Detailed information of TA system
Overview
TA module
| Type | I | Classification (family/domain) | sprA1-sprA1AS/- |
| Location | 565228..565408 | Replicon | chromosome |
| Accession | NZ_CP089157 | ||
| Organism | Staphylococcus aureus strain UNC_SaCF30 | ||
Toxin (Protein)
| Gene name | sprA1 | Uniprot ID | - |
| Locus tag | LTK17_RS02660 | Protein ID | WP_001801861.1 |
| Coordinates | 565228..565323 (+) | Length | 32 a.a. |
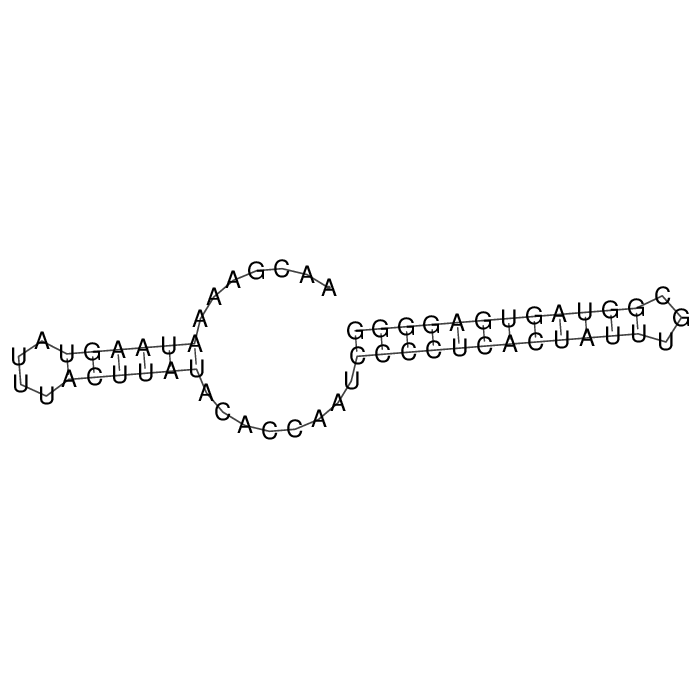
Antitoxin (RNA)
| Gene name | sprA1AS | ||
| Locus tag | - | ||
| Coordinates | 565351..565408 (-) |
Genomic Context
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|
Associated MGEs
| MGE detail |
Similar MGEs |
Relative position |
MGE Type | Cargo ARG | Virulence gene | Coordinates | Length (bp) |
|---|
Relative position:
(1) inside: TA loci is completely located inside the MGE;
(2) overlap: TA loci is partially overlapped with the MGE;
(3) flank: The TA loci is located in the 5 kb flanking regions of MGE.
Sequences
Toxin
Download Length: 32 a.a. Molecular weight: 3554.32 Da Isoelectric Point: 10.4449
>T227729 WP_001801861.1 NZ_CP089157:565228-565323 [Staphylococcus aureus]
VMLIFVHIIAPVISGCAIAFFSYWLSRRNTK
VMLIFVHIIAPVISGCAIAFFSYWLSRRNTK
Download Length: 96 bp
>T227729 NZ_CP123024:2719962-2720069 [Escherichia coli]
ATGACGCTCGCGCAGTTTGCCATGACTTTCTGGCACGACCTGGCGGCACCGATCCTGGCGGGAATTATTACCGCAGCGAT
TGTCGGCTGGTGGCGTAACCGGAAGTAA
ATGACGCTCGCGCAGTTTGCCATGACTTTCTGGCACGACCTGGCGGCACCGATCCTGGCGGGAATTATTACCGCAGCGAT
TGTCGGCTGGTGGCGTAACCGGAAGTAA
Antitoxin
Download Length: 58 bp
>AT227729 NZ_CP089157:c565408-565351 [Staphylococcus aureus]
AACGAAAATAAGTATTTACTTATACACCAATCCCCTCACTATTTGCGGTAGTGAGGGG
AACGAAAATAAGTATTTACTTATACACCAATCCCCTCACTATTTGCGGTAGTGAGGGG
Similar Proteins
Only experimentally validated proteins are listed.
| Protein | Organism | Identities (%) | Coverage (%) | Ha-value |
|---|
| Protein | Organism | Identities (%) | Coverage (%) | Ha-value |
|---|