt ID | _ |
| Length | 694 a.a. | PDB ID | _ |
| Note | Predicted by oriTfinder 2.0 | ||
T4CP protein sequence
Download Length: 694 a.a. Molecular weight: 78137.93 Da Isoelectric Point: 8.0177
>WP_001284076.1 MULTISPECIES: conjugative transfer system coupling protein TraD [Enterobacterales]
MSDSKRTNLHAQENFYRPILEYRSASILLICSVSMLYMGLSSDGLDIAPIVLFTSILLFLLCLYRCKTAA
PFLMAHWRVFKRHFMFVSLDSLRVINKSNFFSNERKYRQLVQDYQNKNKDIPERKSYFCDGFEWGPEHAD
RAYQIANLSSDKREIELPFVFNPIKRHFDAMARKMGGSNAIFAVERREPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVVVVIDPKNDAEWRESLMEEAKTLGLPFYKFHPGQPASSVCIDVCNTYTNVSD
LTSRLLSLVTVPGEVNPFVQYAKALVSNVISGLSYIEKKPSIYLIHKNMKSHMSIVNLTVKVMESCYARY
YGYDVWTEKVKYVANETLPVRFKRLAEWFTAHFMNYEGSEQIDWLDTVSQLIDYSMSDPEHMAKMTAGIM
PVFDMLIEKPLNELLSPNPNSVSSREIVTSEGMFSTGGVLYISLDGLSNPDTAAAISQLIMSDLTSCAGS
RYNAQDGDMSANSRISIFVDEAHSAINNPMINLLAQGRAAKIALFICTQTISDFIAAASVETANRITGLC
NNYISLRVNDTPTQTLVVENFGKSAISTNMVTYTTGSETSLPHNNFSGSISERKQTTLEESIPKDLLGQV
PMFHIVARLQDGRKVVGQIPIAVAEKQMKPNTTLSEMLFKKAGKVTLRQNLDIKNLNKFLRKLH
MSDSKRTNLHAQENFYRPILEYRSASILLICSVSMLYMGLSSDGLDIAPIVLFTSILLFLLCLYRCKTAA
PFLMAHWRVFKRHFMFVSLDSLRVINKSNFFSNERKYRQLVQDYQNKNKDIPERKSYFCDGFEWGPEHAD
RAYQIANLSSDKREIELPFVFNPIKRHFDAMARKMGGSNAIFAVERREPIFVTEDNWFGHTLITGNVGTG
KTVLQRLLSISMLHLGHVVVVIDPKNDAEWRESLMEEAKTLGLPFYKFHPGQPASSVCIDVCNTYTNVSD
LTSRLLSLVTVPGEVNPFVQYAKALVSNVISGLSYIEKKPSIYLIHKNMKSHMSIVNLTVKVMESCYARY
YGYDVWTEKVKYVANETLPVRFKRLAEWFTAHFMNYEGSEQIDWLDTVSQLIDYSMSDPEHMAKMTAGIM
PVFDMLIEKPLNELLSPNPNSVSSREIVTSEGMFSTGGVLYISLDGLSNPDTAAAISQLIMSDLTSCAGS
RYNAQDGDMSANSRISIFVDEAHSAINNPMINLLAQGRAAKIALFICTQTISDFIAAASVETANRITGLC
NNYISLRVNDTPTQTLVVENFGKSAISTNMVTYTTGSETSLPHNNFSGSISERKQTTLEESIPKDLLGQV
PMFHIVARLQDGRKVVGQIPIAVAEKQMKPNTTLSEMLFKKAGKVTLRQNLDIKNLNKFLRKLH
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
Host bacterium
| ID | 1714 | GenBank | NZ_JACGGB010000019 |
| Plasmid name | F2W|unnamed1 | Incompatibility group | IncHI2 |
| Plasmid size | 81567 bp | Coordinate of oriT [Strand] | 19305..19558 [-] |
| Host baterium | Enterobacter asburiae strain F2W |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |