Detailed information of oriT
oriT
The information of the oriT region
| oriTDB ID | 100423 |
| Name | oriT_plasmid1_DO |
| Organism | Enterococcus faecium DO |
| Sequence Completeness | intact |
| NCBI accession of oriT (coordinates [strand]) | NC_017961 (23823..24037 [+], 215 nt) |
| oriT length | 215 nt |
| IRs (inverted repeats) | 61..76, 80..95 (ACTTAACCCCCCGTAT..ACAGGGGGGTACAAAT) 123..134, 139..150 (GAAAATCCTTTG..CAAGGGATTTAC) |
| Location of nic site | 135..136 |
| Conserved sequence flanking the nic site |
TGG|T |
| Note | predicted by the oriTfinder |
oriT sequence
Download Length: 215 nt
>oriT_plasmid1_DO
AAGCGGAAGTCGCAGGTGTGGACTGATCTTGCTGGCTGGTGTGGCAATAGCCACGCCAGCACTTAACCCCCCGTATCTAACAGGGGGGTACAAATCGACAGGAAACAGTCAAAAAAACATTAGAAAATCCTTTGGTTACAAGGGATTTACAAAATTTCAGCGTATGTCAAATGGGCTTTAAAAGTTGACATACGCCTTTTTGATTGGAGGGATTT
AAGCGGAAGTCGCAGGTGTGGACTGATCTTGCTGGCTGGTGTGGCAATAGCCACGCCAGCACTTAACCCCCCGTATCTAACAGGGGGGTACAAATCGACAGGAAACAGTCAAAAAAACATTAGAAAATCCTTTGGTTACAAGGGATTTACAAAATTTCAGCGTATGTCAAATGGGCTTTAAAAGTTGACATACGCCTTTTTGATTGGAGGGATTT
Visualization of oriT structure
oriT secondary structure
Predicted by RNAfold.
Download structure file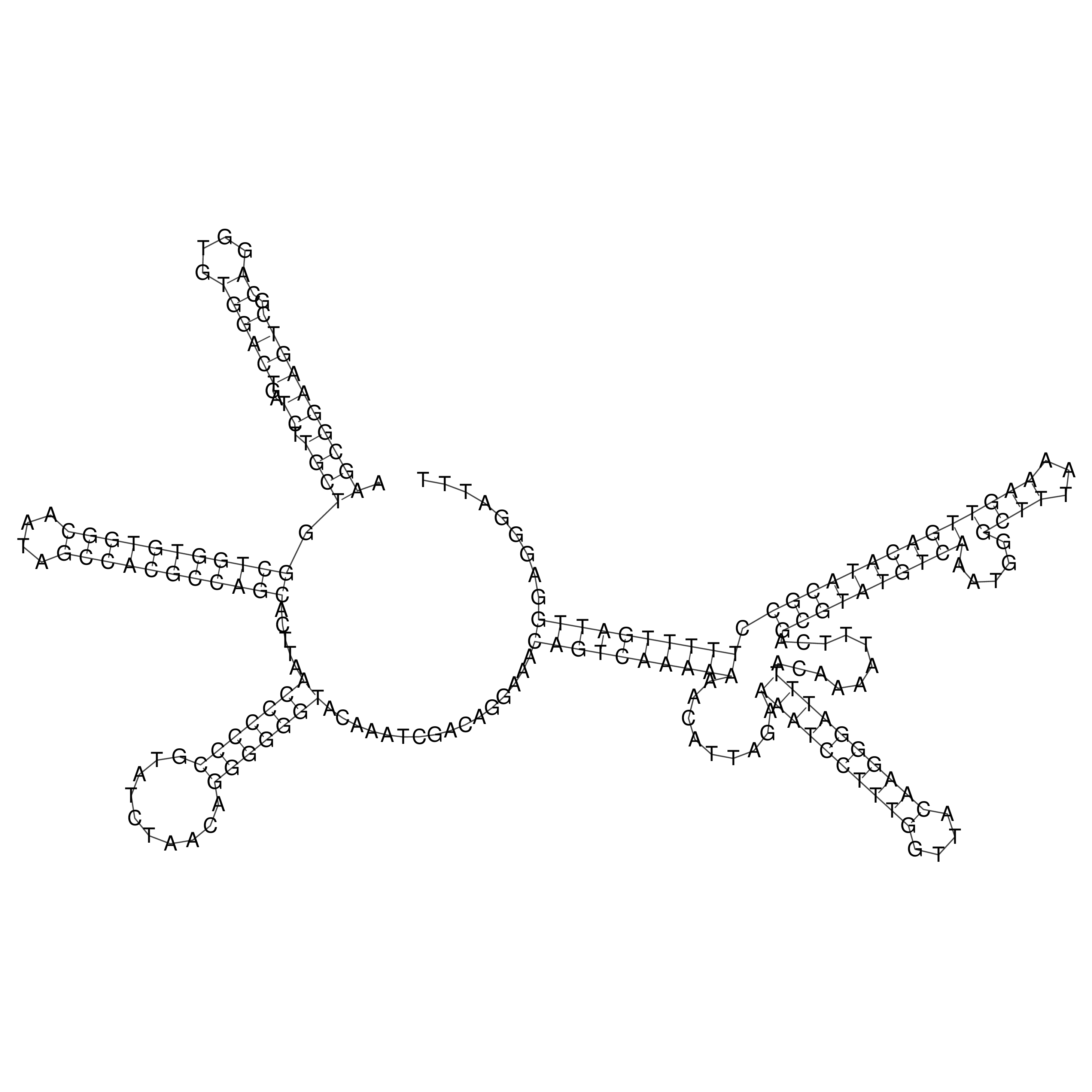
Relaxase
