
| 1043 | |
| ICEValE0601 | |
| Vibrio alginolyticus E0601 | |
| 106165 | |
| 46.04 | |
| prfC | |
| - | |
| - | |
| KT072768 (complete ICE sequence in this GenBank file) | |
| - | |
| 1..106165 | |
| coordinates: 5724..5830; oriTDB id: 200056 TATCGAGACGCCAAACAGTGATTGTTACGGCAGTTTTACGTTTGGCGTTTCGATCCAAAAGCCAAACGGA TAGTGGTTTTGGCTTTCGGGGTTAATTGGATGGGGAA | |
| coordinates: 38216..40396; Locus tag: ICEValE0601_046; Family: MOBH |
The graph information of ICEValE0601 components from KT072768 | |||||
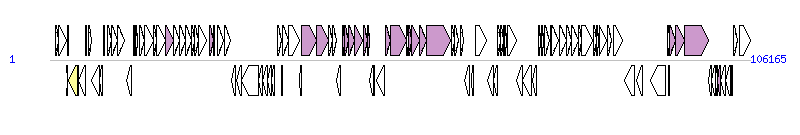 | |||||
| Complete gene list of ICEValE0601 from KT072768 | |||||
| # | Gene | Coordinates [+/-], size (bp) | Product (GenBank annotation) | *Reannotation | |
| 1 | hipB | 906..1226 [+], 321 | putative Antitoxin protein,HipB | ||
| 2 | hipA | 1219..2433 [+], 1215 | putative toxin protein, HipA | ||
| 3 | ICEValE0601_003 | 2583..2696 [-], 114 | Hypothetical protein | ||
| 4 | xis | 2659..2853 [+], 195 | Recombination directionality factor, Xis | ||
| 5 | int | 2865..4106 [-], 1242 | Intergrase | Integrase | |
| 6 | s002 | 4108..4377 [-], 270 | Hypothetical protein, S002 | ||
| 7 | s003 | 4380..5354 [-], 975 | Rod shape determination protein,S003 | ||
| 8 | ICEValE0601_008 | 5450..5572 [+], 123 | Hypothetical protein | ||
| 9 | mobI | 5819..6262 [+], 444 | Hypothetical protein, MobI | ||
| 10 | rumB | 6273..7541 [-], 1269 | Error-prone repair protein, RumB | ||
| 11 | rumA | 7549..7998 [-], 450 | Error-prone repair protein, RumA | ||
| 12 | ICEValE0601_012 | 8118..8255 [+], 138 | Hypothetical protein | ||
| 13 | s024 | 8654..9313 [+], 660 | DNA polymerase III, S024 | ||
| 14 | ICEValE0601_014 | 9357..10274 [+], 918 | Transposase | ||
| 15 | ICEValE0601_015 | 10312..11352 [+], 1041 | DDE endonuclease | ||
| 16 | ICEValE0601_016 | 11552..12373 [-], 822 | Transposase | ||
| 17 | ICEValE0601_017 | 12652..12867 [+], 216 | Flp pilus assembly protein | ||
| 18 | cpaA | 13025..13297 [+], 273 | Type IV prepilin peptidase, CpaA | ||
| 19 | cpaB | 13517..14323 [+], 807 | Flp pilus assembly protein, CpaB | ||
| 20 | cpaC | 14335..15663 [+], 1329 | Type II/IV secretion system protein, CpaC | ||
| 21 | ICEValE0601_021 | 15660..16190 [+], 531 | Hypothetical protein | ||
| 22 | cpaE | 16199..17449 [+], 1251 | Type II/IV secretion system ATPase, CpaE | ||
| 23 | cpaF | 17446..18735 [+], 1290 | Type II/IV secretion system protein, CpaF | TrbB, T4SS component | |
| 24 | tadB | 18732..19652 [+], 921 | Flp pilus assembly protein, TadB | ||
| 25 | tadC | 19642..20550 [+], 909 | Type II/IV secretion system protein, TadC | ||
| 26 | tadD | 20543..21439 [+], 897 | Flp pilus assembly protein, TadD | ||
| 27 | tadE | 21420..21938 [+], 519 | Flp pilus assembly membrane protein, TadE | ||
| 28 | tadF | 21919..22458 [+], 540 | Flp pilus assembly surface protein, TadF | ||
| 29 | tadG | 22461..23789 [+], 1329 | Flp pilus assembly protein, TadG | ||
| 30 | ICEValE0601_030 | 24157..24798 [+], 642 | Outer membrane protein | IcmN, T4SS component | |
| 31 | ICEValE0601_031 | 24846..24995 [+], 150 | Hypothetical protein | ||
| 32 | ICEValE0601_032 | 25384..26322 [+], 939 | Transposase | ||
| 33 | ICEValE0601_033 | 26457..27377 [+], 921 | Transposase | ||
| 34 | ICEValE0601_034 | 27453..28127 [-], 675 | Two-component system response regulator | ||
| 35 | ICEValE0601_035 | 28237..29115 [-], 879 | Hypothetical protein | ||
| 36 | papC | 29105..31567 [-], 2463 | P pilus assembly protein, porin, PapC | ||
| 37 | ICEValE0601_037 | 31569..32252 [-], 684 | Hypothetical protein | ||
| 38 | papD | 32249..32980 [-], 732 | P pilus assembly protein, PapD | ||
| 39 | ICEValE0601_039 | 33046..33540 [-], 495 | Hypothetical protein | ||
| 40 | ICEValE0601_040 | 33626..34120 [-], 495 | Hypothetical protein | ||
| 41 | ICEValE0601_041 | 34430..35065 [+], 636 | Threonine efflux protein | ||
| 42 | ICEValE0601_042 | 35075..35197 [-], 123 | Hypothetical protein | ||
| 43 | ICEValE0601_043 | 35306..36118 [+], 813 | 4-hydroxyphenylpyruvate dioxygenase | ||
| 44 | ICEValE0601_044 | 36115..37824 [+], 1710 | Type III restriction endonuclease | ||
| 45 | ICEValE0601_045 | 37860..38162 [-], 303 | Hypothetical protein | ||
| 46 | traI | 38216..40396 [+], 2181 | Conjugative transfer protein, TraI | Relaxase, MOBH Family | |
| 47 | traD | 40445..42265 [+], 1821 | Conjugative transfer protein, TraD | TraD_F, T4SS component | |
| 48 | s091 | 42275..42835 [+], 561 | Conjugative transfer protein, S091 | ||
| 49 | traJ | 42822..43457 [+], 636 | Conjugative transfer protein, TraJ | ||
| 50 | ICEValE0601_050 | 43484..44071 [-], 588 | Hypothetical protein | ||
| 51 | traL | 44360..44641 [+], 282 | Conjugative transfer pilus assembly protein, TraL | TraL_F, T4SS component | |
| 52 | traE | 44650..45264 [+], 615 | Conjugative transfer pilus assembly protein, TraE | TraE_F, T4SS component | |
| 53 | traK | 45248..46144 [+], 897 | Conjugative transfer pilus assembly protein, TraK | TraK_F, T4SS component | |
| 54 | traB | 46147..47436 [+], 1290 | Conjugative transfer pilus assembly protein, TraB | TraB_F, T4SS component | |
| 55 | traV | 47511..48083 [+], 573 | Conjugative transfer protein, TraV | TraV_F, T4SS component | |
| 56 | traA | 48080..48466 [+], 387 | Conjugative transfer protein, TraA | TraA_F, T4SS component | |
| 57 | ICEValE0601_057 | 48507..49013 [-], 507 | Acetyltransferase | ||
| 58 | ICEValE0601_058 | 49004..49270 [-], 267 | Hypothetical protein | ||
| 59 | ICEValE0601_059 | 49581..50684 [-], 1104 | Transposase | ||
| 60 | dsbC | 50955..51647 [+], 693 | Thiol:disulfide involved in conjugative transfer,DsbC | TrbB_I, T4SS component | |
| 61 | traC | 51648..54047 [+], 2400 | Conjugative transfer pilus assembly protein, TraC | TraC_F, T4SS component | |
| 62 | ICEValE0601_062 | 54040..54387 [+], 348 | Conjugative transfer protein 345 | ||
| 63 | trhF | 54440..54883 [+], 444 | Conjugative signal peptidase, TrhF | TraF, T4SS component | |
| 64 | traW | 54894..56018 [+], 1125 | Conjugative transfer pilus assembly protein, TraW | TraW_F, T4SS component | |
| 65 | traU | 56050..57030 [+], 981 | Conjugative transfer pilus assembly protein, TraU | TraU_F, T4SS component | |
| 66 | traN | 57033..60725 [+], 3693 | Conjugative transfer protein, TraN | TraN_F, T4SS component | |
| 67 | ICEValE0601_067 | 60956..61288 [+], 333 | Hypothetical protein | ||
| 68 | ICEValE0601_068 | 61319..62008 [+], 690 | Hypothetical protein | ||
| 69 | ICEValE0601_069 | 62324..62752 [+], 429 | Transposase | ||
| 70 | ICEValE0601_070 | 62798..63697 [-], 900 | Transposase | ||
| 71 | ICEValE0601_071 | 63929..64231 [-], 303 | Hypothetical protein | ||
| 72 | betT | 64540..66183 [+], 1644 | High-affinity choline uptake protein, BetT | ||
| 73 | ICEValE0601_073 | 66379..67236 [-], 858 | Mechanosensitive channel-related protein | ||
