
| 1022 | |
| ICEPmiCHN2416 | |
| Proteus mirabilis 09MAS2416 | |
| 92,556 | |
| 46.71 | |
| - | |
| Resistance to ampicillin, azithromycin, chloramphenicol, cefazolin, ciprofloxacin, gentamycin, kanamycin, STRtrimethoprim-sulfamethoxazole, sulfamethoxazole, tetracycline | |
| - | |
| KX243407 (complete ICE sequence in this GenBank file) | |
| - | |
| 1..92556 | |
| coordinates: 3686..3792; oriTDB id: 200056 TATCGAGACGCCAAACAGTGATTGTTACGGCAGTTTTACGTTTGGCGTTTCGATCCAAAAGCCAAACGGA TAGTGGTTTTGGCTTTTGGGGTTAATTGGATGGGGAA | |
| coordinates: 34802..36952; Gene: traI; Family: MOBH |
The graph information of ICEPmiCHN2416 components from KX243407 | |||||
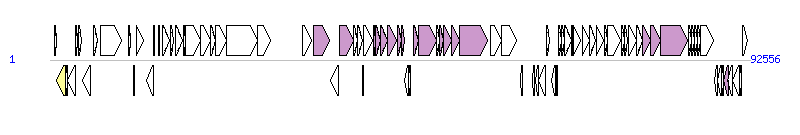 | |||||
| Complete gene list of ICEPmiCHN2416 from KX243407 | |||||
| # | Gene | Coordinates [+/-], size (bp) | Product (GenBank annotation) | *Reannotation | |
| 1 | - | 616..810 [+], 195 | hypothetical protein | ||
| 2 | int | 827..2068 [-], 1242 | integrase | Integrase | |
| 3 | - | 2070..2339 [-], 270 | hypothetical protein | ||
| 4 | - | 2342..3316 [-], 975 | Rod shape determination protein | ||
| 5 | - | 3412..3579 [+], 168 | hypothetical protein | ||
| 6 | - | 3781..4224 [+], 444 | hypothetical protein | ||
| 7 | rumB | 4235..5407 [-], 1173 | Error-prone, lesion bypass DNA polymerase V (UmuC) | ||
| 8 | tnp | 5691..6284 [+], 594 | Transposase | ||
| 9 | tnpA | 6638..9478 [+], 2841 | Transposase | ||
| 10 | - | 10376..10642 [+], 267 | hypothetical protein | ||
| 11 | - | 11037..11162 [-], 126 | hypothetical protein | ||
| 12 | - | 11441..12424 [+], 984 | hypothetical protein | ||
| 13 | - | 12785..13648 [-], 864 | Beta-lactamase | ||
| 14 | - | 13679..13810 [+], 132 | hypothetical protein | ||
| 15 | - | 14377..14499 [+], 123 | hypothetical protein | ||
| 16 | - | 14882..15790 [+], 909 | DNA polymerase III | ||
| 17 | - | 15990..16289 [+], 300 | hypothetical protein | ||
| 18 | - | 16607..17530 [+], 924 | hypothetical protein | ||
| 19 | - | 17679..17906 [+], 228 | transcriptional regulator | ||
| 20 | - | 17906..19960 [+], 2055 | Type I restriction-modification system, DNA-methyltransferase subunit M | ||
| 21 | - | 19957..21252 [+], 1296 | conserved hypothetical protein | ||
| 22 | - | 21255..21917 [+], 663 | hypothetical protein | ||
| 23 | - | 21914..23323 [+], 1410 | type I restriction-modification system, S subunit, putative | ||
| 24 | - | 23333..27466 [+], 4134 | protein kinase, putative | ||
| 25 | - | 27463..29109 [+], 1647 | hypothetical protein | ||
| 26 | herA | 33403..34638 [+], 1236 | Bipolar DNA helicase HerA | ||
| 27 | traI | 34802..36952 [+], 2151 | Conjugative transfer protein TraI, relaxase | Relaxase, MOBH Family | |
| 28 | - | 37078..38181 [-], 1104 | putative virulence protein | ||
| 29 | traD | 38315..40150 [+], 1836 | IncF plasmid conjugative transfer protein TraD | TraD_F, T4SS component | |
| 30 | - | 40160..40720 [+], 561 | Conjugative transfer protein 234 | ||
| 31 | - | 40707..41342 [+], 636 | Conjugative transfer protein s043 | ||
| 32 | - | 41316..41435 [-], 120 | hypothetical protein | ||
| 33 | fic | 41481..42587 [+], 1107 | Fic family protein | ||
| 34 | traL | 42777..43058 [+], 282 | IncF plasmid conjugative transfer pilus assembly protein TraL | TraL_F, T4SS component | |
| 35 | traE | 43055..43681 [+], 627 | IncF plasmid conjugative transfer pilus assembly protein TraE | TraE_F, T4SS component | |
| 36 | traK | 43665..44561 [+], 897 | IncF plasmid conjugative transfer pilus assembly protein TraK | TraK_F, T4SS component | |
| 37 | traB | 44564..45853 [+], 1290 | IncF plasmid conjugative transfer pilus assembly protein TraB | TraB_F, T4SS component | |
| 38 | traV | 45928..46500 [+], 573 | Conjugative transfer protein TraV | TraV_F, T4SS component | |
| 39 | traA | 46497..46883 [+], 387 | Conjugative transfer protein TraA | TraA_F, T4SS component | |
| 40 | - | 46924..47430 [-], 507 | Acetyltransferase | ||
| 41 | - | 47421..47687 [-], 267 | hypothetical protein | ||
