Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 100827 |
| Name | oriT_pCE-R2-11-0435_92 |
| Organism | Salmonella enterica subsp. enterica serovar Heidelberg strain CE-R2-11-0435 |
| Sequence Completeness | intact |
| NCBI accession of oriT (coordinates [strand]) | NZ_CP016520 (38923..39032 [+], 110 nt) |
| oriT length | 110 nt |
| IRs (inverted repeats) | IR1: 24..31, 35..42 (GTCGGGGC..GCCCTGAC) IR2: 49..65, 72..88 (GTAATTGTAATAGCGTC..GACGGTATTACAATTAC) |
| Location of nic site | 96..97 |
| Conserved sequence flanking the nic site |
CATCCTG|T |
| Note | predicted by the oriTfinder |
oriT sequence
Download Length: 110 nt
>oriT_pCE-R2-11-0435_92
TCACTTCAGGCTCCTTACGGGGTGTCGGGGCGAAGCCCTGACCAGATGGTAATTGTAATAGCGTCGCGTGTGACGGTATTACAATTACACATCCTGTCCCGTTTTTCAGG
TCACTTCAGGCTCCTTACGGGGTGTCGGGGCGAAGCCCTGACCAGATGGTAATTGTAATAGCGTCGCGTGTGACGGTATTACAATTACACATCCTGTCCCGTTTTTCAGG
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file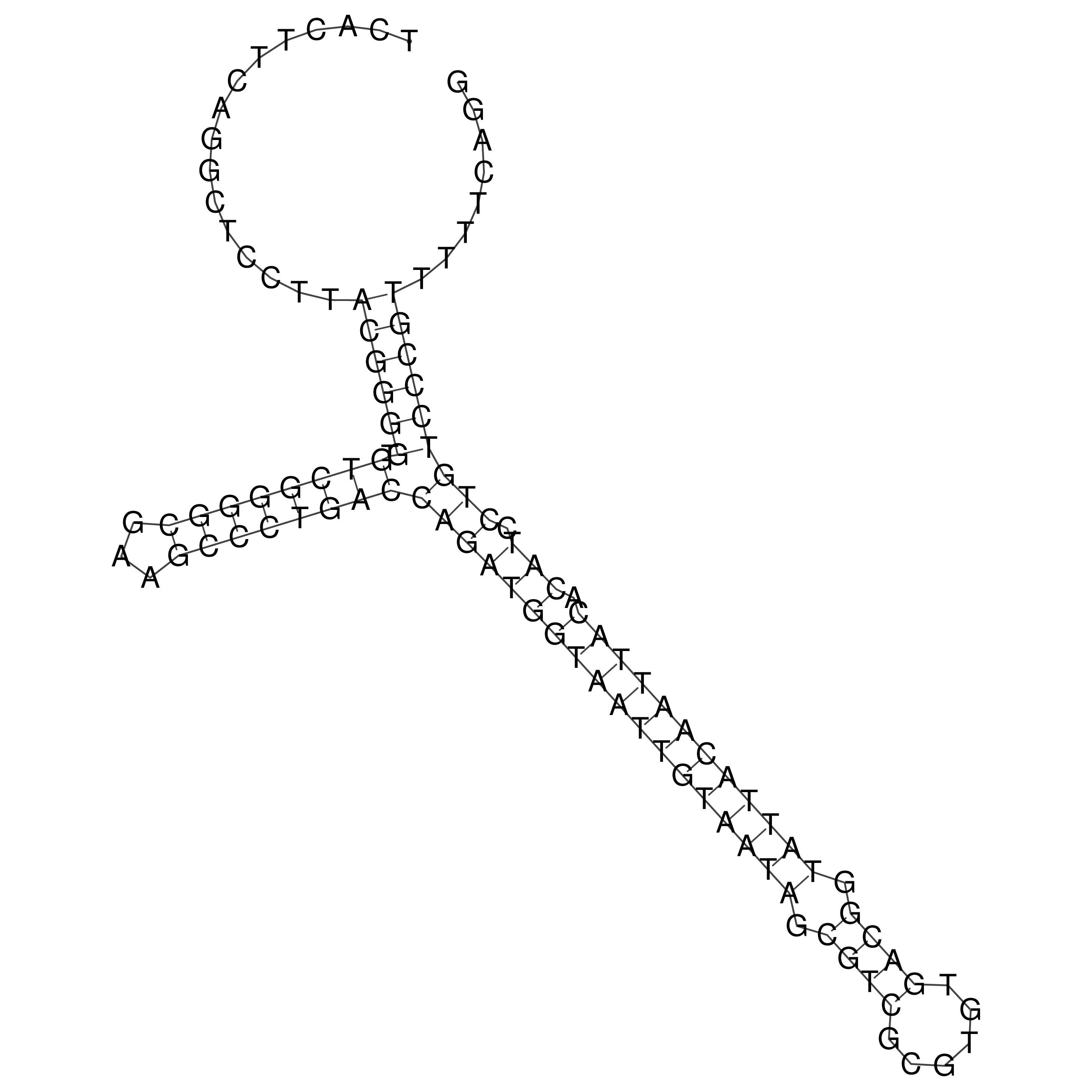
Relaxase
| ID | 899 | GenBank | WP_001405892 |
| Name | NikB_pCE-R2-11-0435_92 |
UniProt ID | _ |
| Length | 899 a.a. | PDB ID | |
| Note | putative relaxase | ||
Relaxase protein sequence
Download Length: 899 a.a. Molecular weight: 103883.32 Da Isoelectric Point: 7.3373
>WP_001405892.1 MULTISPECIES: IncI1-type relaxase NikB [Enterobacteriaceae]
MNAVIPKKRRDGKSSFEDLVSYVSVRDDMTDEELDLSSSSQAEQPHRSRFSRLVDYATRLRNESFVALVD
VMKDGCEWVNFYGVTCFHNCTSLETAAADMEYIAQQAHYAKDNTDPVFHYILSWQAHESPRPEQIYDSVR
HTLKSLGLGEHQYVSAVHTDTDNLHVHVAVNRVHPVTGYLNCLSWSQEKLSRACRELELKHGFAPDNGCW
VHAPGNRIVRKTAVERDRQNAWTRGKKQTFREYVAQTAVAGLRSEPVNDWLSLHRRLAEDGLYLSQMDGK