| ID | 494 | GenBank | YP_006377333 |
| Name | Pre_plasmid1_DO |
UniProt ID | Q3Y252 |
| Length | 420 a.a. | PDB ID | |
| Note | putative relaxase | ||
Relaxase protein sequence
Download Length: 420 a.a. Molecular weight: 49660.33 Da Isoelectric Point: 9.9693
>YP_006377333.1 plasmid recombinase enzyme Pre (plasmid) [Enterococcus faecium DO]
MSYAVCRMQKVKSAGLKGMQFHNQRERKSRTNDDIDHERTRENYDLKNDKNIDYNERVKEIIESQKTGTR
KTRKDAVLVNELLVTSDRDFFEQLDPGEQKRFFEESYKLFSERYGKQNIAYATVHNDEQTPHMHLGVVPM
RDGKLQGKNVFNRQELLWLQDKFPEHMKKQGFELKRGERGSDRKHIETAKFKKQTLEKEIDFLEKNLAVK
KDEWTAYSDKVKSDLEVPAKRHMKSVEVPTGEKSMFGLGKEIMKTEKKPTKNVVISERDYKNLVTAARDN
DRLKQHVRNLMSTDMAREYKKLSKEHGQVKEKYSGLVERFNENVNDYNELLEENKSLKSKISDLKRDVSL
IYESTKEFLKERTDGLKAFKNVFKGFVDKVKDKTAQFQEKHDLEPKKNEFELTHNREVKKERSRDQGMSL
MSYAVCRMQKVKSAGLKGMQFHNQRERKSRTNDDIDHERTRENYDLKNDKNIDYNERVKEIIESQKTGTR
KTRKDAVLVNELLVTSDRDFFEQLDPGEQKRFFEESYKLFSERYGKQNIAYATVHNDEQTPHMHLGVVPM
RDGKLQGKNVFNRQELLWLQDKFPEHMKKQGFELKRGERGSDRKHIETAKFKKQTLEKEIDFLEKNLAVK
KDEWTAYSDKVKSDLEVPAKRHMKSVEVPTGEKSMFGLGKEIMKTEKKPTKNVVISERDYKNLVTAARDN
DRLKQHVRNLMSTDMAREYKKLSKEHGQVKEKYSGLVERFNENVNDYNELLEENKSLKSKISDLKRDVSL
IYESTKEFLKERTDGLKAFKNVFKGFVDKVKDKTAQFQEKHDLEPKKNEFELTHNREVKKERSRDQGMSL
Protein domains
Predicted by InterproScan.
Protein structure
| Source | ID | Structure |
|---|---|---|
| AlphaFold DB | Q3Y252 |
T4SS
T4SS were predicted by using oriTfinder2.
Region 1: 21718..35886
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| HMPREF0351_RS13140 (HMPREF0351_12728) | 18022..18921 | + | 900 | WP_025192414 | protein rep | - |
| HMPREF0351_RS13145 (HMPREF0351_12729) | 18944..19624 | - | 681 | WP_002328852 | IS6-like element IS1216 family transposase | - |
| HMPREF0351_RS13150 (HMPREF0351_12730) | 19782..20404 | - | 623 | Protein_19 | recombinase family protein | - |
| HMPREF0351_RS13155 (HMPREF0351_12731) | 20675..21187 | - | 513 | WP_000774078 | hypothetical protein | - |
| HMPREF0351_RS13160 (HMPREF0351_12734) | 21718..22032 | + | 315 | WP_000420682 | YdcP family protein | orf23 |
| HMPREF0351_RS13165 (HMPREF0351_12735) | 22048..22434 | + | 387 | WP_000985015 | YdcP family protein | orf23 |
| HMPREF0351_RS13170 (HMPREF0351_12736) | 22463..23848 | + | 1386 | WP_000813488 | FtsK/SpoIIIE domain-containing protein | virb4 |
| HMPREF0351_RS13175 (HMPREF0351_12737) | 23851..24003 | + | 153 | WP_000879507 | hypothetical protein | - |
| HMPREF0351_RS13180 (HMPREF0351_12738) | 24026..25231 | + | 1206 | WP_000398284 | MobT family relaxase | - |
| HMPREF0351_RS13185 (HMPREF0351_12739) | 26015..27925 | + | 1911 | WP_002286940 | group II intron reverse transcriptase/maturase | - |
| HMPREF0351_RS13190 (HMPREF0351_12740) | 28029..28253 | + | 225 | WP_032509114 | hypothetical protein | orf19 |
| HMPREF0351_RS13195 (HMPREF0351_12741) | 28370..28867 | + | 498 | WP_000342539 | antirestriction protein ArdA | - |
| HMPREF0351_RS13200 (HMPREF0351_12742) | 28956..29348 | + | 393 | WP_000723888 | conjugal transfer protein | orf17a |
| HMPREF0351_RS13205 (HMPREF0351_12743) | 29332..31779 | + | 2448 | WP_000331160 | ATP-binding protein | virb4 |
| HMPREF0351_RS13210 (HMPREF0351_12744) | 31782..33959 | + | 2178 | WP_014748747 | YtxH domain-containing protein | orf15 |
| HMPREF0351_RS13215 (HMPREF0351_12745) | 33956..34957 | + | 1002 | WP_001574272 | bifunctional lysozyme/C40 family peptidase | orf14 |
| HMPREF0351_RS13220 (HMPREF0351_12746) | 34954..35886 | + | 933 | WP_001224319 | conjugal transfer protein | orf13 |
| HMPREF0351_RS13225 | 36131..36247 | + | 117 | WP_001791010 | tetracycline resistance determinant leader peptide | - |
Host bacterium
| ID | 884 | GenBank | NC_017961 |
| Plasmid name | plasmid1_DO | Incompatibility group | _ |
| Plasmid size | 36262 bp | Coordinate of oriT [Strand] | 23823..24037 [+] |
| Host baterium | Enterococcus faecium DO |
Cargo genes
| Drug resistance gene | tetracycline resistance |
| Virulence gene | - |
| Metal resistance gene | - |
| Degradation gene | - |
| Symbiosis gene | - |
| Anti-CRISPR | - |