| 42 | - | 48056..48748 [+], 693 | Thiol:disulfide involved in conjugative transfer | TrbB_I, T4SS component | |
| 43 | traC | 48749..51148 [+], 2400 | IncF plasmid conjugative transfer pilus assembly protein TraC | TraC_F, T4SS component | |
| 44 | - | 51141..51488 [+], 348 | Conjugative transfer protein 345 | ||
| 45 | trhF | 51472..51984 [+], 513 | Conjugative signal peptidase TrhF | TraF, T4SS component | |
| 46 | traW | 51995..53119 [+], 1125 | IncF plasmid conjugative transfer pilus assembly protein TraW | TraW_F, T4SS component | |
| 47 | traU | 53103..54131 [+], 1029 | IncF plasmid conjugative transfer pilus assembly protein TraU | TraU_F, T4SS component | |
| 48 | traN | 54134..57826 [+], 3693 | IncF plasmid conjugative transfer protein TraN | TraN_F, T4SS component | |
| 49 | - | 58285..59703 [+], 1419 | hypothetical protein | ||
| 50 | herA | 59700..61739 [+], 2040 | Bipolar DNA helicase HerA | ||
| 51 | dns | 62148..62498 [-], 351 | Endonuclease I precursor | ||
| 52 | - | 63846..64109 [-], 264 | Il-IS_2, transposase | ||
| 53 | - | 64131..64643 [-], 513 | Lipoprotein signal peptidase | ||
| 54 | - | 64647..65543 [-], 897 | Cobalt-zinc-cadmium resistance protein CzcD | ||
| 55 | merR | 65639..66046 [+], 408 | Cd(II)/Pb(II)-responsive transcriptional regulator | ||
| 56 | - | 66276..66878 [-], 603 | hypothetical protein | ||
| 57 | - | 67009..67122 [-], 114 | hypothetical protein | ||
| 58 | - | 67245..67571 [+], 327 | hypothetical protein | ||
| 59 | ssb | 67587..68006 [+], 420 | Single-stranded DNA-binding protein | ||
| 60 | bet | 68086..68904 [+], 819 | Recombination protein BET | ||
| 61 | - | 68987..69130 [+], 144 | hypothetical protein | ||
| 62 | - | 69191..70207 [+], 1017 | hypothetical protein | ||
| 63 | - | 70417..71376 [+], 960 | Aerobic cobaltochelatase CobS subunit | ||
| 64 | - | 71376..72143 [+], 768 | hypothetical protein | ||
| 65 | - | 72242..73195 [+], 954 | Cobalamine biosynthesis protein | ||
| 66 | - | 73257..73697 [+], 441 | hypothetical protein | ||
| 67 | - | 73767..75422 [+], 1656 | Plasmid associated gene product | ||
| 68 | - | 75505..76002 [+], 498 | DNA repair protein RadC | ||
| 69 | - | 76002..76343 [+], 342 | hypothetical protein | ||
| 70 | - | 76435..77508 [+], 1074 | putative primase | ||
| 71 | - | 77598..78302 [+], 705 | hypothetical protein | ||
| 72 | traF | 78395..79339 [+], 945 | IncF plasmid conjugative transfer pilus assembly protein TraF | TraF_F, T4SS component | |
| 73 | traH | 79342..80730 [+], 1389 | IncF plasmid conjugative transfer pilus assembly protein TraH | TraH_F, T4SS component | |
| 74 | traG | 80734..84303 [+], 3570 | IncF plasmid conjugative transfer protein TraG | TraG_F, T4SS component | |
| 75 | merR | 84401..84790 [+], 390 | Mercuric resistance operon regulatory protein | ||
| 76 | merT | 84875..85222 [+], 348 | Mercuric transport protein, MerT | ||
| 77 | merP | 85282..85557 [+], 276 | Periplasmic mercury(+2) binding protein | ||
| 78 | merC | 85567..85980 [+], 414 | Mercuric transport protein, MerC | ||
| 79 | merA | 86018..87691 [+], 1674 | Dihydrolipoamide dehydrogenase | ||
| 80 | eexR | 87797..88228 [-], 432 | EexR | ||
| 81 | setC | 88283..88816 [-], 534 | Transcriptional activator | ||
| 82 | setD | 88813..89112 [-], 300 | Transcriptional activator | ||
| 83 | - | 89109..89657 [-], 549 | Soluble lytic murein transglycosylase and related regulatory proteins | Orf169_F, T4SS component | |
| 84 | - | 89644..90306 [-], 663 | hypothetical protein | ||
| 85 | - | 90293..91162 [-], 870 | hypothetical protein | ||
| 86 | - | 91218..91469 [-], 252 | hypothetical protein | ||
| 87 | setR | 91587..92234 [+], 648 | Putative cI prophage repressor protein | ||
|
(1) Li X; Du Y; Du P; Dai H; Fang Y; Li Z; Lv N; Zhu B; Kan B; Wang D (2016). SXT/R391 integrative and conjugative elements in Proteus species reveal abundant genetic diversity and multidrug resistance. Sci Rep. 6:37372. [PubMed:27892525] |