FLVMDGWDRNREGVQLDSFGPSWCAEKLMKKMGDYTPVPKDIFSQVEAPGRYNPDFIAADVRPEKIAETE
SLQQYACRHLGERLPEMAREGRLENCQAIHRTLAEAGLWMRVQHGHLVICDGYDHNQTPVRADSVWSLLT
LDNVNQLDGGWQPVPTDIFRQVTPTERFRGRRMESCPATDKEWHRMRTGTGPQGAIKRELFSDKESLWGY
SISHCSPQIEEMITQGEFTWQRCHELFAQQGLMLQKQHHGLVVVDAFNHEQTPVKASSIHPDLTLGRAEP
QAGPFVSAPADLFDRVQPESRYNPELAVSDRYGVSSKRDPMLRRQRREARAEARADLRARYLAWREQWRK
PDLRYGERCREIHQACRLRKSHIRAQYDDPALRKLHYHIAEVQRMQALIRLKEDIRDERQKLIADGKWYP
PSYRQWVEIQAAQGDRAAVSQLRGWDYRDRRKDRSRTTTTDRCVVLCEPGGTPVYGNTGDLEARLQKNGS
VRFRDRRTGEFVCTDYGDRVVFRNHHDRNALADKLDLIAPVLFGRDPRMGFEPEGNDKQFNQVFAEMVAW
HNVTGRTGHEDYRITRPDVDHHREGSERYYRDYIAANSNDDASLPPPEQDKRWEPPSPG
MNAVIPKKRRDGKSSFEDLVSYVSVRDDMTDEELDLSSSSQAEQPHRSRFSRLVDYATRLRNESFVALVD
VMKDGCEWVNFYGVTCFHNCTSLETAAADMEYIAQQAHYAKDNTDPVFHYILSWQAHESPRPEQIYDSVR
HTLKSLGLGEHQYVSAVHTDTDNLHVHVAVNRVHPVTGYLNCLSWSQEKLSRACRELELKHGFAPDNGCW
VHAPGNRIVRKTAVERDRQNAWTRGKKQTFREYVAQTAVAGLRSEPVNDWLSLHRRLAEDGLYLSQMDGK
FLVMDGWDRNREGVQLDSFGPSWCAEKLMKKMGDYTPVPKDIFSQVEAPGRYNPDFIAADVRPEKIAETE
SLQQYACRHLGERLPEMAREGRLENCQAIHRTLAEAGLWMRVQHGHLVICDGYDHNQTPVRADSVWSLLT
LDNVNQLDGGWQPVPTDIFRQVTPTERFRGRRMESCPATDKEWHRMRTGTGPQGAIKRELFSDKESLWGY
SISHCSPQIEEMITQGEFTWQRCHELFAQQGLMLQKQHHGLVVVDAFNHEQTPVKASSIHPDLTLGRAEP
QAGPFVSAPADLFDRVQPESRYNPELAVSDRYGVSSKRDPMLRRQRREARAEARADLRARYLAWREQWRK
PDLRYGERCREIHQACRLRKSHIRAQYDDPALRKLHYHIAEVQRMQALIRLKEDIRDERQKLIADGKWYP
PSYRQWVEIQAAQGDRAAVSQLRGWDYRDRRKDRSRTTTTDRCVVLCEPGGTPVYGNTGDLEARLQKNGS
VRFRDRRTGEFVCTDYGDRVVFRNHHDRNALADKLDLIAPVLFGRDPRMGFEPEGNDKQFNQVFAEMVAW
HNVTGRTGHEDYRITRPDVDHHREGSERYYRDYIAANSNDDASLPPPEQDKRWEPPSPG
Protein domains
Predicted by InterproScan.
