Detailed information of TA system
Overview
TA module
| Type | I | Classification (family/domain) | txpA-ratA/- |
| Location | 2905894..2906117 | Replicon | chromosome |
| Accession | NZ_AP026721 | ||
| Organism | Enterococcus faecalis strain SVR2330 | ||
Toxin (Protein)
| Gene name | txpA | Uniprot ID | - |
| Locus tag | PO855_RS14395 | Protein ID | WP_075551663.1 |
| Coordinates | 2906016..2906117 (-) | Length | 34 a.a. |
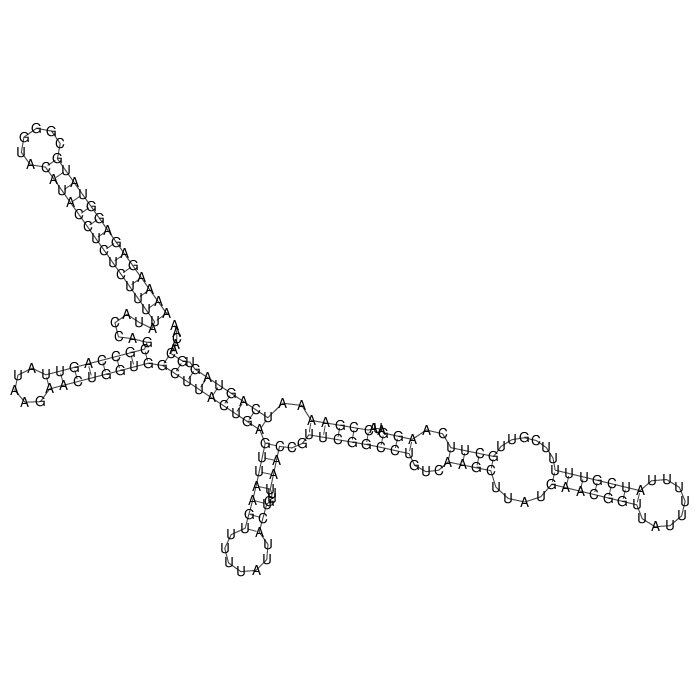
Antitoxin (RNA)
| Gene name | ratA | ||
| Locus tag | - | ||
| Coordinates | 2905894..2906071 (+) |
Genomic Context
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|
Associated MGEs
| MGE detail |
Similar MGEs |
Relative position |
MGE Type | Cargo ARG | Virulence gene | Coordinates | Length (bp) |
|---|
Relative position:
(1) inside: TA loci is completely located inside the MGE;
(2) overlap: TA loci is partially overlapped with the MGE;
(3) flank: The TA loci is located in the 5 kb flanking regions of MGE.
Sequences
Toxin
Download Length: 34 a.a. Molecular weight: 3599.34 Da Isoelectric Point: 4.5869
>T42619 WP_075551663.1 NZ_AP026721:c2906117-2906016 [Enterococcus faecalis]
MSIEAALELMISFAAFVALLIFGILEATKNDKK
MSIEAALELMISFAAFVALLIFGILEATKNDKK
Download Length: 102 bp
>T42619 NZ_AP026721:c2906117-2906016 [Enterococcus faecalis]
TTGTCAATCGAAGCGGCATTGGAATTGATGATTAGTTTTGCAGCGTTTGTTGCACTACTGATTTTCGGTATCCTTGAAGC
AACGAAAAACGATAAAAAATAA
TTGTCAATCGAAGCGGCATTGGAATTGATGATTAGTTTTGCAGCGTTTGTTGCACTACTGATTTTCGGTATCCTTGAAGC
AACGAAAAACGATAAAAAATAA
Antitoxin
Download Length: 178 bp
>AT42619 NZ_AP026721:2905894-2906071 [Enterococcus faecalis]
AAAAGAGAGGTATGCGGGTACATACCTCTCTTTTATACCAGCGCCAGTTATAAGAACTGGTGGCTTACTGAGTTAAGTTT
TTATTACTGTTTAACCGTTCGGCCTGTCAAGCTTATGAACGGTTATTTTTTATCGTTTTTCGTTGCTTCAAGGATACCGA
AAATCAGTAGTGCAACAA
AAAAGAGAGGTATGCGGGTACATACCTCTCTTTTATACCAGCGCCAGTTATAAGAACTGGTGGCTTACTGAGTTAAGTTT
TTATTACTGTTTAACCGTTCGGCCTGTCAAGCTTATGAACGGTTATTTTTTATCGTTTTTCGTTGCTTCAAGGATACCGA
AAATCAGTAGTGCAACAA
Similar Proteins
Only experimentally validated proteins are listed.
| Protein | Organism | Identities (%) | Coverage (%) | Ha-value |
|---|
| Protein | Organism | Identities (%) | Coverage (%) | Ha-value |
|---|