| 74 | yrbG | 67314..67796 [-], 483 | Inner membrane protein, YrbG | ||
| 75 | ICEValE0601_075 | 67888..68370 [+], 483 | Transposase | ||
| 76 | ICEValE0601_076 | 68387..68776 [+], 390 | Transposase | ||
| 77 | ICEValE0601_077 | 68757..68882 [+], 126 | Hypothetical protein | ||
| 78 | ICEValE0601_078 | 68980..69189 [+], 210 | Hypothetical protein | ||
| 79 | ICEValE0601_079 | 69390..70694 [+], 1305 | RNA-dependent DNA polymerase | ||
| 80 | ICEValE0601_080 | 70727..71605 [-], 879 | DDE endonuclease | ||
| 81 | ICEValE0601_081 | 71709..73040 [-], 1332 | Transposase | ||
| 82 | ICEValE0601_082 | 73168..73770 [-], 603 | Hypothetical protein, S063 | ||
| 83 | s089 | 74138..74464 [+], 327 | Hypothetical protein,S089 | ||
| 84 | ssb | 74480..74899 [+], 420 | Single-stranded DNA-binding protein,Ssb | ||
| 85 | bet | 74979..75797 [+], 819 | Recombination protein, Bet | ||
| 86 | orfZ | 75879..76022 [+], 144 | Hypothetical protein, OrfZ | ||
| 87 | exo | 76084..77100 [+], 1017 | Recombination related exonuclease, Exo | ||
| 88 | cobS | 77310..78269 [+], 960 | Aerobic cobaltochelatase, CobS | ||
| 89 | ICEValE0601_089 | 78269..79036 [+], 768 | Hypothetical protein | ||
| 90 | ICEValE0601_090 | 79135..80088 [+], 954 | Cobalamine biosynthesis protein | ||
| 91 | ICEValE0601_091 | 80150..80590 [+], 441 | Hypothetical protein | ||
| 92 | ICEValE0601_092 | 80660..82315 [+], 1656 | Plasmid associated protein | ||
| 93 | radC | 82398..82895 [+], 498 | DNA repair protein, RadC | ||
| 94 | ICEValE0601_094 | 82895..83236 [+], 342 | Hypothetical protein | ||
| 95 | ICEValE0601_095 | 83328..84401 [+], 1074 | Putative primase | ||
| 96 | ICEValE0601_096 | 84490..85197 [+], 708 | Hypothetical protein | ||
| 97 | ICEValE0601_097 | 85515..86771 [+], 1257 | Glucose-1-phosphate adenylyltransferase | ||
| 98 | ICEValE0601_098 | 87126..88598 [-], 1473 | DNA polymerase | ||
| 99 | ICEValE0601_099 | 88886..89887 [-], 1002 | Transposase | ||
| 100 | ICEValE0601_100 | 91090..93288 [-], 2199 | Diguanylate cyclase | ||
| 101 | ICEValE0601_101 | 93591..93713 [+], 123 | Hypothetical protein | ||
| 102 | ICEValE0601_102 | 93844..93969 [-], 126 | Hypothetical protein | ||
| 103 | traF | 93953..94885 [+], 933 | Conjugative transfer pilus assembly protein, TraF | TraF_F, T4SS component | |
| 104 | traH | 94888..96276 [+], 1389 | Conjugative transfer pilus assembly protein, TraH | TraH_F, T4SS component | |
| 105 | traG | 96280..99849 [+], 3570 | Conjugative transfer protein, TraG | TraG_F, T4SS component | |
| 106 | eex | 99882..100313 [-], 432 | Exclusion system protein, Eex | ||
| 107 | setC | 100368..100901 [-], 534 | Transcriptional activator, SetC | ||
| 108 | setD | 100898..101197 [-], 300 | Transcriptional activator, SetD | ||
| 109 | ICEValE0601_109 | 101194..101742 [-], 549 | LysM/invasin protein | Orf169_F, T4SS component | |
| 110 | s083 | 101729..102391 [-], 663 | Hypothetical protein, S083 | ||
| 111 | s084 | 102378..103247 [-], 870 | Hypothetical protein, S084 | ||
| 112 | setQ | 103303..103554 [-], 252 | Hypothetical protein, SetQ | ||
| 113 | setR | 103672..104319 [+], 648 | cI prophage repressor protein, SetR | ||
| 114 | prfC | 104576..106165 [+], 1590 | Peptide chain release factor 3 | ||
| (1) Luo P; He X; Wang Y; Liu Q; Hu C (2016). Comparative genomic analysis of six new-found integrative conjugative elements (ICEs) in Vibrio alginolyticus. BMC Microbiol. 0.721527778. [PubMed:27145747] |