Protein structure
No available structure.
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 44392..82680
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| BAT85_RS23425 (BAT85_23190) | 42108..44399 | - | 2292 | WP_001289282 | F-type conjugative transfer protein TrbC | - |
| BAT85_RS23430 (BAT85_23195) | 44392..45462 | - | 1071 | WP_000151582 | IncI1-type conjugal transfer protein TrbB | trbB |
| BAT85_RS23435 (BAT85_23200) | 45481..46689 | - | 1209 | WP_000121273 | IncI1-type conjugal transfer protein TrbA | trbA |
| BAT85_RS23440 (BAT85_23205) | 46981..47133 | + | 153 | WP_001303307 | Hok/Gef family protein | - |
| BAT85_RS23445 (BAT85_23210) | 47205..47456 | - | 252 | WP_001291965 | hypothetical protein | - |
| BAT85_RS26505 | 47955..48050 | + | 96 | WP_001303310 | DinQ-like type I toxin DqlB | - |
| BAT85_RS26175 | 48115..48291 | - | 177 | WP_001054897 | hypothetical protein | - |
| BAT85_RS23450 (BAT85_23215) | 48683..48892 | + | 210 | WP_000062603 | HEAT repeat domain-containing protein | - |
| BAT85_RS23455 (BAT85_23220) | 48964..49614 | - | 651 | WP_001178506 | plasmid IncI1-type surface exclusion protein ExcA | - |
| BAT85_RS23460 (BAT85_23225) | 49688..51856 | - | 2169 | WP_000698357 | DotA/TraY family protein | traY |
| BAT85_RS23465 (BAT85_23230) | 51953..52537 | - | 585 | WP_001037987 | IncI1-type conjugal transfer protein TraX | - |
| BAT85_RS23470 (BAT85_23235) | 52566..53768 | - | 1203 | WP_001189160 | IncI1-type conjugal transfer protein TraW | traW |
| BAT85_RS25200 | 53735..54349 | - | 615 | WP_000337399 | IncI1-type conjugal transfer protein TraV | traV |
| BAT85_RS23475 (BAT85_23240) | 54349..57393 | - | 3045 | WP_065641822 | IncI1-type conjugal transfer protein TraU | traU |
| BAT85_RS23480 (BAT85_23245) | 57483..58283 | - | 801 | WP_001164788 | IncI1-type conjugal transfer protein TraT | traT |
| BAT85_RS23485 (BAT85_23250) | 58267..58455 | - | 189 | WP_001277255 | putative conjugal transfer protein TraS | - |
| BAT85_RS23490 (BAT85_23255) | 58519..58923 | - | 405 | WP_000086959 | IncI1-type conjugal transfer protein TraR | traR |
| BAT85_RS23495 (BAT85_23260) | 58974..59501 | - | 528 | WP_001055569 | conjugal transfer protein TraQ | traQ |
| BAT85_RS23500 (BAT85_23265) | 59501..60205 | - | 705 | WP_000801919 | IncI1-type conjugal transfer protein TraP | traP |
| BAT85_RS23505 (BAT85_23270) | 60205..61494 | - | 1290 | WP_001272000 | conjugal transfer protein TraO | traO |
| BAT85_RS23510 (BAT85_23275) | 61497..62480 | - | 984 | WP_001191879 | IncI1-type conjugal transfer protein TraN | traN |
| BAT85_RS23515 (BAT85_23280) | 62491..63183 | - | 693 | WP_000138551 | DotI/IcmL family type IV secretion protein | traM |
| BAT85_RS23520 (BAT85_23285) | 63180..63527 | - | 348 | WP_001055900 | conjugal transfer protein | traL |
| BAT85_RS23525 (BAT85_23290) | 63545..67312 | - | 3768 | WP_001141542 | LPD7 domain-containing protein | - |
