Detailed information of TA system
Overview
TA module
| Type | I | Classification (family/domain) | sprA1-sprA1AS/- |
| Location | 216271..216451 | Replicon | chromosome |
| Accession | NZ_CP062401 | ||
| Organism | Staphylococcus aureus strain NAS_OP_098 | ||
Toxin (Protein)
| Gene name | sprA1 | Uniprot ID | - |
| Locus tag | IF770_RS01420 | Protein ID | WP_001801861.1 |
| Coordinates | 216356..216451 (-) | Length | 32 a.a. |
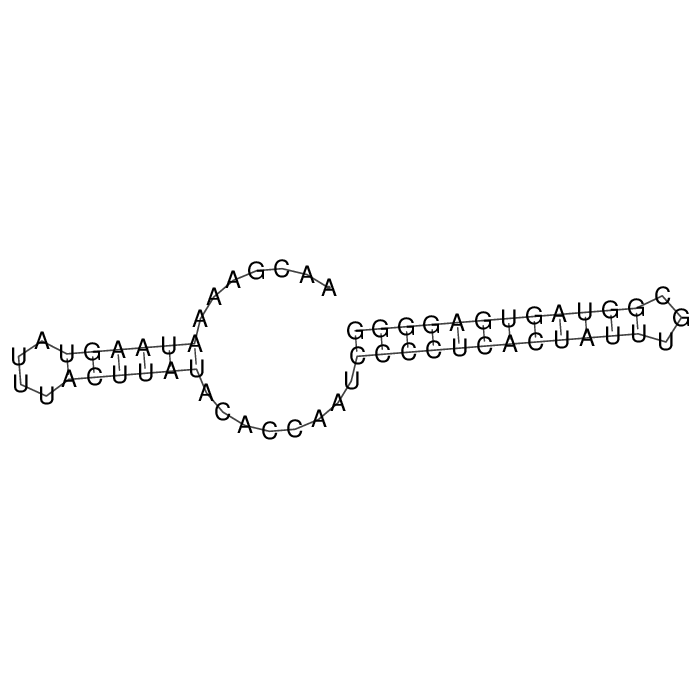
Antitoxin (RNA)
| Gene name | sprA1AS | ||
| Locus tag | - | ||
| Coordinates | 216271..216328 (+) |
Genomic Context
| Locus tag | Coordinates | Strand | Size (bp) | Protein ID | Product | Description |
|---|---|---|---|---|---|---|
| IF770_RS01390 | 211980..212114 | + | 135 | WP_001791797.1 | hypothetical protein | - |
| IF770_RS01395 | 212278..213834 | + | 1557 | WP_000028669.1 | type I restriction-modification system subunit M | - |
| IF770_RS01400 | 213827..215056 | + | 1230 | WP_000072627.1 | restriction endonuclease subunit S | - |
| IF770_RS01405 | 215598..216008 | + | 411 | WP_223200549.1 | transposase | - |
| IF770_RS01410 | 215993..216154 | + | 162 | Protein_243 | transposase | - |
| IF770_RS01415 | 216132..216233 | - | 102 | WP_001792025.1 | hypothetical protein | - |
| - | 216271..216328 | + | 58 | - | - | Antitoxin |
| IF770_RS01420 | 216356..216451 | - | 96 | WP_001801861.1 | type I toxin-antitoxin system Fst family toxin PepA1 | Toxin |
| IF770_RS01425 | 216596..217608 | + | 1013 | Protein_246 | IS3 family transposase | - |
| IF770_RS01430 | 217806..218378 | - | 573 | WP_000414216.1 | hypothetical protein | - |
| IF770_RS01435 | 218479..218820 | - | 342 | WP_000627540.1 | DUF3969 family protein | - |
| IF770_RS01440 | 218861..219487 | - | 627 | WP_000669024.1 | hypothetical protein | - |
| IF770_RS01445 | 219562..220557 | - | 996 | WP_000070642.1 | DUF4352 domain-containing protein | - |
| IF770_RS01450 | 220638..221288 | - | 651 | WP_001795210.1 | excalibur calcium-binding domain-containing protein | - |
Associated MGEs
| MGE detail |
Similar MGEs |
Relative position |
MGE Type | Cargo ARG | Virulence gene | Coordinates | Length (bp) |
|---|---|---|---|---|---|---|---|
| - | inside | Prophage | - | selk / hlgA / lukD | 141725..240470 | 98745 |
Relative position:
(1) inside: TA loci is completely located inside the MGE;
(2) overlap: TA loci is partially overlapped with the MGE;
(3) flank: The TA loci is located in the 5 kb flanking regions of MGE.
Sequences
Toxin
Download Length: 32 a.a. Molecular weight: 3554.32 Da Isoelectric Point: 10.4449
>T174891 WP_001801861.1 NZ_CP062401:c216451-216356 [Staphylococcus aureus]
VMLIFVHIIAPVISGCAIAFFSYWLSRRNTK
VMLIFVHIIAPVISGCAIAFFSYWLSRRNTK
Download Length: 96 bp
>T174891 NZ_CP081701:2803454-2803561 [Escherichia coli]
ATGACGCTCGCGCAGTTTGCCATGATTTTCTGGCACGACCTGGCAGCACCGATCCTGGCGGGAATTATTACCGCAGCGAT
TGTCGGCTGGTGGCGTAACCGGAAGTAA
ATGACGCTCGCGCAGTTTGCCATGATTTTCTGGCACGACCTGGCAGCACCGATCCTGGCGGGAATTATTACCGCAGCGAT
TGTCGGCTGGTGGCGTAACCGGAAGTAA
Antitoxin
Download Length: 58 bp
>AT174891 NZ_CP062401:216271-216328 [Staphylococcus aureus]
AACGAAAATAAGTATTTACTTATACACCAATCCCCTCACTATTTGCGGTAGTGAGGGG
AACGAAAATAAGTATTTACTTATACACCAATCCCCTCACTATTTGCGGTAGTGAGGGG
Similar Proteins
Only experimentally validated proteins are listed.
| Protein | Organism | Identities (%) | Coverage (%) | Ha-value |
|---|
| Protein | Organism | Identities (%) | Coverage (%) | Ha-value |
|---|