The graph information of SGI1-PmSCO components from JX121639 |
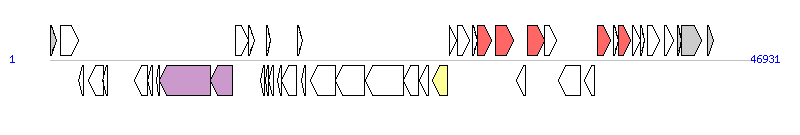 |
| Complete gene list of SGI1-PmSCO from JX121639 |
| # | Gene | Coordinates [+/-], size (bp) | Product | *Reannotation  |
| 1 | thdF | 1..468 [+], 468 | tRNA modification GTPase | |
| 2 | int | 672..1889 [+], 1218 | integrase | |
| 3 | xis | 1886..2254 [-], 369 | excisionase | |
| 4 | rep | 2612..3565 [-], 954 | replication protein | |
| 5 | S004 | 3552..3842 [-], 291 | hypothetical protein | |
| 6 | overlap with orfA characteristic of IS3 family members | 5654..6526 [-], 873 | IS1359 transposase; OrfB | |
| 7 | - | 6523..6867 [-], 345 | IS1359 transposase; OrfA | |
| 8 | S010 | 7118..7372 [-], 255 | hypothetical protein | |
| 9 | S011 | 7372..10776 [-], 3405 | conjugative transfer pilus assembly protein TraG | TraG_F, T4SS component |
| 10 | S012 | 10780..12204 [-], 1425 | conjugal transfer pilus assembly protein TraH | TraH_F, T4SS component |
| 11 | S013 | 12448..13305 [+], 858 | hypothetical protein | |
| 12 | S014 | 13302..13697 [+], 396 | hypothetical protein | |
| 13 | S015 | 14112..14381 [-], 270 | hypothetical protein | |
| 14 | S016 | 14459..14644 [-], 186 | hypothetical protein | |
| 15 | S017 | 14504..14758 [+], 255 | hypothetical protein | |
| 16 | S018 | 14655..14972 [-], 318 | hypothetical protein | |
| 17 | S019 | 15230..15526 [-], 297 | hypothetical protein | |
| 18 | S020 | 15530..16495 [-], 966 | phage integrase family protein | |
| 19 | S021 | 16588..16923 [+], 336 | hypothetical protein | |
| 20 | S022 | 16835..17110 [-], 276 | hypothetical protein | |
| 21 | S023 | 17471..19111 [-], 1641 | helicase | |
| 22 | S024 | 19129..21054 [-], 1926 | ATP-dependent endonuclease | |
| 23 | S025 | 21132..23675 [-], 2544 | subtilisin-like peptidase | |
| 24 | S026 | 23696..24685 [-], 990 | AAA ATPase, central domain protein | |
| 25 | res | 24764..25348 [-], 585 | resolvase | |
| 26 | intI1 | 25622..26635 [-], 1014 | IntI1 integrase | Integrase |
| 27 | aacCA5 | 26793..27269 [+], 477 | aminoglycoside acetyltransferase | |
| 28 | aadA7 | 27345..28142 [+], 798 | aminoglycoside adenyltransferase | |
| 29 | qacEdelta1 | 28306..28653 [+], 348 | QacEdelta1 | |
| 30 | sul1 delta | 28647..29618 [+], 972 | Sul1delta fusion protein | AR |
| 31 | floRc | 29835..31049 [+], 1215 | chloramphenicol and florfenicol resistance protein | AR |
| 32 | tetR(G) | 31256..31882 [-], 627 | TetR(G) | |
| 33 | tetA(G) | 32034..33161 [+], 1128 | TetA(G) | AR |
| 34 | - | 33182..33973 [+], 792 | putative LysR-type transcriptional regulator | |
| 35 | rcr3 | 34065..35549 [-], 1485 | Rcr3 | |
| 36 | groEL/intI1 | 35824..36477 [-], 654 | GroEL/Integrase fusion protein | |
| 37 | pse-1 | 36683..37549 [+], 867 | PSE-1 beta-lactamase | AR |
| 38 | qacEdelta1 | 37766..38113 [+], 348 | QacEdelta1 | |
| 39 | sul1 | 38107..38946 [+], 840 | Sul1 dihydropteroate synthase | AR |
| 40 | - | 39074..39574 [+], 501 | acetyltransferase | |
| 41 | - | 39598..39885 [+], 288 | hypothetical protein | |
| 42 | tnpA6100 | 40051..40845 [+], 795 | IS6100 transposase | |
| 43 | S044 | 41193..41822 [+], 630 | hypothetical protein | |
| 44 | hipB | 42098..42355 [+], 258 | transcriptional regulator | |
| 45 | hipA | 42356..43669 [+], 1314 | regulatory protein | |
| 46 | PMI3124 | 44072..44504 [+], 433 | membrane protein | |