| BAT85_RS23530 (BAT85_23295) | 67402..67953 | - | 552 | WP_000014584 | phospholipase D family protein | - |
| BAT85_RS25205 | 67968..68258 | - | 291 | WP_001372180 | hypothetical protein | traK |
| BAT85_RS23535 (BAT85_23300) | 68255..69403 | - | 1149 | WP_001024972 | plasmid transfer ATPase TraJ | virB11 |
| BAT85_RS23540 (BAT85_23305) | 69400..70218 | - | 819 | WP_000646097 | IncI1-type conjugal transfer lipoprotein TraI | traI |
| BAT85_RS23545 (BAT85_23310) | 70215..70673 | - | 459 | WP_001079808 | IncI1-type conjugal transfer lipoprotein TraH | - |
| BAT85_RS23550 (BAT85_23315) | 71068..71652 | - | 585 | WP_000977522 | histidine phosphatase family protein | - |
| BAT85_RS23555 (BAT85_23320) | 71712..72914 | - | 1203 | WP_000976353 | conjugal transfer protein TraF | - |
| BAT85_RS23560 (BAT85_23325) | 73000..73824 | - | 825 | WP_001545755 | conjugal transfer protein TraE | traE |
| BAT85_RS23565 (BAT85_23330) | 73975..75129 | - | 1155 | WP_001139958 | site-specific integrase | - |
| BAT85_RS26415 | 75128..75481 | + | 354 | WP_001393368 | hypothetical protein | - |
| BAT85_RS26040 | 76650..76898 | + | 249 | WP_001349157 | hypothetical protein | - |
| BAT85_RS23570 (BAT85_23335) | 76895..78187 | - | 1293 | WP_001619585 | shufflon system plasmid conjugative transfer pilus tip adhesin PilV | - |
| BAT85_RS23575 (BAT85_23340) | 78187..78843 | - | 657 | WP_001193549 | A24 family peptidase | - |
| BAT85_RS23580 (BAT85_23345) | 78828..79388 | - | 561 | WP_000014005 | lytic transglycosylase domain-containing protein | virB1 |
| BAT85_RS23585 (BAT85_23350) | 79398..80012 | - | 615 | WP_000908226 | type 4 pilus major pilin | - |
| BAT85_RS23590 (BAT85_23355) | 80029..81114 | - | 1086 | WP_001208802 | type II secretion system F family protein | - |
| BAT85_RS23595 (BAT85_23360) | 81127..82680 | - | 1554 | WP_000362206 | ATPase, T2SS/T4P/T4SS family | virB11 |
| BAT85_RS23600 (BAT85_23365) | 82691..83143 | - | 453 | WP_001247337 | type IV pilus biogenesis protein PilP | - |
| BAT85_RS23605 (BAT85_23370) | 83130..84425 | - | 1296 | WP_000752780 | type 4b pilus protein PilO2 | - |
| BAT85_RS23610 (BAT85_23375) | 84418..86100 | - | 1683 | WP_000748143 | PilN family type IVB pilus formation outer membrane protein | - |
| BAT85_RS23615 (BAT85_23380) | 86114..86551 | - | 438 | WP_000539806 | type IV pilus biogenesis protein PilM | - |
| BAT85_RS25245 | 86551..87618 | - | 1068 | WP_001360287 | type IV pilus biogenesis lipoprotein PilL | - |
Host bacterium
| ID | 1287 | GenBank | NZ_CP016520 |
| Plasmid name | pCE-R2-11-0435_92 | Incompatibility group | IncI1 |
| Plasmid size | 92323 bp | Coordinate of oriT [Strand] | 38923..39032 [+] |
| Host baterium | Salmonella enterica subsp. enterica serovar Heidelberg strain CE-R2-11-0435 |
Cargo genes
| Drug resistance gene | - |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |